Active Learning in Drug Discovery: A Comparative Guide to Efficient Hit Identification Strategies
This article provides a comprehensive comparison of active learning (AL) strategies for hit discovery in drug development.
Active Learning in Drug Discovery: A Comparative Guide to Efficient Hit Identification Strategies
Abstract
This article provides a comprehensive comparison of active learning (AL) strategies for hit discovery in drug development. Aimed at researchers and scientists, it explores the foundational principles of AL as a solution to the high costs and inefficiencies of traditional high-throughput screening. The content details various methodological approaches, including uncertainty sampling, diversity-based selection, and hybrid models, supported by recent case studies across diverse targets like WDR5, SARS-CoV-2 Mpro, and CDK2. It further offers practical guidance on troubleshooting common challenges, optimizing AL workflows, and validating model performance. By synthesizing evidence from current literature, this guide serves as a strategic resource for implementing efficient, AL-driven hit discovery campaigns that significantly enrich hit rates and reduce experimental burden.
What is Active Learning and Why Does it Revolutionize Hit Discovery?
In the high-stakes field of drug discovery, the transition from traditional passive screening to intelligent, iterative active learning represents a fundamental paradigm shift in research methodology. Active learning (AL) is a machine learning framework that strategically selects the most informative data points for experimental testing, thereby compressing discovery timelines and optimizing resource allocation in hit identification [1]. Unlike passive approaches that rely on static datasets and predetermined screening libraries, active learning creates a dynamic, self-improving cycle where each experimental result informs the selection of subsequent experiments [2]. This methodological evolution is particularly critical in early-stage research where the chemical search space is vast and experimental resources are constrained. By prioritizing compounds that maximize information gain, active learning systems efficiently navigate multidimensional optimization landscapes to identify promising hit candidates with fewer experimental iterations [3]. This guide provides a comprehensive comparison of active learning strategies, delivering quantitative performance assessments and implementable experimental protocols for drug discovery researchers seeking to adopt these transformative approaches.
Active Learning Fundamentals: Core Strategies and Mechanisms
Defining the Active Learning Workflow
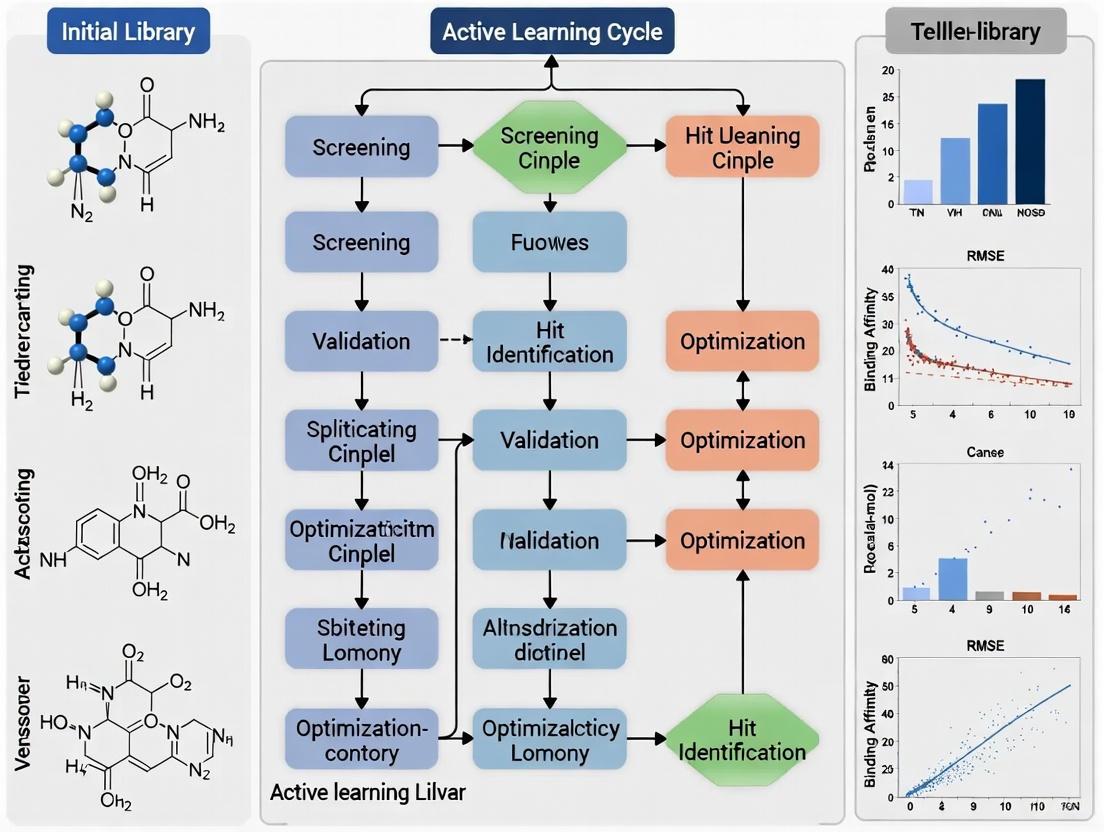
Active learning operates through an iterative feedback loop that progressively refines predictive models by incorporating strategically selected experimental data. The fundamental AL cycle begins with an initial, often small, labeled dataset used to train a preliminary machine learning model. This model then evaluates a larger pool of unlabeled candidate compounds, selecting the most "informative" samples for experimental validation based on specific query strategies [2]. Newly acquired experimental data is incorporated into the training set, updating the model for the next cycle. This continuous process of model prediction, strategic experimentation, and knowledge integration enables researchers to rapidly converge toward high-potential chemical regions while avoiding redundant testing.
The diagram below illustrates this continuous, iterative workflow:
Key Query Strategies for Hit Discovery
Active learning strategies employ various mathematical frameworks to identify which experiments will yield the maximum information gain, with the optimal approach often dependent on specific research goals and dataset characteristics:
- Uncertainty Sampling: Selects compounds where the model's prediction confidence is lowest, effectively targeting decision boundaries where clarification most improves model accuracy. In regression tasks for property prediction, methods like Monte Carlo dropout provide variance-based uncertainty estimates [2].
- Diversity Sampling: Prioritizes compounds that differ substantially from already tested examples, ensuring broad exploration of chemical space and preventing oversampling of similar molecular scaffolds.
- Expected Model Change: Selects data points that would most significantly alter the current model parameters if their labels were known, favoring instances with maximum potential influence.
- Hybrid Approaches: Combine multiple criteria, such as uncertainty and diversity, to balance exploration of unknown regions with refinement of promising areas [2]. The RD-GS strategy exemplifies this approach, demonstrating superior early-phase performance in benchmark studies [2].
Comparative Analysis of Active Learning Platforms and Performance
Quantitative Benchmarking of Active Learning Strategies
Recent comprehensive benchmarking studies reveal significant performance differences among active learning strategies when applied to materials and drug discovery problems. These evaluations typically measure how rapidly models achieve target accuracy levels as the labeled dataset grows, providing crucial insights for strategy selection.
Table 1: Performance Benchmark of Active Learning Strategies in Regression Tasks
| AL Strategy Category | Representative Methods | Early-Stage Performance | Data Efficiency | Key Advantages |
|---|---|---|---|---|
| Uncertainty-Driven | LCMD, Tree-based-R | Clearly outperforms baseline | High | Targets knowledge gaps effectively |
| Diversity-Hybrid | RD-GS | Superior to geometry-only | High | Balances exploration & exploitation |
| Geometry-Only | GSx, EGAL | Underperforms early on | Moderate | Simple implementation |
| Random Sampling | N/A | Baseline for comparison | Low | No computational overhead |
The benchmark analysis demonstrates that uncertainty-driven and diversity-hybrid strategies provide substantial early advantages, selecting more informative samples that accelerate model improvement [2]. As the labeled set grows, performance gaps between strategies typically narrow, indicating diminishing returns from advanced AL approaches under conditions of abundant data [2].
Leading AI-Driven Discovery Platforms Implementing Active Learning
Several established and emerging platforms have successfully integrated active learning principles into end-to-end drug discovery workflows, with documented progression of AI-designed candidates to clinical stages.
Table 2: AI-Driven Drug Discovery Platforms with Active Learning Components
| Platform/Company | Core AL Approach | Therapeutic Area | Clinical Stage | Key Achievement |
|---|---|---|---|---|
| Insilico Medicine | Generative chemistry + target discovery | Idiopathic pulmonary fibrosis | Phase IIa | First AI-generated drug to clinical trials (18-month discovery) [4] |
| Exscientia | Automated precision chemistry | Oncology, Immuno-oncology | Phase I/II | AI-designed candidates with ~70% faster design cycles [4] |
| Schrödinger | Physics-enabled ML design | Autoimmune diseases | Phase III | TYK2 inhibitor (zasocitinib) advancing to late-stage trials [4] |
| Recursion | Phenomics-first screening | Multiple disease areas | Multiple phases | Integrated AL with automated chemistry post-merger [4] |
| Atomwise | Structure-based deep learning | Multi-target | Preclinical | Screens billions of compounds via AtomNet architecture [5] |
These platforms exemplify the translation of active learning methodologies from theoretical concepts to practical drug discovery engines. The merger of Recursion and Exscientia in 2024 exemplifies the strategic trend toward integrating complementary AL capabilities—combining phenomic screening with generative chemistry into a unified active learning framework [4].
Experimental Protocols for Active Learning Implementation
Protocol 1: High-Throughput Screening with Active Learning
This protocol outlines the methodology for implementing active learning in high-throughput screening campaigns, optimized for identifying novel hit compounds against specific therapeutic targets.
Step 1: Initial Library Design and Feature Representation
- Procedure: Curate diverse compound library representing broad chemical space. Calculate molecular descriptors (fingerprints, molecular weight, logP, topological polar surface area) and store in structured database.
- Quality Control: Apply drug-like filters (Lipinski's Rule of Five, PAINS exclusion) to remove problematic compounds. Standardize representation using IUPAC conventions.
- Tools: RDKit or OpenBabel for descriptor calculation, KNIME or Pipeline Pilot for workflow automation.
Step 2: Baseline Model Training
- Procedure: Randomly select 0.5-1% of library (500-1000 compounds) for initial screening. Train ensemble model (random forest or gradient boosting) using 5-fold cross-validation.
- Validation: Establish performance benchmarks (R², MAE) against held-out test set of 20% of initial data.
- Tools: Scikit-learn, AutoML frameworks for automated model selection [2].
Step 3: Iterative Active Learning Cycle
- Procedure:
- Use uncertainty sampling (predictive variance) to identify 384 compounds for next screening batch.
- Conduct high-throughput assays (fluorescence, absorbance, or luminescence-based).
- Incorporate results into training dataset.
- Retrain model with expanded dataset.
- Repeat for 5-10 cycles or until performance metrics plateau.
- Batch Size: Optimize based on screening capacity (typically 384-well format).
- Stopping Criterion: Model performance plateau (ΔMAE < 5% between cycles) or maximum budget allocation.
- Procedure:
Step 4: Hit Validation and Triaging
- Procedure: Confirm actives from final model predictions using orthogonal assay methods. Apply medicinal chemistry filters for lead-like properties.
- Counter-Screening: Test against related targets to assess selectivity.
- Dose-Response: Determine IC₅₀ values for confirmed hits using 10-point concentration series.
This protocol typically reduces experimental requirements by 40-70% compared to conventional high-throughput screening while maintaining comparable hit rates [2] [3].
Protocol 2: Reaction Condition Optimization Using Active Learning
This methodology applies active learning to optimize chemical reaction conditions for parallel synthesis, particularly valuable for building compound libraries or improving synthetic routes for hit-to-lead optimization.
Step 1: Experimental Design Space Definition
- Procedure: Identify critical reaction parameters (catalyst, solvent, temperature, concentration, ligand) and define plausible ranges for each variable.
- Experimental Units: Design microtiter plates with 96-384 reaction wells covering parameter combinations.
- Objective Function: Define optimization criteria (yield, purity, enantioselectivity).
Step 2: Bayesian Optimization Implementation
- Procedure:
- Use Gaussian process regression to model reaction outcome as function of parameters.
- Employ expected improvement acquisition function to select next experiment batch.
- Run reactions using liquid handling automation.
- Analyze outcomes via UPLC-MS/HPLC.
- Update model with results.
- Parallelization: Evaluate 4-8 parameter combinations simultaneously in plate format.
- Procedure:
Step 3: Complementary Condition Identification
- Procedure: Apply maximum uncertainty sampling to identify diverse, high-performing condition sets that cover broader substrate scope [3].
- Validation: Test complementary condition sets across diverse substrate panels to verify generality.
Research demonstrates that this active learning approach identifies high-coverage reaction condition sets with 60% fewer experiments than traditional grid searches while achieving broader substrate compatibility [3].
The strategic relationships and workflow for this protocol are illustrated below:
Essential Research Reagent Solutions for Active Learning Implementation
Successful implementation of active learning workflows requires specific reagent systems and instrumentation to enable rapid iteration between computational prediction and experimental validation.
Table 3: Essential Research Reagents and Platforms for Active Learning
| Reagent/Platform Category | Specific Examples | Function in AL Workflow | Key Considerations |
|---|---|---|---|
| Compound Libraries | Diversity-oriented synthesis libraries, DNA-encoded libraries | Provides chemical space for AL exploration | Library size, diversity, drug-like properties |
| Biochemical Assay Kits | Kinase-Glo, ADP-Glo assays, fluorescence polarization kits | Enables high-throughput target-based screening | Sensitivity, dynamic range, DMSO tolerance |
| Cell-Based Assay Systems | Reporter gene assays, viability assays (CellTiter-Glo) | Provides phenotypic context for hit validation | Relevance to disease physiology, reproducibility |
| Automated Liquid Handlers | Tecan Veya, Eppendorf Research 3 neo pipette | Enables reproducible compound transfer and assay assembly | Precision, throughput, integration capabilities [6] |
| 3D Cell Culture Systems | mo:re MO:BOT platform, organoid technologies | Enhances biological relevance of phenotypic data | Reproducibility, scalability, physiological accuracy [6] |
| Protein Production Systems | Nuclera eProtein Discovery System | Rapid generation of protein targets for screening | Speed, yield, membrane protein capability [6] |
| Data Integration Platforms | Cenevo, Sonrai Analytics Discovery Platform | Unifies experimental data for AL model training | Metadata capture, interoperability, AI readiness [6] |
The transition from passive screening to intelligent, iterative active learning represents a fundamental advancement in hit discovery methodology. The comparative data presented in this guide demonstrates that uncertainty-driven and hybrid active learning strategies consistently outperform both passive approaches and random sampling, particularly during early phases of discovery campaigns where data scarcity presents the greatest challenge. Implementation success depends on selecting AL strategies aligned with specific project objectives—uncertainty sampling for rapid hit identification versus diversity-based approaches for comprehensive chemical space exploration. The integration of automated experimental systems with robust data capture infrastructure emerges as a critical enabler, allowing the full potential of active learning cycles to be realized. As AI-driven platforms continue to advance, adopting these iterative approaches will become increasingly essential for maintaining competitive advantage in drug discovery.
High-Throughput Screening (HTS) has long been the established cornerstone of early drug discovery, relying on the automated experimental testing of hundreds of thousands of physical compounds to identify initial "hits" [7]. However, this brute-force approach carries immense and often prohibitive financial and temporal costs. A single HTS campaign can cost hundreds of thousands of dollars and requires significant investments in specialized infrastructure: miniaturized assay formats (e.g., 384- or 1536-well plates), sophisticated robotics for liquid handling, and high-capacity plate readers [8] [7]. Furthermore, the hit rate in a typical HTS is notoriously low, often less than 1%, meaning vast resources are expended to find a very small number of useful starting points [8]. This high-cost, low-efficiency problem inherent to traditional HTS powerfully justifies the shift toward Artificial Intelligence (AI)-driven Active Learning (AL) strategies.
Active Learning describes a machine learning paradigm in which the algorithm intelligently selects the most informative data points to test next, creating a iterative "design-make-test-analyze" loop [1]. By prioritizing experiments that maximize learning and minimize redundancy, AL aims to drastically reduce the number of experiments and compounds required to identify high-quality hits. This guide provides an objective comparison of these two approaches, presenting experimental data and protocols to help researchers evaluate their relative merits.
Comparative Performance: HTS vs. Active Learning
The following tables summarize key performance metrics and characteristics of HTS and AL, compiled from recent large-scale studies.
Table 1: Quantitative Comparison of Screening Performance Between HTS and AI/AL Approaches
| Performance Metric | Traditional HTS | AI/Active Learning | Key Findings from Experimental Data |
|---|---|---|---|
| Typical Hit Rate | 0.001% - 0.15% [9] [8] | ~6.7% - 7.6% (AtomNet study) [9] | A 318-target study showed AI consistently achieved hit rates orders of magnitude higher than HTS benchmarks [9]. |
| Active Compound Recovery | Requires screening >99% of library | 70-90% of actives found screening only 35-50% of library [8] | Iterative screening recovers the vast majority of active compounds while testing a fraction of the collection [8]. |
| Campaign Cost | "Hundreds of thousands of dollars" per campaign [8] | Significantly lower physical testing costs; higher computational cost | AL reduces the primary cost driver: the number of physical compounds that must be synthesized and tested [8] [9]. |
| Chemical Space Explored | Limited to existing physical libraries (10^5 - 10^6 compounds) | Access to virtual, synthesis-on-demand libraries (10^9 - 10^12 compounds) [9] | AI screens a chemical space thousands of times larger than HTS, accessing novel scaffolds not in any physical library [9]. |
Table 2: Characteristics and Resource Requirements of HTS vs. Active Learning
| Characteristic | Traditional HTS | Active Learning |
|---|---|---|
| Primary Approach | Experimental, brute-force screening of a full static library | Computational, iterative selection of informative subsets |
| Automation Focus | Liquid handling robotics, plate readers | Algorithmic selection and model retraining |
| Key Assay Metric | Z'-factor (0.5-1.0 indicates excellent assay) [7] | Model performance (e.g., F1 score, predictive accuracy for hit identification) [10] |
| Data Utilization | Single-use for a single campaign; often underutilized | Cumulative; each experiment improves the model for subsequent cycles |
| Scaffold Novelty | Limited to known and available chemotypes | High; capable of generating novel, drug-like scaffolds not based on known bioactives [9] |
Experimental Evidence for Active Learning Efficiency
Protocol: Iterative Screening for Hit Finding
A seminal study demonstrated a practical AL protocol for hit identification, which can be implemented with standard computational resources [8].
- Step 1: Initial Diverse Set Screening. The process begins by screening a small, diverse subset (e.g., 10-15%) of the compound library, selected using a MaxMin algorithm to ensure broad chemical coverage [8].
- Step 2: Model Training. The results (active/inactive labels) from this initial batch are used to train a machine learning model. The study found Random Forest (RF) algorithms performed best, even on standard desktops [8].
- Step 3: Exploitation and Exploration. The trained model predicts hit probabilities for all remaining unscreened compounds. The next batch for testing is selected using an 80/20 rule: 80% of the batch is filled with compounds ranked highest for predicted activity (exploitation), while 20% are chosen randomly from the remainder to explore under-sampled chemical space and improve the model (exploration) [8].
- Step 4: Iterative Model Refinement. Steps 2 and 3 are repeated for a small number of iterations (e.g., 3-6 cycles). With each iteration, the model becomes more accurate at predicting active compounds [8].
Results: This workflow, screening just 35% of the total library over three iterations, recovered a median of 70% of all active compounds. Increasing the screened portion to 50% raised the median recovery to 80% of actives [8]. This demonstrates a massive reduction in experimental effort for a minimal loss in potential hits.
Protocol: Large-Scale Virtual Screening with the AtomNet Model
A 2024 study involving 318 targets provides robust, large-scale evidence for replacing initial HTS with convolutional neural networks (CNNs) [9].
- Step 1: Virtual Screening. The AtomNet CNN, a structure-based deep learning model, was used to score protein-ligand interactions for billions of compounds in a virtual, synthesis-on-demand library [9].
- Step 2: Algorithmic Compound Selection. The top-ranked molecules were clustered to ensure diversity. The highest-scoring exemplars from each cluster were selected algorithmically, without manual cherry-picking [9].
- Step 3: Experimental Validation. Selected compounds were synthesized and tested in biochemical or cell-based assays at contract research organizations (CROs). Assays included standard additives (e.g., Tween-20, DTT) to mitigate common interference artifacts [9].
Results: Across 22 internal drug discovery projects, the approach achieved a 91% success rate in identifying reconfirmed hits. The average dose-response hit rate was 6.7%, vastly exceeding typical HTS hit rates. Crucially, this success was demonstrated for targets without known binders, high-quality X-ray structures, or both, addressing a historical limitation of computational methods [9].
Visualizing the Workflows
The fundamental difference between HTS and AL lies in their workflow structure. The diagrams below illustrate the linear, resource-heavy nature of HTS versus the adaptive, efficient cycle of AL.
HTS Linear Process - Figure 1: The traditional HTS process is a linear, single-pass workflow that requires screening an entire library before any analysis, leading to high upfront costs.
AL Iterative Cycle - Figure 2: The Active Learning workflow is an iterative cycle where experimental data continuously refines a model, which then intelligently selects the next most valuable experiments, dramatically increasing efficiency.
The Scientist's Toolkit: Essential Research Reagents & Solutions
Successful implementation of an AL strategy, particularly the experimental validation phase, relies on key reagents and tools. The following table details essential components for setting up the necessary screening assays.
Table 3: Key Research Reagent Solutions for Screening Assays
| Reagent / Solution | Function in Screening | Application Notes |
|---|---|---|
| Transcreener ADP2 Assay | Universal biochemical assay for detecting ADP production; applicable to kinases, ATPases, GTPases, and other enzymes. | Flexible detection: can use Fluorescence Polarization (FP), Fluorescence Intensity (FI), or TR-FRET readouts. Enables potency (IC50) and residence time measurements [7]. |
| Miniaturized Assay Plates (384-/1536-well) | High-density microplates that minimize reagent consumption and enable automated, high-throughput testing. | Standard format for modern HTS and follow-up screening. Requires compatible liquid handling robotics and plate readers [7]. |
| Assay Interference Mitigants | Reagents like Tween-20, Triton-X 100, and Dithiothreitol (DTT) added to assays to reduce false positives. | Counteract common compound interference mechanisms such as aggregation, promiscuous inhibition, and oxidation [9]. |
| Target-Specific Biochemical Kits | Pre-optimized assay systems for specific target classes (e.g., kinases, proteases, epigenetic targets). | Reduce development time and ensure robust performance (high Z'-factor) for primary screening [7]. |
| Cell-Based Assay Reagents | Reagents for cell viability (e.g., Cell Titer-Glo), reporter gene assays, and high-content imaging. | Critical for phenotypic screening and assessing compound activity in a more physiologically relevant context [8] [7]. |
The empirical data and comparative analysis presented in this guide build a compelling case. The high-cost problem of Traditional HTS—characterized by low hit rates, immense physical screening costs, and limited chemical exploration—is no longer a necessary burden in hit discovery. Active Learning and other AI-driven approaches offer a validated, efficient, and powerful alternative. By adopting an iterative, intelligence-guided strategy, researchers can substantially reduce costs and timelines while accessing richer, more novel chemical space, ultimately accelerating the journey from concept to clinical candidate.
In the resource-intensive field of drug discovery, active learning has emerged as a powerful strategy to accelerate hit identification. This guide compares two advanced active learning frameworks—one leveraging generative AI and another employing a multi-task balanced-ranking strategy—against non-iterative high-throughput screening (HTS), providing the experimental data and protocols to underpin your research decisions.
Performance Comparison: Active Learning Strategies vs. Traditional HTS
The table below summarizes the key performance metrics of two distinct active learning methodologies compared to a primary HTS screen, demonstrating the significant efficiency gains of an iterative approach.
Table 1: Experimental Performance Metrics Across Discovery Strategies
| Strategy / Framework Name | Core Approach | Target Protein(s) | Hit Rate | Key Experimental Validation | Reference |
|---|---|---|---|---|---|
| Generative AI with Active Learning | VAE with nested AL cycles guided by chemoinformatic & physics-based oracles [11] | CDK2, KRAS | 8 out of 9 synthesized molecules showed in vitro activity (1 nanomolar) [11] | Synthesis & bioassay of generated molecules; CDK2: 8/9 active; KRAS: 4 in silico actives identified [11] | [11] |
| ChemScreener | Multi-task active learning with Balanced-Ranking acquisition [12] | WDR5 | Average 5.91% (Range: 3-10%) [12] | 44 hits advanced to dose-response; over 50% of top hits validated as binders by DSF; 3 novel scaffold series identified [12] | [12] |
| Primary HTS (Baseline) | Non-iterative screening of a large compound library [12] | WDR5 | 0.49% [12] | N/A (Baseline for comparison) | [12] |
Detailed Experimental Protocols
To ensure reproducibility and provide depth for scientific evaluation, here are the detailed methodologies for the two featured active learning frameworks.
Protocol 1: Generative AI with Nested Active Learning Cycles
This protocol is designed for de novo molecular generation and optimization for a specific target [11].
- 1. Data Representation & Initial Training: Represent training molecules as tokenized SMILES strings, converted into one-hot encoding vectors. The Variational Autoencoder (VAE) is first trained on a general molecular dataset to learn viable chemistry, then fine-tuned on a target-specific initial training set [11].
- 2. Workflow Execution (Nested Active Learning): The core of the method involves two nested cycles [11]:
- Inner AL Cycle (Chemical Optimization): The VAE generates new molecules. An oracle comprising chemoinformatic predictors evaluates them for drug-likeness, synthetic accessibility (SA), and dissimilarity from the training set. Molecules passing these filters are added to a "temporal-specific set" used to fine-tune the VAE. This cycle repeats, iteratively improving chemical properties [11].
- Outer AL Cycle (Affinity Optimization): After several inner cycles, the accumulated molecules in the temporal-specific set are evaluated by a physics-based affinity oracle (e.g., molecular docking simulations). Molecules with favorable docking scores are promoted to a "permanent-specific set," which is used for the next round of VAE fine-tuning, initiating a new outer cycle with nested inner cycles [11].
- 3. Candidate Selection & Validation: After multiple outer AL cycles, the most promising molecules from the permanent-specific set undergo rigorous filtration. This includes advanced molecular modeling (e.g., PELE simulations for binding pose refinement) and Absolute Binding Free Energy (ABFE) calculations. Top candidates are then selected for chemical synthesis and in vitro bioassay validation [11].
Protocol 2: ChemScreener's Balanced-Ranking for HTS Follow-Up
This protocol is designed for efficiently screening large, diverse chemical libraries after a primary HTS, using iterative single-dose assays [12].
- 1. Primary HTS & Model Initialization: Conduct a primary high-throughput screen to establish a baseline hit rate and gather initial activity data. Use this data to initialize a multi-task predictive model [12].
- 2. Iterative Screening & Model Updating: For each iterative cycle [12]:
- Prediction: The model predicts activity for all compounds not yet tested.
- Selection (Balanced-Ranking): A subset of compounds is selected for experimental testing based on an acquisition function that balances exploration (prioritizing compounds with high model uncertainty, often calculated via ensemble methods) and exploitation (prioritizing compounds with high predicted activity). This strategy enriches hit rates while exploring novel chemistry [12].
- Model Refinement: The selected compounds are tested in a single-dose assay (e.g., HTRF). The new experimental data (activity labels) is added to the training set, and the model is retrained before the next cycle [12].
- 3. Hit Validation & Characterization: Consolidate all hits from the iterative cycles and retest them alongside close analogs in a full dose-response assay. Confirm binding through orthogonal biophysical methods (e.g., Differential Scanning Fluorimetry - DSF) and cluster confirmed hits to identify novel scaffold series [12].
Workflow Visualization
The following diagrams illustrate the logical structure of the two core active learning principles, using the specified color palette.
Generative AI Active Learning Workflow
Balanced-Ranking Active Learning Workflow
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 2: Key Reagents and Materials for Active Learning-Driven Hit Discovery
| Item | Function / Application in the Workflow |
|---|---|
| WDR5 Protein | The target protein used in the HTRF-based iterative screening campaign to identify potential inhibitors [12]. |
| CDK2 / KRAS Proteins | Oncology-related target proteins used for benchmarking the generative AI workflow, involving docking and bioassays [11]. |
| HTRF Assay Kit | Used for the primary and iterative single-dose screens (e.g., for WDR5) to rapidly quantify compound activity and provide data for model refinement [12]. |
| Differential Scanning Fluorimetry (DSF) | An orthogonal, biophysical method used post-screening to validate true binding of identified hits to the target protein (e.g., WDR5), countering assay artifacts [12]. |
| Molecular Docking Software | Serves as the physics-based affinity oracle in the generative AI workflow, providing a computationally efficient estimate of binding potential to prioritize compounds [11]. |
| VAE & Predictive Models | The core computational engines. The VAE generates novel molecules, while the predictive models (e.g., Random Forest, Deep Learning) forecast activity and uncertainty for the screening library [12] [11]. |
In the face of vast chemical space and constrained research budgets, active learning (AL) has emerged as a transformative machine learning approach for hit discovery. AL operates through an iterative feedback process that selectively identifies the most valuable data points for experimental testing, effectively optimizing resource allocation [13]. This methodology stands in stark contrast to traditional high-throughput screening (HTS), which treats all compounds equally and often wastes resources on testing chemically redundant or uninformative molecules. The fundamental premise of AL is that by prioritizing uncertainty and diversity in compound selection, models can learn more efficiently, requiring far fewer experimental cycles to identify promising hit compounds [14].
The strategic implementation of AL addresses three critical challenges in modern drug discovery: the exponentially expanding chemical space that exceeds practical testing capacity, the high costs and time requirements of wet-lab experimentation, and the scarcity of labeled bioactivity data for model training [13]. By framing hit discovery as an iterative, model-guided exploration rather than a one-shot screening endeavor, AL enables research teams to accelerate project timelines, significantly enrich hit rates from chemical libraries, and substantially reduce operational costs associated with compound acquisition and testing.
Quantitative Performance Comparison
Empirical studies across multiple drug discovery campaigns demonstrate that active learning strategies consistently outperform traditional screening approaches across key performance metrics. The following tables synthesize quantitative results from recent implementations, highlighting the significant advantages of AL in hit discovery.
Table 1: Comparative Performance of Active Learning vs. Traditional Screening
| Screening Approach | Average Hit Rate | Hit Rate Improvement | Number of Hits Identified | Library Size | Reference |
|---|---|---|---|---|---|
| Primary HTS Screen | 0.49% | Baseline | Not specified | Not specified | [12] |
| ChemScreener AL | 5.91% (avg: 3-10%) | 12x increase | 104 hits | 1,760 compounds | [12] |
| Transcriptomics AL | 13-17x higher | 13-17x increase | Significant increase | Not specified | [15] |
| Random Sampling | Baseline | Baseline | Baseline | Various | [16] |
| Active Learning (various strategies) | Significantly higher | 30-70% more efficient than random | Increased | Various | [16] [14] |
Table 2: Efficiency Gains of Active Learning Strategies
| Performance Metric | Traditional Screening | Active Learning | Improvement | Application Context |
|---|---|---|---|---|
| Labeling efficiency | Baseline | 30-70% reduction in labels needed | Significant | General ML [14] |
| Hit identification speed | Baseline | Much earlier identification | Substantial | Anti-cancer drug screening [16] |
| Model performance gain | Baseline | Faster improvement per labeled sample | Significant | Drug response prediction [16] |
| Experimental cost | High | Reduced through focused experimentation | Substantial | Virtual screening [13] |
The data reveal that AL implementations achieve substantially higher hit rates compared to conventional methods. The ChemScreener workflow demonstrated particularly impressive results, increasing hit rates from a baseline of 0.49% in primary HTS to an average of 5.91% (ranging from 3-10% across cycles) [12]. This represents an approximate 12-fold enrichment in hit discovery efficiency. Similarly, an AL framework leveraging transcriptomics for phenotypic screening outperformed state-of-the-art models, translating to a 13-17x increase in phenotypic hit-rate across two hematological discovery campaigns [15].
Beyond hit rate enrichment, AL methodologies demonstrate remarkable efficiency in resource utilization. Research indicates that well-designed AL pipelines can reduce labeling requirements by 30-70% while maintaining or improving model performance compared to exhaustive screening approaches [14]. This efficiency translates directly to cost savings through reduced compound testing, smaller library requirements, and shorter discovery timelines.
Experimental Protocols and Methodologies
ChemScreener Workflow for WDR5 Inhibitor Discovery
The ChemScreener experimental protocol exemplifies a sophisticated implementation of active learning for hit discovery. The methodology employed a multi-task active learning workflow designed for early drug discovery across large, diverse chemical libraries [12]. The process commenced with an initial training set of known bioactive compounds, followed by iterative cycles of model prediction and experimental validation.
Balanced-Ranking Acquisition Strategy: ChemScreener's core innovation lies in its acquisition function, which leverages ensemble uncertainty to balance exploration of novel chemistry with exploitation of predicted activity [12]. This strategy simultaneously prioritizes compounds with high predicted activity against WDR5 while ensuring chemical diversity by selecting structures from underrepresented regions of chemical space. The ensemble model generated multiple predictions for each compound, with disagreement among models serving as a proxy for uncertainty.
Experimental Validation Cycle: Each AL iteration consisted of several key steps: (1) model training on existing bioactivity data; (2) prediction on untested compounds in the library; (3) selection of compounds for testing using the Balanced-Ranking strategy; (4) experimental testing via HTRF assays; and (5) model updating with new experimental results [12]. This cycle repeated five times in the WDR5 case study, with each iteration refining the model's understanding of structure-activity relationships.
Hit Confirmation Protocol: Promising hits from single-dose HTRF screens underwent rigorous validation through multiple orthogonal assays. The confirmation workflow included: (1) compound consolidation with close analogs; (2) retesting in dose-response format; (3) counter-screening in HTRF assays to exclude artifacts; and (4) validation of binding via differential scanning fluorimetry (DSF) [12]. This comprehensive approach ensured that identified hits represented genuine binders rather than assay artifacts.
Anti-Cancer Drug Response Prediction Framework
A comprehensive investigation of AL strategies for anti-cancer drug response prediction provides another exemplary protocol. This study focused on constructing drug-specific response prediction models for cancer cell lines, with the dual objectives of improving prediction model performance and efficiently identifying effective treatments [16].
Cell Line Selection Strategies: The researchers implemented and compared multiple AL approaches for selecting cell lines for screening, including: (1) random sampling (baseline); (2) greedy sampling (selecting cell lines with highest predicted sensitivity); (3) uncertainty sampling (prioritizing predictions with highest model uncertainty); (4) diversity sampling (maximizing representation of different cancer types); and (5) hybrid approaches combining uncertainty and diversity criteria [16].
Data Sources and Processing: The analysis utilized the Cancer Therapeutics Response Portal v2 (CTRP) dataset, encompassing 494 drugs, 812 cell lines, and over 318,000 dose-response experiments [16]. Cell lines were represented by multi-omic features, including gene expression, mutations, and copy number variations, while drugs were encoded using molecular fingerprints and descriptors.
Evaluation Metrics: Performance was assessed using two primary metrics: (1) the number of identified hits (validated responsive treatments) selected during the AL process, and (2) the performance of response prediction models trained on the data selected by each strategy [16]. The results demonstrated that most AL strategies significantly outperformed random selection for identifying effective treatments, with hybrid approaches generally showing the most robust performance across diverse drug classes.
Workflow Visualization
Active Learning Cycle for Hit Discovery
The following diagram illustrates the iterative feedback loop that forms the core of active learning methodologies in drug discovery:
Active Learning Cycle for Hit Discovery
Balanced-Ranking Acquisition Strategy
ChemScreener's innovative Balanced-Ranking strategy combines exploration and exploitation through the following decision process:
Balanced-Ranking Acquisition Strategy
Research Reagent Solutions Toolkit
Successful implementation of active learning for hit discovery requires specialized research tools and reagents. The following table details essential components used in the featured studies:
Table 3: Essential Research Reagents and Tools for Active Learning-Driven Hit Discovery
| Reagent/Tool | Function in Workflow | Application Example |
|---|---|---|
| HTRF Assay Kits | Measure compound-protein interaction in high-throughput format | WDR5-binding confirmation in ChemScreener study [12] |
| Cancer Cell Line Panels | Provide diverse biological context for compound screening | CTRP database with 812 cell lines for response prediction [16] |
| Molecular Fingerprints | Encode chemical structures for machine learning models | Extended-connectivity fingerprints for structure-activity modeling [13] |
| Differential Scanning Fluorimetry (DSF) | Validate binding through thermal stability shifts | Orthogonal confirmation of WDR5 binders [12] |
| Transcriptomics Profiling | Generate multi-omic features for cell line characterization | Predictive features for phenotypic screening [15] |
| Automated Screening Systems | Enable high-throughput experimental testing | Implementation of iterative AL cycles [12] |
| Ensemble Modeling Software | Generate predictions with uncertainty estimates | Balanced-ranking acquisition in ChemScreener [12] |
Comparative Analysis of Active Learning Strategies
The implementation details of AL strategies significantly impact their performance in hit discovery applications. Different sampling approaches offer distinct advantages and limitations:
Uncertainty Sampling prioritizes compounds where the model shows highest prediction uncertainty, typically targeting decision boundary regions [14]. This approach efficiently improves model accuracy but may overfocus on outliers or noisy data points. In anti-cancer drug response prediction, uncertainty sampling demonstrated particular effectiveness for early identification of responsive treatments [16].
Diversity Sampling selects compounds that maximize structural or functional diversity in the training set, ensuring broad coverage of chemical space [14]. This approach mitigates the redundancy inherent in large chemical libraries but may deliver slower improvements in hit rates compared to uncertainty-focused methods.
Hybrid Approaches combine multiple criteria to balance competing objectives. The Balanced-Ranking strategy used in ChemScreener exemplifies this category, simultaneously considering predicted activity (exploitation) and model uncertainty (exploration) [12]. Similarly, research in other domains has successfully combined uncertainty sampling with clustering to ensure diverse selection of informative samples [14]. These hybrid methods generally demonstrate more robust performance across diverse drug targets and chemical libraries.
Committee-Based Strategies employ multiple models to quantify disagreement as a measure of uncertainty [14]. While computationally intensive, this approach can yield more reliable uncertainty estimates than single-model methods, particularly for complex structure-activity relationships.
The comparative performance of these strategies depends on factors including target biology, chemical library diversity, and available training data. The research consistently indicates that most AL strategies significantly outperform random selection, with hybrid approaches generally delivering the most balanced performance across multiple optimization objectives [16].
The accumulated evidence from recent studies firmly establishes active learning as a transformative methodology for hit discovery in drug development. Through strategic compound selection and iterative model refinement, AL implementations consistently achieve substantial hit rate enrichment, significant timeline acceleration, and meaningful cost reduction compared to traditional screening approaches.
The case studies examined demonstrate that AL can increase hit rates by an order of magnitude—from under 0.5% in conventional HTS to 3-10% in AL-guided campaigns [12]—while simultaneously reducing experimental requirements by 30-70% [14]. These improvements directly address the fundamental challenges of modern drug discovery: navigating vast chemical spaces with constrained resources.
The successful application of AL across diverse target classes (including WDR5 and various anti-cancer targets) and screening methodologies (binding assays, phenotypic screens) underscores its versatility and generalizability [12] [16] [15]. As drug discovery continues to confront increasingly challenging targets and growing chemical spaces, the strategic implementation of active learning methodologies will become increasingly essential for maintaining research productivity and therapeutic innovation.
A Practical Guide to Active Learning Strategies and Their Real-World Applications
In the high-stakes field of drug discovery, researchers face the monumental challenge of identifying potential therapeutic compounds from libraries containing billions of molecules. Traditional high-throughput screening methods are prohibitively expensive and time-consuming, often requiring substantial resources to evaluate even a fraction of available chemical space. Active learning has emerged as a powerful strategy to address this inefficiency by enabling iterative, data-driven selection of the most informative compounds for experimental testing. Within this paradigm, uncertainty sampling represents a foundational approach that prioritizes compounds for which the current predictive model exhibits maximum uncertainty, thereby targeting samples most likely to improve model performance with each iteration.
The application of uncertainty sampling in drug discovery is particularly valuable in scenarios characterized by extreme class imbalance, such as synergistic drug combination screening where synergistic pairs represent only 1.47-3.55% of all possible combinations [17]. In such contexts, random sampling strategies waste significant resources on non-informative examples, while well-designed uncertainty sampling methods can discover 60% of synergistic drug pairs by exploring only 10% of the combinatorial space [17]. This efficiency gain translates directly to reduced experimental costs and accelerated research timelines, making uncertainty sampling an indispensable tool for modern drug development pipelines.
Theoretical Foundations of Uncertainty Sampling
Core Principles and Mechanisms
Uncertainty sampling operates on a fundamentally simple yet powerful principle: in pool-based active learning, an algorithm sequentially queries the labels of those instances for which its current prediction model is maximally uncertain [18] [19]. This approach stands in contrast to other active learning strategies that might prioritize representativeness or diversity. The underlying assumption is that by resolving the model's areas of greatest uncertainty, each newly acquired data point will provide maximum information gain, leading to more efficient model improvement with fewer labeled examples.
The effectiveness of uncertainty sampling hinges on properly defining and quantifying "uncertainty" within the specific context of the prediction task and loss function [19]. Traditional probabilistic measures include:
- Least confidence sampling: Selecting instances where the model assigns the highest probability to any class is lowest
- Margin sampling: Choosing examples with the smallest difference between the two highest class probabilities
- Entropy sampling: Prioritizing instances with maximum class probability entropy
Recent theoretical work has established that uncertainty sampling essentially optimizes against an "equivalent loss" that depends on both the chosen uncertainty measure and the original loss function [19]. This perspective provides a mathematical foundation for understanding the behavior and performance of different uncertainty sampling variants.
Uncertainty Types and Their Implications
A critical advancement in uncertainty sampling theory recognizes that not all uncertainty is equivalent. Modern approaches distinguish between epistemic uncertainty (reducible uncertainty stemming from limited training data) and aleatoric uncertainty (irreducible uncertainty inherent in the data itself) [18]. This distinction is particularly relevant in drug discovery, where epistemic uncertainty might indicate promising exploration areas for model improvement, while high aleatoric uncertainty might signal inherently noisy or unpredictable biological systems.
Evidential Deep Learning (EDL) represents one approach to separately modeling these uncertainty types. As implemented in the EviDTI framework for drug-target interaction prediction, EDL provides direct uncertainty quantification without relying on computationally expensive random sampling [20]. This capability allows researchers to not only identify uncertain predictions but also understand the nature of that uncertainty, enabling more informed decision-making about which compounds to prioritize for experimental validation.
Comparative Analysis of Uncertainty Sampling Strategies
Standard Uncertainty Sampling Methods
Traditional uncertainty sampling methods form the foundation upon which more advanced techniques are built. These approaches typically rely on the probabilistic outputs of classification models to identify uncertain instances. In the context of drug discovery, these methods have been applied to various prediction tasks including drug-target interactions, synergy prediction, and molecular property estimation.
The fundamental limitation of these standard approaches lies in their potential to create sample imbalance in multi-class scenarios, where high-frequency or high-complexity classes become overrepresented while low-frequency classes suffer from insufficient representation [21]. This distributional imbalance can severely constrain model performance, resulting in significantly diminished predictive capability for underrepresented molecular classes and ultimately affecting overall accuracy. Despite this limitation, standard uncertainty sampling remains widely used due to its computational efficiency and straightforward implementation.
Evidential Deep Learning for Uncertainty Quantification
The EviDTI framework represents a significant advancement in uncertainty-aware modeling for drug-target interaction prediction [20]. By employing evidential deep learning, EviDTI addresses a critical challenge in traditional deep learning models: the tendency to produce overconfident and incorrect predictions for novel, unseen drug-target interactions. The framework integrates multiple data dimensions, including drug 2D topological graphs, 3D spatial structures, and target sequence features, through a specialized architecture comprising protein feature encoders, drug feature encoders, and an evidential layer.
Experimental results demonstrate EviDTI's competitive performance against 11 baseline models across three benchmark datasets: DrugBank, Davis, and KIBA [20]. On the challenging KIBA dataset, characterized by significant class imbalance, EviDTI outperformed the best baseline model by 0.6% in accuracy, 0.4% in precision, 0.3% in Matthews correlation coefficient, and 0.4% in F1 score [20]. More importantly, the well-calibrated uncertainty estimates provided by EviDTI's evidential approach enable prioritization of drug-target interactions with higher confidence predictions for experimental validation, potentially enhancing the efficiency of drug discovery pipelines.
Category-Enhanced Uncertainty Sampling
To address the class imbalance limitations of traditional uncertainty sampling, Wang et al. proposed an enhanced approach that integrates category information with uncertainty measures [21]. This method employs a pre-trained VGG16 architecture and cosine similarity metrics to efficiently extract category features without requiring additional model training. The framework combines these features with traditional uncertainty measures to ensure balanced sampling across classes while maintaining computational efficiency.
In object detection tasks, this category-enhanced approach achieves competitive mean average precision scores while ensuring balanced category representation [21]. For image classification, the method achieves accuracy comparable to state-of-the-art approaches while reducing computational overhead by up to 80% [21]. Although developed for computer vision applications, the underlying principle of incorporating category information to mitigate sampling bias has direct relevance to drug discovery, particularly in scenarios involving multiple target classes or therapeutic areas.
Calibrated Uncertainty Sampling
A recently proposed innovation addresses the critical issue of miscalibrated uncertainty estimates in deep neural networks [22]. Standard uncertainty sampling approaches can be misled when a model's uncertainty estimates are poorly calibrated on unlabeled data, leading to suboptimal sample selection and reduced active learning performance. The calibrated uncertainty sampling method estimates calibration errors and queries samples with the highest calibration error before leveraging the model's uncertainty estimates.
Theoretical analysis shows that active learning with this acquisition function eventually leads to a bounded calibration error on both the unlabeled pool and unseen test data [22]. Empirically, the approach surpasses other acquisition function baselines by achieving lower calibration and generalization errors across pool-based active learning settings. This focus on calibration is particularly relevant to drug discovery, where reliable confidence estimates are essential for making costly decisions about which compounds to synthesize and test experimentally.
Thompson Sampling and Roulette Wheel Selection
For screening ultralarge combinatorial libraries, Thompson sampling has emerged as a valuable probabilistic search method that operates in reagent space rather than product space [23]. This approach associates each chemical building block with a probability distribution, sampling promising building blocks more frequently to guide the search toward productive areas of chemical space. Recent enhancements to this method introduce a roulette wheel selection approach combined with thermal cycling to balance greedy search and diversity-driven exploration.
In extensive benchmarking involving 2.18 billion evaluations across 20 reactions applied to 109 shape-based virtual screens, the enhanced Thompson sampling approach matched greedy scheme performance on two-component libraries and outperformed it on most three-component libraries [23]. The method parallelizes with approximately linear scaling, enabling practical screening of ultralarge combinatorial spaces containing billions or trillions of compounds. This capability is particularly valuable for hit expansion in early drug discovery, where efficiently navigating vast chemical spaces is essential.
Table 1: Performance Comparison of Uncertainty Sampling Methods in Drug Discovery Applications
| Method | Key Innovation | Application Context | Performance Advantages | Limitations |
|---|---|---|---|---|
| Standard Uncertainty Sampling | Querying by model uncertainty | General classification tasks | Computational efficiency, straightforward implementation | Prone to class imbalance, uncalibrated uncertainties |
| EviDTI (Evidential Deep Learning) | Integrated uncertainty quantification | Drug-target interaction prediction | Competitive accuracy (82.02% on DrugBank), well-calibrated uncertainty estimates | Architectural complexity, computational requirements |
| Category-Enhanced Sampling | Integration of category information | Multi-class scenarios | Reduces computational overhead by up to 80%, mitigates class imbalance | Requires category feature extraction |
| Calibrated Uncertainty Sampling | Explicit calibration error estimation | Pool-based active learning | Lower calibration and generalization errors | Additional computation for calibration estimation |
| Enhanced Thompson Sampling | Roulette wheel selection with thermal cycling | Ultralarge combinatorial library screening | Identifies >90% of top molecules with 0.1-1% library evaluation, linear scaling | Performance varies by library composition |
Experimental Protocols and Methodologies
EviDTI Framework Implementation
The EviDTI framework employs a multi-modal architecture that integrates diverse molecular representations for enhanced drug-target interaction prediction [20]. The implementation comprises three main components:
Protein Feature Encoder: This module utilizes the protein language pre-trained model ProtTrans as the initial encoder to generate target representations. These representations undergo further feature extraction through a light attention module, providing insights into local interactions at the residue level.
Drug Feature Encoder: This component processes both 2D topological information and 3D structural information of drug molecules. For 2D representations, the pre-trained model MG-BERT generates initial encodings that are subsequently processed by a 1D convolutional neural network. For 3D structural information, the spatial structure is converted into an atom-bond graph and a bond-angle graph, with representations obtained through a GeoGNN module.
Evidential Layer: The concatenated target and drug representations are fed into this layer, which outputs the parameter α used to calculate both prediction probability and corresponding uncertainty value.
Experimental validation followed rigorous protocols using three benchmark datasets: DrugBank, Davis, and KIBA [20]. These datasets were randomly divided into training, validation, and test sets in a ratio of 8:1:1. Performance was evaluated using seven metrics: accuracy, recall, precision, Matthews correlation coefficient, F1 score, area under the ROC curve, and area under the precision-recall curve.
Active Learning for Synergistic Drug Combination Discovery
The experimental protocol for active learning in synergistic drug discovery involves several key components [17]. Researchers typically use the Oneil dataset, which contains 15,117 measurements comprising 38 drugs and 29 cell lines with 3.55% synergistic drug pairs (defined as Loewe synergy score >10). The active learning process proceeds iteratively through multiple batches, with model updates between batches incorporating newly acquired experimental data.
Critical protocol parameters include:
- Batch size: Smaller batch sizes (typically 1-5% of total samples) yield higher synergy discovery rates
- Molecular features: Morgan fingerprints with addition operations demonstrated highest prediction performance
- Cellular features: Gene expression profiles from GDSC database significantly improve prediction quality
- AI algorithms: Ranging from parameter-light (logistic regression) to parameter-heavy (transformers)
This protocol demonstrated that 1,488 measurements scheduled with active learning recovered 60% (300 out of 500) synergistic combinations, saving 82% of experimental resources compared to random screening (which would require 8,253 measurements to obtain the same number of synergies) [17].
Enhanced Thompson Sampling for Library Screening
The experimental methodology for enhanced Thompson sampling with roulette wheel selection involves several stages [23]:
Warmup Cycle: Reagents from each reaction component are placed in a matrix, with each reagent selected for a minimum number of molecules. This stage establishes initial probability distributions for building blocks.
Search Cycle: The method samples random scores from probability distributions of each building block, selects building blocks with highest sampled scores, combines them to produce virtual reaction products, and evaluates these products using 3D similarity searches.
Probability Update: Evaluation results are used to adjust probability distributions of related building blocks using a Boltzmann-weighted average rather than arithmetic mean.
The benchmarking process involved 109 queries against twenty distinct 1-million-compound libraries using ROCS (Rapid Overlay of Chemical Structures) for shape-based similarity assessment [23]. This extensive evaluation encompassed 2.18 billion assessments to validate method performance across diverse chemical spaces.
Table 2: Experimental Results Across Different Uncertainty Sampling Applications
| Application Domain | Dataset | Baseline Performance | Uncertainty Sampling Performance | Key Metric |
|---|---|---|---|---|
| Drug-Target Interaction Prediction | DrugBank | Varies by baseline model | Accuracy: 82.02%, Precision: 81.90% | MCC: 64.29% [20] |
| Drug-Target Interaction Prediction | Davis | Varies by baseline model | Exceeds best baseline by 0.8% in accuracy, 0.6% in precision | MCC: +0.9%, F1: +2% [20] |
| Drug-Target Interaction Prediction | KIBA | Varies by baseline model | Outperforms best baseline by 0.6% in accuracy, 0.4% in precision | MCC: +0.3%, F1: +0.4% [20] |
| Synergistic Drug Discovery | Oneil | Random sampling requires 8,253 measurements for 300 synergies | Active learning finds 300 synergies in 1,488 measurements | 82% resource saving [17] |
| Combinatorial Library Screening | 20 virtual libraries | Exhaustive screening required | Identifies >90% of top 100 molecules with 0.1-1% evaluation | Linear scaling with CPUs [23] |
Workflow Visualization: Uncertainty Sampling in Drug Discovery
The following diagram illustrates the generalized workflow for uncertainty sampling in drug discovery applications, integrating elements from the EviDTI framework, synergistic combination discovery, and combinatorial library screening:
Uncertainty Sampling Workflow in Drug Discovery
Table 3: Key Research Reagents and Computational Tools for Uncertainty Sampling Implementation
| Resource Category | Specific Tool/Resource | Function in Uncertainty Sampling | Application Context |
|---|---|---|---|
| Bioactivity Datasets | DrugBank Database | Provides known drug-target interactions for model training and validation | Drug-target interaction prediction [20] |
| Bioactivity Datasets | Davis Dataset | Contains kinase inhibition data for evaluation | Drug-target interaction benchmarking [20] |
| Bioactivity Datasets | KIBA Dataset | Provides kinase inhibitor bioactivity scores | Method validation on unbalanced data [20] |
| Bioactivity Datasets | Oneil Dataset | Contains drug combination synergy measurements | Synergistic drug discovery [17] |
| Computational Tools | ProtTrans | Protein language model for sequence feature extraction | Protein representation learning [20] |
| Computational Tools | MG-BERT | Molecular graph pre-training model for 2D structure encoding | Drug representation learning [20] |
| Computational Tools | GeoGNN | Geometric deep learning for 3D molecular structure processing | 3D drug representation [20] |
| Computational Tools | ROCS (Rapid Overlay of Chemical Structures) | 3D shape-based similarity screening | Combinatorial library screening [23] |
| Experimental Platforms | High-throughput screening systems | Automated experimental validation of selected compounds | All experimental applications |
| Cellular Model Systems | GDSC (Genomics of Drug Sensitivity in Cancer) | Provides gene expression profiles for cellular context | Synergy prediction in specific environments [17] |
Uncertainty sampling strategies represent powerful approaches for optimizing compound selection in drug discovery, offering significant efficiency improvements over traditional screening methods. The comparative analysis presented in this guide demonstrates that while standard uncertainty sampling provides a solid foundation, specialized approaches such as evidential deep learning, category-enhanced sampling, and enhanced Thompson sampling address specific limitations and application scenarios.
The experimental data consistently shows that well-implemented uncertainty sampling can achieve 60-90% of potential discoveries while evaluating only 10% or less of total available compounds [20] [17]. This efficiency gain translates directly to reduced research costs and accelerated timelines, making uncertainty sampling an increasingly essential component of modern drug discovery pipelines.
Future developments in uncertainty sampling will likely focus on improved uncertainty calibration, better integration of multi-modal data sources, and enhanced handling of extreme class imbalance scenarios. As these methods continue to evolve, their integration with emerging technologies such as hybrid AI-quantum computing approaches [24] promises to further expand their capabilities and applications in pharmaceutical research.
The concept of chemical space, a theoretical framework for organizing molecular diversity, is foundational to modern cheminformatics and drug discovery. With an estimated >10^60 potential drug-like molecules, the chemical universe is too vast to explore exhaustively [25]. This reality makes the strategic selection of diverse molecular subsets a critical task for discovering novel bioactive compounds and functional materials. Diversity-based selection aims to ensure that screened compounds are not just numerous but also broadly representative of unexplored chemical territories, thereby maximizing the probability of identifying new hits with unique properties and mechanisms of action.
In hit discovery research, diversity-based strategies are often evaluated alongside other active learning (AL) approaches, which iteratively select compounds for screening based on model predictions. The core thesis of this guide is that while all AL strategies offer efficiency gains over random screening, diversity-based methods provide a unique and essential advantage by systematically promoting exploration over exploitation. This ensures that molecular libraries do not merely grow in size but expand meaningfully in their coverage of chemical space, a distinction highlighted by recent studies questioning whether the rapid increase in database size directly translates to increased diversity [26].
This guide provides a comparative analysis of diversity-based selection against other prominent active learning strategies, supported by quantitative performance data and detailed experimental protocols.
Comparative Analysis of Active Learning Strategies
Active learning strategies for drug screening aim to optimize the experimental selection process to achieve one or both of two primary objectives: improving the performance of drug response prediction models and efficiently identifying effective treatments (hits) [27]. The table below summarizes the core operational principles of key strategies.
Table 1: Key Active Learning Strategies for Hit Discovery
| Strategy | Primary Selection Principle | Main Advantage | Typical Use Case |
|---|---|---|---|
| Diversity-Based | Selects compounds that are most dissimilar to previously tested molecules or to each other [25]. | Maximizes exploration and broad coverage of chemical space. | Early-stage discovery when little is known about the structure-activity relationship. |
| Uncertainty Sampling | Selects compounds for which the prediction model is most uncertain [27]. | Rapidly improves model accuracy in local regions around decision boundaries. | When a preliminary model exists and needs refinement. |
| Greedy Sampling | Selects compounds predicted to be most active (e.g., highest predicted IC50) [27]. | Directly maximizes the short-term yield of confirmed hits. | Hit confirmation stages after initial active regions are identified. |
| Hybrid (e.g., Uncertainty + Diversity) | Combines multiple criteria, such as uncertainty and diversity, in the selection process [27]. | Balances exploration (diversity) and exploitation (uncertainty/greedy). | A robust default choice for balanced campaign performance. |
| Random Sampling | Selects compounds randomly from the library. | Provides an unbiased baseline; simple to implement. | Baseline for comparing the performance of other strategies. |
Quantitative comparisons from a comprehensive investigation of anti-cancer drug screening reveal the relative performance of these strategies. The study evaluated approaches based on two key metrics: the number of identified hits (responses validated to be responsive) and the performance of the drug response prediction model built on the acquired data [27].
Table 2: Performance Comparison of Active Learning Strategies in Anti-Cancer Drug Screening [27]
| Strategy | Hit Identification (Relative to Random) | Model Performance | Overall Efficacy |
|---|---|---|---|
| Diversity-Based | Significant Improvement | Good, improves for some drugs | High for broad exploration |
| Uncertainty Sampling | Significant Improvement | Good, improves for some drugs | High for model refinement |
| Greedy Sampling | Moderate Improvement | Limited improvement | Medium, risks early convergence |
| Hybrid Approaches | Significant Improvement | Good, more consistent improvement | Very High, balanced performance |
| Random Sampling | (Baseline) | (Baseline) | Low |
A key real-world application is the ChemScreener workflow, which employs a Balanced-Ranking acquisition strategy. This multi-task active learning approach leverages ensemble uncertainty to explore novel chemistry while maintaining hit rate enrichment. In an iterative screen targeting the WDR5 protein, ChemScreener achieved an average hit rate of 5.91% (with cycles reaching 3–10%), a substantial increase from the primary HTS screen baseline of 0.49%. This demonstrates the power of combining exploration with targeted activity prediction, leading to the identification of multiple novel scaffold series [12].
Essential Tools and Metrics for Diversity Assessment
The Scientist's Toolkit: Research Reagent Solutions
Implementing a diversity-based selection strategy requires a suite of computational tools and metrics. The following table details the essential components of the research toolkit.
Table 3: Essential Research Reagents and Tools for Diversity Analysis
| Tool / Descriptor | Type | Primary Function | Relevance to Diversity |
|---|---|---|---|
| Molecular Fingerprints (e.g., ECFP, MACCS) [26] [25] | Structural Descriptor | Encodes molecular structure as a binary bit string. | Serves as the foundational representation for calculating structural similarity and diversity. |
| Tanimoto Coefficient [25] | Similarity Metric | Calculates the similarity between two fingerprint vectors. | The most common metric for pairwise similarity; 1 - Tanimoto is used as a distance/dissimilarity measure. |
| iSIM (Intrinsic Similarity Method) [26] | Computational Framework | Efficiently calculates the average pairwise similarity within a massive library in O(N) time. | Provides a global, single-value metric (iT) for a library's internal diversity, enabling comparison of entire databases. |
| Graph Neural Networks (GNNs) [25] | Machine Learning Model | Learns vector representations of molecules that capture both structural and property information. | Generates rich molecular descriptors that can be used for property-aware diversity selection. |
| BitBIRCH Algorithm [26] | Clustering Algorithm | Efficiently groups extremely large numbers of molecular fingerprints. | Enables "granular" analysis of chemical space by identifying natural clusters within a library. |
| Submodular Functions (e.g., Log-Determinant) [25] | Mathematical Framework | Quantifies the diversity of a set of molecules (e.g., as the volume spanned by their vectors). | Allows for efficient, near-optimal diverse subset selection with mathematical performance guarantees. |
Quantifying Diversity and Strategy Performance
The performance of hit discovery strategies is measured using a range of metrics, tailored to the specific goal, whether it is classification or regression.
- For Hit Identification (Classification):
- Hit Rate: The percentage of screened compounds that are confirmed as active. This is a direct measure of screening efficiency [12].
- Scaffold Diversity: The number of distinct molecular frameworks or core structures identified among the hits. A higher count indicates broader exploration success [12].
- For Model Performance (Regression/Prediction):
- Root Mean Squared Error (RMSE): Measures the average difference between predicted and actual activity values. It is in the same units as the target variable, making it interpretable [28].
- R-Squared (R²): Represents the proportion of variance in the response variable that is explained by the model. A higher value indicates a better fit [28].
Experimental Protocols for Strategy Evaluation
Protocol 1: Time-Evolution Analysis of Chemical Library Diversity
This protocol, derived from recent research, assesses how the chemical diversity of public libraries has evolved over time, independent of a specific screening campaign [26].
- Data Collection: Obtain sequential historical releases of a chemical database (e.g., ChEMBL releases 1 through 33).
- Molecular Representation: Standardize molecular structures and encode them using one or more fingerprint types (e.g., ECFP4).
- Global Diversity Calculation: For each library release, calculate the iT (iSIM Tanimoto) value. This metric represents the average pairwise Tanimoto similarity within the library, where a lower iT indicates greater diversity [26].
- Granular Cluster Analysis: Apply the BitBIRCH clustering algorithm to each release to identify and track the formation and growth of distinct molecular clusters over time [26].
- Complementary Similarity Analysis: For each molecule in a release, calculate its complementary similarity (the iT of the library after its removal). This identifies "medoid" molecules (low complementary similarity, central to the library) and "outlier" molecules (high complementary similarity, on the periphery) [26].
- Trend Interpretation: Analyze trends in iT, cluster count and size, and the diversity of medoid vs. outlier regions to determine if library growth corresponds with true diversity expansion.
Protocol 2: Benchmarking Active Learning Strategies for a Specific Target
This protocol outlines a head-to-head comparison of active learning strategies for a specific drug screening project.
- Initialization:
- Define Compound Library: Select an ultra-large library (e.g., >10^7 compounds) for screening [26].
- Establish Training Data: Start with a small, initial set of compounds with known activity against the target (e.g., WDR5).
- Iterative Active Learning Cycle: Repeat for a predetermined number of cycles or until a performance plateau is reached.
- Model Training: Train a predictive model (e.g., a Graph Neural Network) on all data acquired so far [25].
- Compound Selection: Apply each AL strategy in parallel to select a batch of compounds from the unexplored library.
- Diversity: Use a method like SubMo-GNN, which employs a submodular function (e.g., log-determinant) on GNN-generated molecular vectors to select a maximally diverse subset [25].
- Uncertainty: Select compounds where the ensemble of models shows the highest prediction variance [27].
- Greedy: Select compounds with the highest predicted activity.
- Hybrid: Use a balanced ranking that combines criteria (e.g.,
α * Uncertainty_Score + (1-α) * Diversity_Score).
- Experimental Testing: Acquire and test the selected compounds in a robust assay (e.g., HTRF, DSF).
- Data Integration: Add the new experimental results to the training data.
- Performance Evaluation: After the final cycle, compare the strategies based on the cumulative number of hits identified, the diversity of the hit scaffolds, and the predictive performance (e.g., RMSE) of the models they built.
The following workflow diagram illustrates the key steps and decision points in Protocol 2.
Visualizing the Chemical Space Exploration Logic
The strategic rationale for employing diversity-based selection, especially in contrast to other methods, can be visualized as a decision-making logic for navigating chemical space. The following diagram maps this high-level logic, showing how diversity-based methods prioritize broad exploration to mitigate the risk of overlooking promising regions.
The empirical data clearly demonstrates that while all advanced active learning strategies significantly outperform random and greedy sampling, diversity-based selection holds a unique and critical role in the hit discovery pipeline. Its primary strength lies in systematically ensuring the broad exploration of chemical space, which is a fundamental safeguard against the premature convergence on local optima—a common pitfall of purely exploitation-driven strategies.
For researchers and drug development professionals, the strategic implication is that diversity-based methods are indispensable in the early stages of discovery when the goal is to map the structure-activity landscape and identify novel chemotypes. As campaigns progress, hybrid approaches that balance diversity with model uncertainty or predicted activity often provide the most robust performance, efficiently expanding the chemical frontier while deepening the understanding of promising regions [12] [27]. The ongoing development of sophisticated tools like iSIM, BitBIRCH, and SubMo-GNN provides the computational rigor needed to move beyond simple compound counting and towards a truly strategic, diversity-driven expansion of the explored chemical universe [26] [25].
In the resource-intensive landscape of modern drug discovery, the strategic selection of which experiments to perform is as critical as the experiments themselves. Active learning (AL), a subfield of machine learning, has emerged as a powerful paradigm for optimizing this process by dynamically balancing exploration of the vast chemical space with exploitation of known promising regions. This balanced approach is particularly valuable in hit discovery research, where the high cost of acquiring labeled data through experimental synthesis and characterization creates significant bottlenecks [29]. Traditional discovery methods are expensive, time-consuming, and frequently have a high failure rate, often due to the lack of effective predictive models for identifying suitable drug candidates [30].
Hybrid active learning strategies, which integrate multiple query principles, are reshaping computational drug discovery pipelines. These strategies enable researchers to maximize the informational value of each experimental cycle, thereby accelerating the identification of viable therapeutic compounds. By leveraging adaptive, integrated workflows that connect functional and mechanistic insights, these approaches enhance efficacy in developing novel immune therapeutics and overcoming resistance [31]. This guide provides a comparative analysis of active learning strategies, offering experimental data and methodologies to inform their application in hit discovery research.
Experimental Protocols: Benchmarking Active Learning Strategies
A recent comprehensive benchmark study evaluated 17 active learning strategies within an Automated Machine Learning (AutoML) framework for materials science regression tasks, which share common challenges with drug discovery, such as high data acquisition costs [29]. The following details the core methodology, which can be adapted for drug-target interaction studies.
Methodological Framework
The study employed a pool-based active learning framework in a regression task scenario. The process is iterative and designed to simulate a real-world experimental cycle [29]:
- Initialization: The process begins with a small set of labeled samples ( L{\text{initial}} = { (Xi, Yi) }{i=1}^{l} ) and a large pool of unlabeled samples ( U ). The initial labeled set is typically created by random sampling from the unlabeled dataset.
- Iterative Active Learning Cycle:
- Model Training: An AutoML model is fitted on the current labeled dataset ( L ). The AutoML system automatically searches and optimizes across different model families (e.g., tree-based ensembles, neural networks) and their hyperparameters.
- Query Strategy: An AL strategy selects the most informative sample ( x^* ) from the unlabeled pool ( U ). The strategy is based on principles such as uncertainty, diversity, or expected model change.
- Annotation (Oracle): The selected sample ( x^* ) is passed to an "oracle" (e.g., a human expert, a high-fidelity simulation, or a wet-lab experiment) to obtain its target value ( y^* ). In a drug discovery context, this could involve synthesizing a compound and testing its binding affinity.
- Dataset Update: The newly labeled sample ( (x^, y^) ) is added to the training set: ( L = L \cup { (x^, y^) } ), and removed from the unlabeled pool: ( U = U \setminus { x^* } ).
- Stopping Criterion: The cycle repeats until a predefined budget (e.g., number of compounds tested) is exhausted or a performance threshold is met.
The benchmark used a train-test split of 80:20, with model validation performed automatically within the AutoML workflow using 5-fold cross-validation [29].
Active Learning Strategies Explained
The benchmark evaluated strategies based on four core principles, which can be hybridized [29]:
- Uncertainty Estimation: This strategy selects samples for which the model's prediction is most uncertain. In regression, methods like Monte Carlo Dropout are used to estimate predictive variance as a proxy for uncertainty. It is primarily an exploitation strategy, focusing on refining the model in ambiguous regions of the feature space.
- Diversity: This strategy aims to select a set of samples that are representative of the overall distribution of the unlabeled pool. It is an exploration strategy, ensuring broad coverage of the chemical space and preventing the model from overlooking promising, unexplored regions.
- Expected Model Change Maximization (EMCM): This approach selects the sample that would cause the greatest change in the current model parameters if its label were known. It seeks data points with high potential information gain.
- Representativeness: This principle ensures that the selected samples are not only informative but also representative of the underlying data distribution, avoiding the selection of outliers.
Hybrid strategies, such as RD-GS, combine uncertainty and diversity criteria to balance exploration and exploitation. The benchmark found that these hybrid approaches, along with pure uncertainty-based methods like LCMD and Tree-based-R, were particularly effective in the early, data-scarce phases of learning [29].
Performance Comparison of Active Learning Strategies
The following tables summarize the quantitative performance of different active learning strategies and computational approaches as reported in benchmark studies and recent literature.
Table 1: Performance of Active Learning Strategies in a Materials Science Benchmark (applicable to drug discovery) [29]
| Strategy Type | Example Methods | Early-Stage Performance (Data-Scarce) | Late-Stage Performance (Data-Rich) | Key Characteristics |
|---|---|---|---|---|
| Uncertainty-Based | LCMD, Tree-based-R | Clearly outperforms random sampling | Converges with other methods | Excellent for exploitation; targets model decision boundaries. |
| Diversity-Hybrid | RD-GS | Clearly outperforms random sampling | Converges with other methods | Balances exploration and exploitation; selects representative and informative samples. |
| Geometry-Only | GSx, EGAL | Underperforms compared to uncertainty/hybrid | Converges with other methods | Focuses on exploration and data distribution coverage. |
| Random Sampling | (Baseline) | (Baseline for comparison) | (Baseline for comparison) | Passive, non-strategic approach. |
Table 2: Performance of AI/Hybrid Models in Drug-Target Interaction and Hit Discovery
| Model/Approach | Reported Accuracy/Performance | Key Application & Findings | Reference |
|---|---|---|---|
| Context-Aware Hybrid (CA-HACO-LF) | Accuracy: 0.986 (98.6%) on drug-target interaction prediction. | Proposed for drug discovery; combines ant colony optimization for feature selection with logistic forest classification. | [30] |
| Quantum-Enhanced AI (Insilico Medicine) | 21.5% improvement in filtering non-viable molecules vs. AI-only. | Screened 100M molecules for KRAS-G12D target; identified a compound with 1.4 μM binding affinity. | [24] |
| Generative AI (GALILEO) | 100% in vitro hit rate (12/12 compounds active). | Generated novel antiviral compounds targeting viral RNA polymerases from a 52 trillion molecule space. | [24] |
| FP-GNN Model | Accuracy: 0.91 in determining compound effectiveness. | Used to identify multifunctional antimicrobial compounds. | [30] |
Workflow Visualization: Active Learning in Hit Discovery
The following diagram illustrates the iterative cycle of a pool-based active learning workflow, as implemented in the benchmark study and adapted for a drug discovery context.
The Scientist's Toolkit: Essential Research Reagents & Solutions
The implementation of the described experimental protocols relies on several key computational and experimental resources.
Table 3: Key Research Reagent Solutions for AI-Driven Hit Discovery
| Reagent / Solution | Function in the Workflow | Application Context |
|---|---|---|
| AutoML Platforms | Automates the selection and optimization of machine learning models and their hyperparameters, reducing manual tuning effort. | Essential for the model training step within the active learning cycle, especially when the underlying model may change between iterations [29]. |
| Kaggle 11,000 Medicine Dataset | Provides a structured dataset of drug details for training and benchmarking predictive models for drug-target interactions. | Used as a benchmark dataset for pre-processing, feature extraction (e.g., N-grams, Cosine Similarity), and model validation [30]. |
| High-Content Imaging & Single-Cell Transcriptomics | Provides high-dimensional, phenotypic data for analysis. Enables nuanced biological response measurement in integrated screening approaches. | Critical in phenotypic screening and for validating the functional outcomes of predictions in complex cellular systems [31]. |
| Multi-omics Datasets (Genomics, Proteomics) | Provides a comprehensive framework for linking observed phenotypic outcomes to discrete molecular pathways. | Used to inform target identification and hypothesis refinement in hybrid discovery workflows [31]. |
| Python-based ML Stack (e.g., Scikit-learn, PyTorch) | Provides the programming environment for implementing feature extraction, similarity measurement, and classification models. | The standard open-source ecosystem for building and deploying custom active learning pipelines and AI models like CA-HACO-LF [30]. |
Discussion and Strategic Outlook
The empirical data demonstrates that the choice of active learning strategy is not one-size-fits-all but is highly dependent on the stage of the discovery campaign and the available data budget. In the critical early stages, uncertainty-driven and diversity-hybrid strategies provide a significant performance advantage over passive or purely exploratory methods by maximizing the information gain from each expensive experimental cycle [29]. As the labeled set grows, the marginal benefit of sophisticated AL strategies diminishes, and all methods tend to converge.
The future of hit discovery lies in the continued integration of these balanced AL strategies with even more powerful AI paradigms. The emergence of hybrid AI and quantum computing approaches in 2025 indicates a new era of computational capability, enabling the exploration of chemical spaces at an unprecedented scale and precision [24]. Furthermore, the parallel trend of combining phenotypic and target-based discovery creates a feedback loop where AL can guide exploration in both functional and mechanistic dimensions, accelerating the development of novel therapeutics such as immune checkpoint inhibitors and bispecific antibodies [31]. For researchers, adopting these hybrid, balanced strategies is becoming essential for maintaining a competitive edge in the increasingly complex and data-driven field of drug discovery.
Efficient hit discovery for challenging protein targets like WD Repeat Domain 5 (WDR5) represents a critical bottleneck in early drug discovery. WDR5 is a highly conserved nuclear protein that functions as a molecular scaffold, playing a central role in numerous biological processes through its function in mediating the assembly of large protein complexes. It regulates chromatin modification and gene expression, including the recruitment of c-MYC to chromatin—a key process implicated in the pathogenesis of c-MYC-dependent cancers such as acute myeloid leukemia [32]. Two distinct binding sites have been identified: the WDR5-interacting (WIN) site, which engages with SET-family methyltransferases, and the WDR5-binding motif (WBM) site, responsible for c-MYC recruitment [32].
The chemical space of drug-like compounds is estimated to include around 10^60 possible molecules, making efficient navigation of this vast diversity one of the biggest challenges in drug discovery [32]. This case study examines how ChemScreener's active learning workflow addressed this challenge for WDR5 inhibitor discovery and compares its performance against alternative computational and experimental approaches.
Methodology: ChemScreener's Active Learning Workflow
ChemScreener employs a multi-task active learning workflow specifically designed for early drug discovery across large, diverse chemical libraries. The workflow operates iteratively through a structured process that combines ensemble modeling with strategic compound selection [12] [33].
Experimental Protocol
The core methodology consists of five key phases that create a continuous learning loop:
Ensemble Model Training: Ten independent ChemProp models are trained on all available labeled compound data, providing robust uncertainty estimates through consensus prediction [33].
Compound Scoring & Selection: New compounds are evaluated using a Balanced-Ranking acquisition strategy that leverages ensemble uncertainty to explore novel chemistry while maintaining hit rate enrichment by prioritizing predicted activity [12].
Domain-Specific Filtering: Selected candidates undergo application of domain-specific filters and enforcement of drug-likeness criteria to ensure compound quality and relevance [33].
Experimental Testing: Filtered compounds proceed to experimental validation using standardized biological assays—in this case, HTRF (Homogeneous Time-Resolved Fluorescence) screening for WDR5 inhibition [12].
Data Integration: New experimental results are merged into the training set, and the process repeats with retrained models [33].
For the WDR5 case study, this workflow was implemented across five iterative single-dose HTRF screens, with hit consolidation, retesting of close analogs, and validation through dose-response curves and counter-screening in secondary assays [12].
WDR5 Biological Context and Experimental Design
WDR5 features a distinctive 7-bladed β-propeller architecture with its primary functional sites being the WIN site, which contains a characteristic arginine-binding cavity, and the WBM site on the protein surface [32] [34]. The WIN site mediates interactions with SET1 family methyltransferases, while the WBM site facilitates interactions with MYC and RbBP5 [34]. ChemScreener targeted the WIN site for inhibitor discovery, employing HTRF assays for primary screening followed by surface plasmon resonance (SPR), differential scanning fluorimetry (DSF), and NanoBRET target engagement assays for validation [12] [32].
Performance Comparison: ChemScreener vs Alternative Approaches
Quantitative Performance Metrics
The table below summarizes the experimental outcomes for ChemScreener and two alternative computational approaches for WDR5 inhibitor discovery.
| Approach | Key Features | Screening Efficiency | Hit Rate | Chemical Diversity | Key Outcomes |
|---|---|---|---|---|---|
| ChemScreener (Active Learning) | Ensemble-based active learning; Balanced-Ranking acquisition [12] | 1,760 compounds tested over 5 iterative cycles [12] | 5.91% average (104 hits); 3-10% range [12] | 3 novel scaffold series + 3 singleton scaffolds [12] | 44 compounds advanced to dose-response; over 50% validated as binders by DSF [12] |
| DEL-ML (DNA-Encoded Library + Machine Learning) | DEL screening integrated with machine learning; de novo compound generation [32] | Rapid progression from screening to optimized probe (LH168) [32] | Initial hit MR43378 with nanomolar potency [32] | Focused optimization from single chemical series [32] | LH168 probe: 10 nM EC50 in cells, exceptional selectivity, long residence time [32] |
| Generative AI (Insilico Medicine) | AI-powered molecule generation; physics-based molecular modeling [35] | 60-200 molecules synthesized and tested per program [35] | Lead compound 9c-1 with 35-fold improvement over initial hit [35] | Novel scaffolds generated de novo [35] | Sub-micromolar binding affinity for WDR5-MYC PPI inhibitors [35] |
| Traditional HTS (Reference) | Experimental screening without computational prioritization | Primary screen of large compound library | 0.49% hit rate [12] | Limited by library diversity | Provides baseline for comparison |
Experimental Workflow Comparison
The diagram below illustrates the fundamental differences in methodology between the three computational approaches for WDR5 inhibitor discovery.
The Scientist's Toolkit: Essential Research Reagents and Methods
Successful implementation of WDR5 inhibitor discovery programs requires specific experimental tools and methodologies. The table below details key research reagents and their applications in the evaluated studies.
| Research Tool | Type | Key Applications in WDR5 Studies | Example Implementation |
|---|---|---|---|
| HTRF Assays | Biochemical high-throughput screening | Primary single-dose screening for WIN site inhibitors [12] | 5 iterative screens with dose-response confirmation [12] |
| Surface Plasmon Resonance (SPR) | Biophysical binding kinetics | Determination of binding affinity (KD) and residence time [32] | Characterization of LH168 (KD = 154 nM) and residence time (714s) [32] |
| NanoBRET Target Engagement | Cellular potency assessment | Measurement of cellular target engagement (EC50) [32] | Intact vs. permeabilized cells to assess membrane penetration [32] |
| Differential Scanning Fluorimetry (DSF) | Thermal stability binding assessment | Validation of direct binding to WDR5 protein [12] | Counter-screening to confirm >50% of hits as true binders [12] |
| X-ray Crystallography | Structural biology | Elucidation of binding modes and structure-activity relationships [32] | Co-crystal structure of WDR5 with inhibitors (e.g., PDB ID: 8T5I) [32] |
| Biolayer Interferometry (BLI) | Kinetic binding characterization | Quantification of protein-protein interaction inhibition [34] | Measurement of WDR5 interactions with MYC and RbBP5 peptides [34] |
Discussion: Strategic Implications for Hit Discovery
The comparative analysis reveals distinct strategic advantages for each approach, highlighting how selection should be guided by specific research objectives and resource constraints.
Context-Dependent Approach Selection
ChemScreener's active learning demonstrates particular strength in exploration-focused tasks where the goal is identifying diverse chemotypes from large chemical libraries. Its balanced approach to exploration and exploitation yielded a 12-fold improvement in hit rate over traditional HTS while discovering multiple novel scaffolds [12] [33]. This makes it ideally suited for early-stage projects where chemical starting points are limited or when seeking intellectual property around novel chemotypes.
DEL-ML integration excels in rapid probe development from validated starting points, as demonstrated by the streamlined optimization of MR43378 to the highly selective chemical probe LH168 [32]. The approach leverages the massive screening capacity of DEL technology (millions to billions of compounds) while using machine learning to extrapolate to commercially available chemical space [32]. This approach is particularly valuable for targets with established binding sites but limited chemical matter.
Generative AI platforms show exceptional capability for de novo design of novel chemotypes, especially for challenging targets like the WDR5-MYC protein-protein interaction [35]. The technology's ability to generate molecules with desired properties from scratch rather than selecting from existing libraries represents a paradigm shift, particularly for undruggable targets where conventional approaches have failed.
Experimental Design Considerations
The WDR5 case studies emphasize that computational hit identification requires rigorous experimental validation across multiple orthogonal assays. The most successful implementations combine computational prediction with:
- Dose-response confirmation in primary assays
- Counter-screens to eliminate false positives
- Cellular target engagement assessment
- Binding kinetics characterization
- Structural validation where possible
This multi-faceted validation approach ensured that computational hits translated to biologically relevant inhibitors, with all three approaches delivering chemically tractable, experimentally validated starting points for WDR5 drug discovery.
The ChemScreener case study for WDR5 inhibitor discovery demonstrates that active learning strategies can dramatically improve the efficiency and outcomes of early hit discovery. Compared to traditional HTS, ChemScreener achieved a 12-fold increase in hit rate while identifying multiple novel scaffold series [12]. When evaluated alongside alternative computational approaches, each method exhibits distinct strengths: DEL-ML for rapid probe development from validated hits [32], generative AI for de novo design of novel chemotypes [35], and active learning for optimal exploration of large chemical spaces [12] [33].
The selection of an appropriate hit discovery strategy should be guided by project-specific goals, available chemical starting points, and the nature of the target biology. For research teams seeking to maximize chemical diversity from large screening libraries while maintaining strong enrichment rates, ChemScreener's active learning workflow represents a compelling approach that successfully balances exploration of novel chemistry with exploitation of predicted activity.
The drug discovery process is notoriously protracted, expensive, and prone to failure, traditionally relying on the experimental screening of vast chemical libraries. The emergence of generative artificial intelligence (AI) has catalyzed a paradigm shift, enabling the computational de novo design of novel molecular structures from scratch tailored to specific properties [36] [37]. However, generative models often face significant challenges, including insufficient target engagement, poor synthetic accessibility (SA) of proposed molecules, and limited generalization beyond their training data [11]. To overcome these limitations, researchers are increasingly integrating generative AI with active learning (AL), an iterative machine learning feedback process that intelligently selects the most informative data points for labeling and model refinement [13] [38]. This powerful synergy creates a self-improving cycle where generative models propose novel candidate molecules, and AL strategies guide experimental or computational validation toward the most promising regions of chemical space. This case study provides a comparative analysis of this integrated framework, evaluating its performance against conventional methods and other AI-driven approaches within the context of hit discovery research.
Comparative Analysis of Integrated Frameworks vs. Alternative Approaches
The integration of Generative AI with Active Learning can be conceptualized as a unified framework. However, its implementation varies, leading to distinct performance outcomes. The following analysis compares a representative integrated framework against other common computational drug discovery methods.
Table 1: Performance Comparison of De Novo Drug Design Frameworks
| Framework / Model | Core Approach | Reported Efficacy (Hit Rate) | Novelty & Diversity | Synthetic Accessibility (SA) | Key Advantages | Primary Limitations |
|---|---|---|---|---|---|---|
| VAE + Nested AL (Physics-Based) [11] | Variational Autoencoder with nested AL cycles using chemoinformatic & physics-based oracles. | High (CDK2): 8/9 synthesized molecules showed in vitro activity (≈89% hit rate); 1 nanomolar potency [11]. | High (novel scaffolds generated for CDK2 & KRAS) [11]. | Explicitly optimized via SAscore filter [11]. | High-fidelity, experimentally validated; excels in low-data regimes; balances exploration & exploitation [11]. | Computationally intensive due to physics-based simulations [11]. |
| DRAGONFLY (Interactome Learning) [39] | Graph Transformer & LSTM network trained on a drug-target interactome. | Prospective validation: Potent PPARγ partial agonists with desired selectivity identified [39]. | High (quantitative scaffold & structural novelty) [39]. | Optimized via Retrosynthetic Accessibility Score (RAScore) [39]. | "Zero-shot" learning requires no target-specific fine-tuning; integrates ligand- and structure-based design [39]. | Performance plateaued with USRCAT descriptors beyond ~100 training molecules [39]. |
| Fine-Tuned RNN (Chemical Language Model) [39] | Recurrent Neural Network (RNN) fine-tuned on target-specific data. | Lower than DRAGONFLY in comparative studies [39]. | Lower than DRAGONFLY in comparative studies [39]. | Lower than DRAGONFLY in comparative studies [39]. | Simpler architecture; well-established for sequence-based generation [40]. | Requires target-specific fine-tuning; struggles with single-template learning [39]. |
| Conventional Virtual Screening | Selecting compounds from pre-existing static libraries (e.g., via docking). | Variable; often lower than AL-guided approaches [13] [38]. | Limited to existing chemical space of the library. | Dependent on library composition. | Fast; easy to implement. | Limited exploration; does not generate novel chemotypes [11]. |
Experimental Protocol: The VAE-Nested AL Workflow
The following section details the methodology behind one of the most robust integrated frameworks, which combines a Variational Autoencoder (VAE) with a physics-based Active Learning framework, as validated on CDK2 and KRAS targets [11].
The experimental pipeline is a structured, iterative process designed to continuously generate and refine molecules. The diagram below illustrates the core workflow and logical relationships of this nested AL strategy.
Detailed Methodological Steps
Data Representation and Initial Training:
- Molecular Representation: Training molecules are represented as SMILES strings, which are tokenized and converted into one-hot encoding vectors for model input [11].
- Model Pre-training: The VAE is first trained on a large, general dataset of drug-like molecules to learn the fundamental rules of chemical structure. It is then initially fine-tuned on a target-specific training set to imbue it with basic knowledge of relevant chemotypes [11].
Nested Active Learning Cycles:
- Inner AL Cycle (Guided by Chemoinformatic Oracles): The trained VAE samples the latent space to generate new molecules. These are filtered through a chemoinformatic oracle that evaluates:
- Drug-likeness: Adherence to rules like Lipinski's Rule of Five [37].
- Synthetic Accessibility (SA): Estimated using metrics like SAscore [11] [40].
- Structural Novelty: Assessed by dissimilarity to molecules already in the training set. Molecules passing these filters are added to a "temporal-specific set," which is used to fine-tune the VAE, steering subsequent generation toward more drug-like and synthesizable structures [11].
- Outer AL Cycle (Guided by Physics-Based Oracles): After a set number of inner cycles, molecules accumulated in the temporal set are evaluated by a physics-based oracle. This typically involves:
- Molecular Docking: Simulations to predict the binding pose and affinity of the generated molecules against the target protein's structure. Molecules achieving favorable docking scores are promoted to a "permanent-specific set," which is used for the next round of VAE fine-tuning, directly optimizing for target engagement [11].
- Inner AL Cycle (Guided by Chemoinformatic Oracles): The trained VAE samples the latent space to generate new molecules. These are filtered through a chemoinformatic oracle that evaluates:
Candidate Selection and Experimental Validation:
- Stringent Filtration: Post-AL cycles, the most promising candidates from the permanent set undergo rigorous selection.
- Advanced Molecular Modeling (MM): Techniques like Monte Carlo simulations with Protein Energy Landscape Exploration (PELE) provide a deeper analysis of binding interactions and stability [11].
- Absolute Binding Free Energy (ABFE) Calculations: Used for high-fidelity affinity prediction to prioritize the final candidates for synthesis [11].
- Synthesis and Bioassay: The top-ranking molecules are chemically synthesized and tested in vitro for activity (e.g., IC₅₀ determination) and selectivity, providing experimental validation of the framework's efficacy [11].
The Scientist's Toolkit: Essential Research Reagents and Solutions
The successful implementation of an integrated Generative AI and AL pipeline relies on a suite of computational tools and reagents. The following table details key components referenced in the featured experiments.
Table 2: Key Research Reagents and Computational Solutions
| Category | Item / Software | Primary Function in the Workflow |
|---|---|---|
| Generative Models | Variational Autoencoder (VAE) [11] | Encodes molecules into a continuous latent space; enables smooth interpolation and controlled generation of novel molecular structures. |
| Chemical Language Model (CLM) [39] | Models chemical structures as sequences (e.g., SMILES) for de novo generation; often based on RNNs or Transformers. | |
| Molecular Representations | SMILES (Simplified Molecular-Input Line-Entry System) [11] [41] | A string-based notation for representing molecular structures as text for AI model input. |
| Molecular Graph [41] [39] | Represents atoms as nodes and bonds as edges, preserving molecular topology for graph neural networks. | |
| SELFIES [41] | A robust molecular representation that guarantees 100% valid chemical structures, overcoming SMILES syntax issues. | |
| Active Learning Oracles | Chemoinformatic Oracle (e.g., SAscore) [11] [40] | Computationally predicts the ease of synthesis for a generated molecule, filtering out impractical designs. |
| Physics-Based Oracle (e.g., Molecular Docking) [11] | Predicts the binding mode and affinity of a generated molecule to a protein target, prioritizing molecules with high potential activity. | |
| Validation & Simulation | PELE (Protein Energy Landscape Exploration) [11] | An advanced simulation method used for candidate selection to model protein-ligand dynamics and binding stability. |
| Absolute Binding Free Energy (ABFE) Calculations [11] | Provides high-accuracy predictions of binding affinity for final candidate prioritization before synthesis. | |
| Databases | ChEMBL [39] | A large-scale database of bioactive molecules with drug-like properties, used for model training and validation. |
Architectural Comparison: Integrated Framework vs. Alternatives
The integrated framework's performance can be better understood by examining the fundamental architectural differences between it and other common approaches. The following diagram contrasts the closed-loop, self-improving nature of the Generative AI/AL integration with the linear process of conventional virtual screening and the single-step generation of standalone generative AI.
The empirical data and comparative analysis presented in this case study strongly indicate that the integration of Generative AI with Active Learning represents a superior strategy for de novo hit discovery compared to conventional methods or standalone generative models. The key differentiator is the creation of a closed-loop, self-improving system [41]. Unlike linear processes, this framework uses AL to inject expert knowledge and experimental feedback directly into the generative process, leading to rapid iterative improvement.
The most compelling evidence comes from prospective experimental validations, such as the application of the VAE-nested AL framework to CDK2, which achieved an exceptional hit rate of approximately 89% (8 out of 9 synthesized molecules showed in vitro activity) [11]. This success underscores the framework's ability to efficiently navigate the vast chemical space and prioritize candidates with a high probability of experimental success, thereby significantly accelerating the early stages of drug discovery. For researchers, the choice of framework depends on specific project goals: standalone generative AI or conventional screening may suffice for rapid exploration, but for demanding hit-discovery campaigns against challenging targets with limited data, the integrated Generative AI and Active Learning framework offers a powerful, evidence-backed strategy to enhance efficiency and success rates.
The SARS-CoV-2 main protease (Mpro), also known as 3CLpro, is a critical enzyme in the viral life cycle, responsible for cleaving the viral polyproteins pp1a and pp1ab into functional non-structural proteins essential for viral replication and transcription [42] [43]. Its conservation across coronaviruses and absence of closely related human homologs make it an attractive target for antiviral drug development [44] [43]. The COVID-19 pandemic triggered an unprecedented research effort to discover effective Mpro inhibitors, leading to the exploration of innovative computational approaches, including active learning (AL), to accelerate hit discovery [45].
Active learning represents a paradigm shift in virtual screening, moving beyond traditional one-shot methods to an iterative, guided approach. By strategically selecting which compounds to evaluate with computationally expensive methods, AL aims to maximize the exploration of chemical space while minimizing resources [46]. This case study objectively compares an automated AL workflow for Mpro hit discovery against traditional virtual screening approaches, evaluating their performance, experimental validation, and practical implementation.
Workflow Comparison: AL Versus Traditional Virtual Screening
The FEgrow Active Learning Workflow
The FEgrow software package implements an automated AL workflow specifically designed for structure-based hit expansion [46] [47]. This approach combines molecular growing with iterative model refinement:
- Initialization: The process begins with a protein structure (Mpro) and a known ligand core or fragment hit. FEgrow builds congeneric series by attaching flexible linkers and functional groups (R-groups) from extensive libraries [46].
- Hybrid Scoring: Compounds are scored using a hybrid approach that combines machine learning/molecular mechanics (ML/MM) potential energy functions with the gnina convolutional neural network scoring function to predict binding affinity [46].
- Active Learning Cycle: The system employs an iterative loop where a subset of compounds is evaluated using the expensive FEgrow objective function. These results train a machine learning model that predicts which compounds to evaluate next, progressively refining the search toward promising chemical space [46] [47].
- On-Demand Library Integration: The workflow can be "seeded" with purchasable compounds from on-demand chemical libraries like the Enamine REAL database, ensuring synthetic tractability of proposed hits [46].
Traditional Virtual Screening Approaches
Traditional structure-based methods typically follow a linear workflow:
- Pharmacophore Modeling: Based on known active compounds or protein structure, pharmacophore models are created to represent essential interaction features [44].
- Database Screening: Large compound libraries (often millions to billions of compounds) are screened using the pharmacophore model or molecular docking [42] [44].
- Ranking and Selection: Compounds are ranked by docking scores or similarity metrics, with top candidates selected for experimental testing without iterative refinement [42] [44].
Table 1: Key Characteristics of Mpro Screening Approaches
| Feature | Active Learning Workflow | Traditional Virtual Screening |
|---|---|---|
| Search Strategy | Iterative, guided exploration | Linear, one-shot screening |
| Chemical Space Evaluation | Progressive refinement based on previous results | Exhaustive or random sampling |
| Computational Resource Allocation | Focused on promising regions | Distributed across entire library |
| Adaptability | Improves based on accumulated data | Fixed criteria throughout process |
| Key Tools | FEgrow, gnina, custom AL algorithms [46] | Docking (AutoDock Vina, ICM-Pro), pharmacophore modeling [44] |
| Library Size Handled | Efficient with ultra-large libraries (>1B compounds) [46] | Practical for libraries up to hundreds of millions [44] |
Diagram 1: Workflow comparison between traditional and AL approaches.
Experimental Protocols & Performance Metrics
Implementation of the FEgrow AL Workflow
The FEgrow AL methodology employs specific technical implementations:
- Molecular Growing: Starting from a constrained core, FEgrow grows user-defined R-groups using a library of 2000 linkers and approximately 500 R-groups, optimizing grown conformations with ML/MM potential energy functions within a rigid protein binding pocket [46].
- Active Learning Integration: The workflow interfaces with active learning to efficiently search the combinatorial space of possible linkers and functional groups. The AL algorithm selects compounds for evaluation based on their potential to improve the model, using strategies such as uncertainty sampling or expected improvement [46].
- Objective Functions: Beyond docking scores, the workflow can optimize functions that combine molecular properties (e.g., molecular weight) and protein-ligand interaction profiles (PLIP) derived from crystallographic fragments [46].
Traditional Screening Methodology
The conventional approach for Mpro inhibitor identification typically involves:
- Structure Preparation: Multiple crystallographic structures of SARS-CoV-2 Mpro (e.g., PDB IDs: 7VLP, 7TE0, 7RFS) are prepared by removing water molecules and cofactors, followed by binding site definition based on co-crystallized ligands [44].
- Virtual Screening Protocol: Libraries of hundreds of millions of compounds are screened using tools like AutoDock Vina or ICM-Pro, with compounds ranked by docking scores. Pharmacophore constraints may be applied to filter implausible binding modes [44].
- Hit Selection: Top-ranked compounds are selected based on consensus scoring across multiple protein conformations and visual inspection of binding poses [44].
Quantitative Performance Comparison
Table 2: Experimental Performance Metrics for Mpro Inhibitor Discovery
| Metric | FEgrow AL Workflow [46] | Traditional Fingerprint Screening [42] | Advanced Virtual Screening [44] |
|---|---|---|---|
| Initial Library Size | Combinatorial space of linkers/R-groups | ~1.37 billion compounds screened | ~200 million compounds screened |
| Compounds Selected for Testing | 19 designs purchased and tested | 48 compounds tested | 43 compounds tested |
| Experimentally Confirmed Hits | 3 compounds with weak activity | 21 inhibitors (>50% inhibition at 20μM) | 2 compounds with micromolar activity |
| Hit Rate | 15.8% | 43.8% | 4.7% |
| Best IC50 Values | Weak activity (not quantified) | ~1 μM | Micromolar range |
| Key Structural Features | Similar to COVID Moonshot hits | Isoquinoline motif conserved | Novel scaffolds |
| Computational Resource Efficiency | High (targeted exploration) | Moderate (pre-similarity filtering) | Low (exhaustive docking) |
Research Reagent Solutions Toolkit
Table 3: Essential Research Tools for Mpro Inhibitor Discovery
| Tool/Resource | Type | Function in Research | Example Application |
|---|---|---|---|
| FEgrow Software [46] [47] | Computational Tool | Builds and scores congeneric series in protein binding pockets | Automated de novo design targeting SARS-CoV-2 Mpro |
| Enamine REAL Library [42] [46] | Compound Database | Provides access to billions of readily synthesizable compounds | Source of purchasable compounds for experimental validation |
| RDKit [46] | Cheminformatics Toolkit | Handles molecular operations and conformer generation | Merging molecular components in FEgrow workflow |
| gnina [46] | Deep Learning Scorer | Predicts binding affinity using convolutional neural networks | Scoring grown compounds in FEgrow workflow |
| AutoDock Vina [44] | Molecular Docking | Performs flexible ligand docking against fixed protein | Structure-based virtual screening of large libraries |
| ICM-Pro [44] | Molecular Modeling | Uses Monte Carlo simulation for global energy minimization | High-precision docking and binding site analysis |
| SARS-CoV-2 Mpro Assay [42] [43] | Biochemical Assay | Measures enzymatic inhibition of Mpro activity | Experimental validation of computational hits |
| Mpropred Web-App [48] | ML Predictor | Predicts bioactivity against SARS-CoV-2 Mpro | Rapid pre-screening of compound libraries |
Discussion: Strategic Implications for Hit Discovery
The comparative analysis reveals distinct advantages and limitations for each approach in Mpro inhibitor discovery:
The FEgrow AL workflow demonstrates superior computational efficiency for navigating ultra-large chemical spaces, with the ability to identify viable hits while evaluating only a fraction of the total combinatorial possibilities [46]. Its strength lies in the iterative refinement process, which adapts based on accumulated data to focus resources on promising chemical regions. However, the experimental hit rate of 15.8% with only weak activity suggests potential limitations in the current scoring functions or AL selection criteria for achieving high-potency inhibitors [46].
Traditional fingerprint-based screening achieved the highest hit rate (43.8%) in this comparison, successfully identifying compounds with low micromolar activity [42]. This approach benefited from starting with a high-quality lead compound from the COVID Moonshot consortium and using molecular fingerprint similarity to explore analogous structures. The conservation of the isoquinoline motif across the most potent hits validates this structure-based strategy, though it may limit chemical diversity [42].
Advanced virtual screening methods, while comprehensive, showed the lowest efficiency with a 4.7% hit rate despite screening 200 million compounds [44]. This underscores the challenge of false positives in traditional docking approaches and highlights the potential value of incorporating AL refinement cycles to improve prioritization.
Diagram 2: Strategy selection guide for Mpro inhibitor discovery.
The integration of active learning with structure-based drug design represents a promising evolution in virtual screening methodology for targeting SARS-CoV-2 Mpro. The FEgrow AL workflow demonstrates compelling advantages in computational efficiency and automation, particularly for exploring vast chemical spaces with limited initial structural data [46]. However, traditional similarity-based approaches maintain value when high-quality lead compounds are available, as evidenced by their superior hit rates in specific scenarios [42].
Future developments in AL for hit discovery will likely focus on improving scoring functions through more accurate affinity prediction, incorporating synthetic accessibility directly into the optimization process, and expanding toward multi-target inhibition strategies [49] [50]. The emerging paradigm of pan-coronavirus inhibitor design, targeting Mpro across SARS-CoV-2, SARS-CoV, and MERS-CoV, presents an ideal application for advanced AL approaches that can balance multiple optimization objectives simultaneously [50].
As these technologies mature, the integration of experimental feedback directly into the AL cycle will further close the loop between computational prediction and experimental validation, accelerating the discovery of effective antiviral therapeutics against current and emerging pathogenic threats.
Overcoming Challenges and Fine-Tuning Your Active Learning Workflow
This guide objectively compares active learning strategies for hit discovery research, focusing on their performance in managing the exploration-exploitation trade-off. It provides structured experimental data, detailed protocols, and essential resource information to help researchers select optimal strategies for their drug discovery pipelines.
In modern drug discovery, the exploration-exploitation trade-off is a fundamental strategic challenge. Researchers must balance exploring vast, uncharted chemical spaces to find novel compounds (exploration) against optimizing and refining known hit compounds to improve their properties (exploitation). Active Learning (AL), a subfield of artificial intelligence, has emerged as a powerful methodology to navigate this dilemma efficiently [13]. AL employs an iterative feedback process that selects the most valuable data points for experimental labeling based on model predictions, thereby building high-quality models or discovering desirable molecules with fewer costly experiments [13].
The core challenge AL addresses is the prohibitive cost and time associated with experimentally testing every possible compound. By using machine learning models to guide the selection process, AL helps focus resources on the most informative experiments, whether they lie in regions of high uncertainty (exploration) or high predicted performance (exploitation). This guide compares the performance of prominent AL strategies used in hit discovery, providing a structured framework for decision-making.
Comparative Analysis of Active Learning Strategies
The effectiveness of an AL strategy is highly dependent on the specific goals and constraints of a drug discovery project. The following table summarizes the core characteristics, strengths, and limitations of major strategic families.
Table 1: Comparison of Active Learning Strategic Families for Hit Discovery
| Strategy Family | Core Mechanism | Best-Suited Application in Hit Discovery | Key Advantages | Key Limitations |
|---|---|---|---|---|
| Uncertainty Sampling [13] | Queries data points where the model's prediction is least certain. | Early-stage exploration for scaffold hopping; defining initial Structure-Activity Relationships (SAR). | Simple to implement; highly effective for rapid model improvement; computationally efficient. | Can be myopic; may miss clusters of promising compounds; prone to selecting outliers. |
| Expected Improvement (EI) [51] | Selects points offering the highest expected improvement over the current best candidate. | Hit-to-Lead (H2L) optimization focused on improving a key property like potency or selectivity. | Directly targets performance gain; balances moderate exploration with performance-driven exploitation. | Performance depends on accurate model predictions; can converge prematurely to local optima. |
| Info-p / Information-Based [51] | Maximizes expected information gain about the identity of the best possible compound. | Projects where identifying the single best candidate is critical; requires high statistical confidence. | Asymptotically optimal regret bounds; theoretically grounded for optimal identification. | Computationally intensive; requires sophisticated probabilistic modeling. |
| Multi-Objective & Pareto-Front [51] | Treats exploration and exploitation as separate objectives and selects from the Pareto-optimal front. | Multi-parameter optimization (e.g., balancing potency, solubility, and metabolic stability). | Avoids arbitrary weighting of goals; reveals diverse trade-off options; robust in high dimensions. | Increased complexity in analysis and decision-making; requires defining multiple objectives. |
| Adaptive Bayesian (e.g., BHEEM) [51] | Dynamically adjusts the exploration-exploitation balance using online Bayesian updates of the trade-off parameter. | Dynamic projects where the optimal balance shifts (e.g., from broad screening to focused optimization). | Data-driven adaptation; robust to changing project needs; eliminates need for static parameters. | Implementation complexity; requires expertise in Bayesian modeling and computation. |
Quantitative Performance Benchmarking
The theoretical strengths of these strategies are validated through empirical performance metrics. The following table synthesizes key quantitative findings from computational and experimental studies, providing a basis for comparison.
Table 2: Experimental Performance Metrics of Active Learning Strategies
| Strategy | Reported Performance Metric | Comparative Result | Experimental Context |
|---|---|---|---|
| Uncertainty Sampling [13] | Hit identification efficiency | Identifies ~80% of active compounds by testing only 20% of the library [13]. | Virtual screening for compound-target interaction prediction. |
| Info-p Algorithm [51] | Asymptotic regret | Matches the Lai-Robbins lower bound for asymptotic regret, indicating optimal long-term performance [51]. | Multi-armed bandit simulations for best-arm identification. |
| Pareto-Front Methods [51] | Model accuracy (RMSE) | Achieves 21% lower RMSE than pure exploration and 11% better than pure exploitation in regression tasks [51]. | Dynamic regression and active learning for reliability analysis. |
| Bayesian Hierarchical (BHEEM) [51] | Model accuracy (RMSE) | 21% lower RMSE than pure exploration; 11% better than pure exploitation [51]. | Regression tasks with adaptive trade-off control. |
| Active Learning (General) [52] | Knowledge retention | 93.5% retention for active learners vs. 79% for passive learners [52]. | Corporate safety training studies. |
| Active Learning (General) [52] | Test scores / Performance | 54% higher test scores; 70% average score vs. 45% with passive learning [52]. | Educational research across multiple disciplines. |
Experimental Protocols for Strategy Evaluation
To ensure reproducibility and provide a clear methodology for benchmarking AL strategies, the following detailed experimental protocol is provided. This workflow is adapted from standard practices in chemoinformatics and computational drug discovery [13] [53].
Protocol: Benchmarking AL Strategies for Virtual Screening
1. Objective: To quantitatively compare the performance of different AL query strategies in identifying active compounds from a large virtual chemical library.
2. Materials & Data Preparation:
- Chemical Library: A publicly available dataset with known bioactivity annotations, such as the ChEMBL database. The library should be divided into a initial training set (e.g., 1% of data) and a hold-out test set (the remaining 99%).
- Descriptors/Fingerprints: Calculate molecular descriptors or fingerprints (e.g., ECFP4, RDKit fingerprints) for all compounds to serve as feature vectors.
- Machine Learning Model: A base predictive model, typically a Random Forest or a Support Vector Machine (SVM), is trained on the initial training set to predict activity.
3. Iterative Active Learning Cycle:
- Step 1 - Model Training: Train the base model on the current labeled training set.
- Step 2 - Prediction & Strategy Application: Use the trained model to predict activities and associated uncertainties for all compounds in the unlabeled pool.
- Apply the AL strategy (e.g., Uncertainty Sampling, Expected Improvement) to select a batch of
ncompounds (e.g., 50-100) from the pool for "labeling."
- Apply the AL strategy (e.g., Uncertainty Sampling, Expected Improvement) to select a batch of
- Step 3 - "Labeling": Instead of wet-lab experimentation, the true activity of the selected compounds is retrieved from the hold-out test set to simulate experimental results.
- Step 4 - Database Update: The newly "labeled" compounds are removed from the unlabeled pool and added to the training set.
- Step 5 - Performance Assessment: Evaluate the updated model on the fixed hold-out test set. Record metrics like the Area Under the Curve (AUC), enrichment factors, and the cumulative number of actives discovered.
- Step 6 - Iteration: Repeat steps 1-5 for a fixed number of cycles or until a performance plateau is reached.
4. Analysis:
- Plot the number of actively discovered compounds against the number of iterative cycles or the total number of compounds selected for each strategy. The strategy that identifies the most actives in the fewest cycles is the most efficient for that specific scenario.
Workflow Visualization
The following diagram illustrates the core, high-level feedback loop that is common to all Active Learning applications in hit discovery, from virtual screening to lead optimization [13] [53].
The strategic decision within the AL cycle is the choice of query function. The next diagram maps the decision logic for selecting an appropriate strategy based on project goals and data context, synthesizing recommendations from the comparative analysis [51] [13].
The Scientist's Toolkit: Essential Research Reagents & Solutions
Successfully implementing an AL-driven hit discovery campaign requires both computational and experimental components. The following table details key solutions and their functions [54] [53].
Table 3: Key Research Reagent Solutions for Hit Discovery Workflows
| Tool / Solution | Function in Hit Discovery | Application Context |
|---|---|---|
| CETSA (Cellular Thermal Shift Assay) [55] | Validates direct target engagement of compounds in intact cells and tissues, bridging biochemical potency and cellular efficacy. | Target engagement confirmation; mechanistic studies. |
| Transcreener Assays [54] | Provides homogeneous, high-throughput measurement of enzyme activity (e.g., kinases, GTPases) via fluorescence detection. | Biochemical high-throughput screening (HTS) and hit-to-lead assays. |
| Cell Painting Assay [53] | A high-content, morphological profiling assay that can identify subtle biological effects of compounds beyond the primary target. | Phenotypic screening; assessment of off-target effects. |
| AI-Powered Protein Language Models [53] | Predicts properties of therapeutic proteins/antibodies (e.g., affinity, stability) to prioritize candidates for synthesis. | Biologics and antibody discovery; reducing library size. |
| Automated Lab Informatics Platforms [53] | Integrates data from diverse assays (biochemical, cell-based, computational) into standardized formats for AL model consumption. | Data harmonization; enabling cross-modal AL. |
The strategic balancing of exploration and exploitation via Active Learning is no longer a theoretical advantage but a practical necessity for efficient hit discovery. As the field advances, the integration of more adaptive, meta-learning approaches and the seamless fusion of AI with automated experimental workflows will further compress discovery timelines [51] [53]. The future lies in self-optimizing discovery systems where the choice of strategy is not a one-time decision but a continuously adaptive process, dynamically responding to the evolving data landscape to maximize the probability of success.
High-throughput screening (HTS) remains a cornerstone of early drug discovery, but its efficiency is often hampered by the resource-intensive nature of testing massive compound libraries [56]. The strategic selection of batch size and composition—the number and specific compounds tested in each iterative cycle—has emerged as a critical factor in determining the success and cost-effectiveness of modern screening campaigns [57] [17]. This guide objectively compares active learning strategies that leverage batch optimization against traditional screening methods, providing supporting experimental data and detailed protocols to inform research practices. As screening paradigms evolve from simplistic "test everything" approaches to intelligent, iterative workflows, understanding these technical considerations becomes essential for researchers aiming to accelerate hit discovery.
Comparative Performance of Screening Strategies
The transition from traditional high-throughput screening to intelligent, iterative approaches represents a significant shift in early drug discovery. The table below provides a quantitative comparison of these strategies based on recent prospective studies and experimental validations.
Table 1: Performance Comparison of Screening Strategies
| Screening Strategy | Typical Batch Size | Library Coverage Required | Hit Rate Enrichment | Key Advantages |
|---|---|---|---|---|
| Traditional HTS | Full library (10^5-10^6 compounds) | 100% | Baseline (e.g., 0.49%) | Comprehensive coverage of chemical library [57] |
| ML-Iterative Screening | 3-5 batches of ~2,000 compounds [57] | 5.9% of 2M library [57] | 43.3% of full HTS hits recovered [57] | Dramatically reduced experimental cost [57] |
| Active Learning (Synergy Screening) | Dynamic, small batches [17] | 10% of combinatorial space [17] | 60% of synergistic pairs found [17] | Optimized for rare event discovery [17] |
| ChemScreener (Active Learning) | N/A | 1,760 compounds total [12] | Hit rate increased to 3-10% (avg. 5.91%) [12] | Identifies diverse chemotypes [12] |
Analysis of Key Differentiators
- Hit Recovery Efficiency: Machine learning-assisted iterative screening demonstrates remarkable efficiency, recovering 43.3% of all primary actives identified in a parallel full HTS while screening just 5.9% of a two-million-compound library [57]. This includes nearly all compound series selected by medicinal chemists, indicating maintained quality alongside reduced quantity.
- Rare Event Discovery: For challenging domains like synergistic drug combination discovery, where positive rates can be as low as 1.47-3.55%, active learning strategies have proven particularly valuable [17]. One study showed that exploring just 10% of the combinatorial space enabled the discovery of 60% of synergistic drug pairs [17].
- Chemical Diversity: Beyond mere hit rates, balanced-ranking acquisition strategies in active learning workflows successfully explore novel chemistry while maintaining hit rate enrichment, leading to the identification of multiple scaffold series and singleton scaffolds from limited screening [12].
Experimental Protocols for Active Learning Screening
Implementing successful active learning-driven screening campaigns requires meticulous experimental design and execution. The following protocols detail key methodologies from recent studies.
Machine Learning-Assisted Iterative HTS Protocol
Application: Prospective screening for salt-inducible kinase 2 (SIK2) inhibitors [57]
Workflow:
- Initialization: Begin with a randomly selected initial batch of compounds from the full library (typically 0.5-1% of total library size).
- Model Training: Train machine learning models (e.g., graph neural networks, random forests) on existing bioactivity data and compound descriptors (e.g., ECFP fingerprints, molecular weight, cLogP).
- Prediction & Prioritization: Use trained models to predict activity for all remaining unscreened compounds and rank them by predicted activity and uncertainty.
- Batch Selection: Select the next batch of compounds for testing using a balanced acquisition function (e.g., prioritizing compounds with high predicted activity and high uncertainty to balance exploration and exploitation).
- Iterative Testing: Synthesize, test, and validate the selected batch in the biological assay system.
- Model Update: Incorporate new experimental results into the training data and retrain models.
- Termination: Repeat steps 3-6 until a predetermined number of cycles are completed or performance metrics plateau.
Key Parameters:
- Batch size: ~2,000 compounds per cycle (representing ~0.1% of a 2M compound library) [57]
- Number of cycles: 3-5 iterations typically sufficient
- Model features: Compound fingerprints, physicochemical properties, and optionally protein-ligand interaction features
Active Learning for Synergistic Drug Combination Screening
Application: Identification of synergistic drug pairs in oncology [17]
Workflow:
- Data Preparation: Compile drug descriptors (Morgan fingerprints, MAP4, MACCS) and cellular features (gene expression profiles from GDSC database).
- Model Selection: Implement neural network architecture with permutation-invariant combination operations (Sum, Max, Bilinear) to handle drug pair inputs.
- Active Learning Loop:
- Use Thompson sampling or upper confidence bound acquisition functions to select the most informative drug-cell combinations for testing.
- Prioritize candidates with high predicted synergy scores and high model uncertainty.
- Conduct experimental testing in high-throughput combination screening platform.
- Update model with new experimental results after each batch.
- Validation: Confirm synergistic effects using validated synergy models (Loewe, Bliss).
Optimization Insights:
- Molecular encoding has limited impact on performance, while cellular environment features significantly enhance predictions [17]
- Small batch sizes with dynamic exploration-exploitation tuning yield highest synergy discovery rates
- As few as 10 carefully selected genes in cellular features can provide sufficient predictive power
Analytical Validation for Screening Hits
Confirmation Workflow:
- Primary Hit Validation: Retest hits with close analogs in concentration-response format [12].
- Counter-Screening: Filter compounds through counter-assays to eliminate technology interference or non-specific binding [12].
- Orthogonal Validation: Confirm binding through biophysical methods like Differential Scanning Fluorimetry (DSF) [12].
- Medicinal Chemistry Triage: Apply expert evaluation to eliminate pan-assay interference compounds (PAINS), promiscuous bioactive compounds, and other problematic chemotypes [56].
Visualizing Active Learning Workflows
Figure 1: Active Learning Screening Workflow. This iterative process dynamically selects screening batches to maximize hit discovery efficiency.
Research Reagent Solutions Toolkit
Successful implementation of optimized screening campaigns requires specific reagents and technologies. The table below details essential materials and their functions in modern screening workflows.
Table 2: Essential Research Reagents and Technologies for Efficient Screening
| Reagent/Technology | Function in Screening | Application Notes |
|---|---|---|
| Acoustic Ejection Mass Spectrometry | Label-free detection for HTS [58] | Enables subsecond analytical cycle times; ideal for cGAS inhibition assays [58] |
| CETSA (Cellular Thermal Shift Assay) | Target engagement validation in intact cells [55] | Confirms dose-dependent stabilization ex vivo and in vivo [55] |
| Gene Expression Profiles (GDSC) | Cellular context features for synergy prediction [17] | 10 selected genes often sufficient for accurate predictions [17] |
| Morgan Fingerprints | Molecular representation for ML models [17] | Circular fingerprints providing structural information for activity prediction [17] |
| HTRF Assay Systems | Biochemical screening platform [12] | Used for primary screening and counter-screening assays [12] |
| Pan-Assay Interference Compound (PAINS) Filters | Computational triage of screening hits [56] | Removes promiscuous bioactive compounds and assay artifacts [56] |
The strategic optimization of batch size and composition represents a paradigm shift in screening efficiency, with active learning approaches demonstrating consistent advantages over traditional methods. The experimental data and protocols presented in this guide provide researchers with practical frameworks for implementing these strategies. As the field evolves, the integration of AI-guided batch selection with advanced analytical technologies and robust experimental validation creates a powerful foundation for accelerating early drug discovery while maintaining scientific rigor and hit quality.
Mitigating Model Bias and Ensuring Robust Performance in Sparse Data Regimes
In hit discovery research, the high cost and time required for experimental screening create a pervasive challenge: vast chemical spaces must be explored with severely limited labeled data. This results in sparse data regimes, where machine learning models are highly susceptible to biased predictions and unreliable performance. Active Learning (AL) has emerged as a powerful paradigm to address this by strategically selecting the most informative experiments, thereby maximizing knowledge gain while minimizing resource expenditure. This guide objectively compares prevalent Active Learning strategies, evaluating their efficacy in mitigating model bias and ensuring robust performance for hit discovery in sparse data environments.
Experimental Protocols for Comparing Active Learning Strategies
To ensure a fair and informative comparison, the evaluated AL strategies were tested under a consistent experimental framework. The following protocols detail the data sources, benchmarked methods, and evaluation criteria used in the cited studies.
Data Sources and Splitting: The primary analysis for anti-cancer drug response was conducted on the Cancer Therapeutics Response Portal v2 (CTRP) dataset, which includes screening data for 494 drugs across 812 cancer cell lines [16]. For materials science benchmarks, nine different formulation design datasets were used, typical of small-sample scenarios due to high data acquisition costs [2]. Data was typically split into training and test sets in an 80:20 ratio, with validation performed automatically within the AutoML workflow using 5-fold cross-validation [2].
Benchmarked Active Learning Strategies: Multiple AL strategies were systematically evaluated and compared against a baseline of random sampling. The core strategies include [16] [2]:
- Uncertainty Sampling: Selects instances where the model's prediction is most uncertain, often using measures like entropy or Monte Carlo Dropout for regression tasks.
- Diversity Sampling: Aims to maximize the diversity of the selected dataset by choosing instances that are most different from the already labeled data.
- Query-by-Committee (QBC): Involves training multiple models and selecting instances where the models disagree the most.
- Expected Model Change: Selects instances that are expected to cause the most significant change to the current model parameters.
- Hybrid Methods: Combine multiple principles, such as a hybrid of diversity and uncertainty (e.g., RD-GS), to balance exploration and exploitation.
Evaluation Criteria: Performance was assessed based on two primary goals of hit discovery [16]:
- Hit Identification Efficiency: The number of responsive treatments (hits) identified early in the screening process.
- Model Performance: The accuracy of the drug response prediction model trained on the selected data, measured by metrics like Mean Absolute Error (MAE) and the Coefficient of Determination (R²) [2].
Comparative Performance of Active Learning Strategies
The table below summarizes the quantitative performance of different AL strategies as reported in benchmark studies.
Table 1: Comparative Performance of Active Learning Strategies
| Active Learning Strategy | Reported Performance vs. Random Sampling | Key Strengths | Key Limitations |
|---|---|---|---|
| Uncertainty Sampling | Outperforms random in hit identification and, for some drugs, model performance [16]. A "reality check" study found entropy (uncertainty) outperformed all other methods in 72.5% of steps [59]. | Highly effective at refining decision boundaries and identifying challenging cases; simple to implement [60] [59]. | Can be myopic, potentially selecting outliers; performance may depend on a well-calibrated model [2]. |
| Diversity Sampling | Shows improvement, particularly when combined with other methods in hybrid strategies [2]. | Ensures broad coverage of the chemical space, reducing the risk of missing novel hit clusters [16]. | May select many trivial examples if not guided by model performance [2]. |
| Query-by-Committee | Effective in identifying hits and improving model performance, saving 70-95% of labeling resources in some material science applications [2]. | Reduces model-specific bias by leveraging ensemble disagreement; enhances model robustness [60]. | Computationally expensive due to training multiple models [60]. |
| Hybrid (Uncertainty + Diversity) | Uncertainty-driven (LCMD) and diversity-hybrid (RD-GS) strategies "clearly outperform" geometry-only heuristics and baseline early in acquisition [2]. | Balances exploration of new regions with exploitation of uncertain areas, leading to more robust performance [16] [2]. | Strategy weighting can be complex; may inherit some limitations of constituent methods. |
Table 2: Experimental Results from a Comprehensive AL Benchmark in Drug Response [16]
| Sampling Method Category | Performance in Identifying Hits | Impact on Model Prediction Performance |
|---|---|---|
| Random (Baseline) | Baseline | Baseline |
| Greedy | Lower than other AL methods | Lower than some AL methods |
| Uncertainty-Based | Significant improvement | Improvement for some drugs |
| Diversity-Based | Significant improvement | Improvement for some drugs |
| Hybrid Approaches | Significant improvement | Improvement for some drugs |
The Active Learning Workflow for Hit Discovery
The following diagram illustrates the standard iterative workflow of an Active Learning cycle, adapted for a hit discovery campaign.
The Scientist's Toolkit: Essential Reagents for AL-Driven Experiments
Implementing an effective AL pipeline for hit discovery requires a combination of data, computational tools, and experimental resources.
Table 3: Key Research Reagent Solutions for AL-Driven Hit Discovery
| Item / Resource | Function / Application | Example / Implementation Note |
|---|---|---|
| Curated Bioactivity Dataset | Serves as the foundational data for initial model training and the unlabeled pool for querying. | Cancer Therapeutics Response Portal (CTRP) [16]; other repositories like ChEMBL or PubChem. |
| Automated Machine Learning (AutoML) | Automates the selection and optimization of machine learning models, reducing manual tuning and mitigating model selection bias. | Integrated into the AL cycle to ensure the surrogate prediction model is consistently optimal at each iteration [2]. |
| Uncertainty Quantification Method | Provides the core metric for uncertainty-based AL strategies. | Techniques like Monte Carlo Dropout or ensemble methods to estimate predictive variance for regression tasks [2]. |
| High-Throughput Screening (HTS) Assay | Acts as the "oracle" in the AL loop, providing experimental validation for the selected compounds. | Must be robust and scalable to provide rapid feedback for the iterative AL process [13]. |
| Diversity & Featurization Tools | Enables diversity-based and hybrid AL strategies by quantifying molecular similarity. | Requires high-quality molecular descriptors (e.g., fingerprints, graph representations) or genomic signatures for cell lines [16]. |
In the demanding context of sparse data regimes for hit discovery, Active Learning strategies demonstrably outperform traditional random or greedy screening approaches. Benchmark studies reveal that while simple uncertainty-based methods like entropy sampling are surprisingly robust and difficult to beat, hybrid strategies that balance uncertainty with diversity often provide the most consistent and efficient path to identifying hits and building accurate predictive models. The choice of an optimal AL strategy is not universal; it depends on the specific dataset, the cost of experimentation, and the primary objective—whether to maximize immediate hit discovery or to build a generically powerful model. Integrating these strategies with modern tools like AutoML and robust experimental workflows offers a principled approach to mitigating model bias and accelerating the drug discovery pipeline.
In the field of hit discovery research, Active Learning (AL) has emerged as a powerful strategy to accelerate the identification of promising drug candidates while minimizing resource-intensive experimental work. AL operates on a simple but profound principle: instead of randomly screening vast chemical libraries, an algorithm iteratively selects the most informative compounds for experimental testing, thereby improving a predictive model with each cycle [61]. However, the performance of AL is highly dependent on its guidance system. Naive AL strategies, which rely solely on statistical uncertainty, often fail to account for the complex realities of biological systems, leading to suboptimal exploration of chemical space.
This guide posits that the integration of deep domain knowledge—specifically from cellular context and structural biology—is the critical differentiator between a merely functional AL strategy and a transformative one. Incorporating this knowledge grounds computational exploration in biological plausibility, steering the search toward compounds that are not only potent but also functionally relevant and developable. This article provides a comparative analysis of contemporary AL strategies, evaluating their performance and practical utility for researchers engaged in early-stage drug discovery.
Domain Knowledge as a Strategic Guide in Active Learning
Integrating domain knowledge into AL moves the process beyond abstract statistical sampling and into a biologically intelligent search. This integration typically occurs in two key areas:
Cellular Context: This refers to the physiological environment in which a drug target operates. AL strategies informed by cellular context prioritize compounds that demonstrate efficacy in complex, living systems rather than just in purified protein assays. For instance, some leading AI-driven platforms incorporate high-content phenotypic screening on real patient-derived samples to ensure translational relevance [4]. Functionally relevant assays, such as the Cellular Thermal Shift Assay (CETSA), provide direct, empirical evidence of target engagement within intact cells, offering a powerful data stream to guide an AL model toward biologically meaningful chemical regions [55].
Structural Biology: This involves the precise three-dimensional structure of a biological target. AL powered by structural knowledge focuses on the physical principles of molecular interaction. The core challenge has been the "generalizability gap"—where models perform poorly on novel protein families not seen in their training data [62]. Innovative approaches now address this by constraining models to learn from the fundamental physicochemical interactions between atom pairs, forcing the AL system to learn transferable principles of molecular binding rather than memorizing structural shortcuts [62]. Physics-based simulations, when combined with machine learning, create a powerful hybrid approach that ensures generated molecules are not only likely to bind but are also physically plausible [4] [63].
The following diagram illustrates how these two domains of knowledge can be integrated into a cohesive AL workflow for hit discovery.
Comparative Analysis of Leading Active Learning Strategies
Different AL implementations vary significantly in how they leverage domain knowledge, which directly impacts their performance and suitability for specific research goals. The table below provides a structured comparison of several prominent strategies based on recent research.
Table 1: Performance Comparison of Active Learning Strategies in Drug Discovery
| Strategy / Model | Core Approach to Domain Knowledge | Reported Performance / Impact | Key Experimental Findings |
|---|---|---|---|
| muTOX-AL [61] | Uses molecular fingerprints and descriptors to quantify structural similarity for mutagenicity prediction. | Reduced required training samples by ~57% compared to random sampling to achieve the same accuracy. | Showed high structural discriminability, selecting molecules with high similarity but opposite properties, efficiently defining the activity boundary. |
| Brown's Generalizable Framework [62] | Focuses model exclusively on the physicochemical interaction space of atom pairs, ignoring overall protein structure to avoid shortcuts. | Established a reliable baseline for generalizability; modest performance gains but high reliability on novel protein targets. | When tested on held-out protein superfamilies, the model maintained performance, unlike contemporary models which showed significant drops. |
| Schrödinger's Hybrid Workflow [63] | Integrates machine learning with physics-based free energy perturbation (FEP) calculations, using AL to optimize simulation protocols. | Explored 23 billion compound designs in 6 days; identified novel, selective scaffolds with >10,000x selectivity. | Automated the traditionally manual process of FEP+ protocol setup, accelerating the discovery of potent and selective inhibitors for targets like EGFR and WEE1. |
| BoltzGen [64] | A unified generative model for structure prediction & design, with built-in physical constraints (e.g., on protein folding). | Generated novel protein binders for 26 therapeutically relevant targets, including "undruggable" ones; validated in 8 wet labs. | Successfully created functional protein binders from scratch, expanding AI's reach from understanding biology to engineering it for hard-to-treat diseases. |
Key Performance Insights from Comparative Data
- Annotation Efficiency is Quantifiable: The direct comparison in Table 1 demonstrates that informed AL strategies like muTOX-AL can dramatically reduce experimental burden. A 57% reduction in required labeled samples translates to significant cost and time savings, compressing the traditional design-make-test-analyze (DMTA) cycle from months to weeks [55] [61].
- Generalizability is a Critical Metric: Performance on familiar benchmarks can be deceptive. As highlighted by Brown's work, the true test of an AL system is its performance on novel protein families, a scenario that mimics real-world discovery for new targets [62]. Strategies that fail this test can lead to expensive, dead-end exploration.
- Scale and Precision are Not Mutually Exclusive: Schrödinger's application shows that AL can manage immense search spaces (billions of designs) without sacrificing the precision offered by physics-based methods. This synergy allows for both broad exploration and confident optimization [63].
Detailed Experimental Protocols for Key Studies
To enable replication and critical assessment, this section details the experimental methodologies from two pivotal studies compared in this guide.
Protocol 1: Rigorous Generalization Test for Structure-Based AL
This protocol, based on the work of Brown (2025), is designed to rigorously evaluate an AL model's ability to predict binding affinity for novel protein targets [62].
- Dataset Curation: Assemble a large and diverse dataset of protein-ligand complexes with experimentally measured binding affinities (e.g., from PDBBind).
- Data Splitting by Protein Similarity: Partition the data at the level of protein superfamilies. Entire superfamilies and all their associated chemical data are completely left out of the training and validation sets to form the test set. This simulates a real-world scenario of predicting interactions for a newly discovered protein.
- Model Architecture (Interaction-Space Focus):
- Input: Instead of the full 3D structure of the protein and ligand, the model is fed only a representation of their interaction space. This is typically a matrix capturing the distance-dependent physicochemical interactions (e.g., van der Waals, electrostatic) between all atom pairs across the protein and ligand.
- Training: The model is trained on the training set to predict binding affinity from this interaction-based input.
- Evaluation: The model's performance is evaluated exclusively on the held-out test set of novel protein superfamilies. Key metrics include the Root Mean Square Error (RMSE) and Pearson Correlation Coefficient (r) between predicted and experimental affinities.
Protocol 2: Active Learning for Molecular Mutagenicity Prediction (muTOX-AL)
This protocol outlines the AL cycle used by muTOX-AL to efficiently build a predictive model for mutagenicity, a critical safety endpoint in drug discovery [61].
- Initialization:
- Dataset: Use a curated mutagenicity dataset (e.g., TOXRIC, with ~7,500 compounds).
- Pool Setup: Start with a very small initial labeled pool (e.g., 200 randomly selected molecules). The remaining compounds form the unlabeled pool.
- Model Training:
- Feature Extraction: Compute molecular fingerprints and descriptors for the labeled compounds.
- Backbone Network: Train a deep learning model (the "backbone") to predict mutagenicity (binary classification).
- Uncertainty Estimation: Simultaneously, train an uncertainty estimation module that takes hidden layer features from the backbone as input.
- The Active Learning Loop:
- Query: Use the trained model to calculate an "informativeness" score (e.g., predictive uncertainty) for every molecule in the unlabeled pool.
- Selection: Select the top k molecules with the highest uncertainty scores.
- Oracle: Send these selected molecules for experimental annotation (e.g., in vitro Ames test), simulating the interaction with a wet lab.
- Update: Add the newly labeled molecules to the training pool and remove them from the unlabeled pool.
- Iteration: Retrain the model with the expanded labeled set and repeat the AL loop until a performance plateau is reached or the annotation budget is exhausted.
The strategic logic of the muTOX-AL protocol is visualized below, highlighting its iterative, human-in-the-loop nature.
The Scientist's Toolkit: Essential Research Reagents and Solutions
Successfully implementing an AL strategy for hit discovery requires a combination of computational tools and experimental assays. The following table details key resources referenced in the compared studies.
Table 2: Key Research Reagent Solutions for Informed Active Learning
| Tool / Assay | Function in the AL Workflow | Relevance to Domain Knowledge |
|---|---|---|
| CETSA (Cellular Thermal Shift Assay) [55] | Provides quantitative, direct measurement of drug-target engagement in intact cells and native tissue environments. | Supplies cellular context by confirming a compound's mechanistic action and binding within a physiologically relevant system, guiding AL toward functionally active chemotypes. |
| TOXRIC Database [61] | A public database of molecular structures with curated mutagenicity labels, used for training and benchmarking predictive models. | Provides structural and toxicological knowledge, allowing AL models to learn and avoid structural motifs associated with genotoxicity, de-risking the discovery pipeline. |
| FEP+ (Free Energy Perturbation) [63] | A physics-based computational method that provides highly accurate predictions of relative binding free energies between related compounds. | Incorporates structural and thermodynamic knowledge to precisely rank compounds, often used as a high-fidelity oracle or validation step within an AL cycle to prioritize synthesis. |
| Phenotypic Screening Platforms [4] | High-content screening (e.g., using patient-derived cells) that measures complex biological outcomes, not just single-target binding. | Informs AL with disease-relevant cellular context, steering compound selection toward those that elicit a desired phenotypic response, thereby improving translational potential. |
| Interaction-Space Modeling Framework [62] | A specialized deep learning architecture for predicting protein-ligand affinity based solely on pairwise physicochemical interactions. | Embeds structural knowledge in a way that enforces generalizability, making the AL strategy robust and reliable when exploring new target classes. |
The comparative analysis presented in this guide unequivocally demonstrates that the strategic incorporation of domain knowledge is a powerful lever for enhancing Active Learning in hit discovery. The most effective contemporary strategies are not purely data-driven; they are hybrid systems that seamlessly integrate:
- Cellular Context from functional assays like CETSA and phenotypic screening to ensure biological relevance.
- Structural Knowledge from physics-based simulations and generalizable interaction models to ensure precision and reliability.
As the field evolves, the distinction between AL and full-cycle drug design is blurring. The advent of generative models like BoltzGen, which can create novel binders from scratch, represents the next frontier: closed-loop systems where AL guides not only screening but also the de novo design of molecules [64]. For researchers, the imperative is clear: adopt AL strategies that are deeply informed by the language of biology and physics. This is the most reliable path to compressing timelines, reducing attrition, and delivering breakthrough therapeutics.
In the high-stakes field of drug discovery, active learning (AL) has emerged as a transformative methodology for navigating vast chemical spaces efficiently. Unlike traditional high-throughput screening (HTS) that relies on brute-force experimental testing, AL employs an iterative, data-driven selection process to identify the most promising compounds for experimental validation. For researchers in hit discovery, establishing a robust benchmarking framework is crucial for selecting the optimal AL strategy, ultimately accelerating the identification of novel therapeutic candidates while significantly reducing resource consumption. This guide provides a structured comparison of prevalent AL strategies, supported by experimental data and practical implementation protocols to equip scientists with the tools needed to design successful AL campaigns.
Quantitative Comparison of Active Learning Strategies
The table below synthesizes performance data from a comprehensive benchmark study that evaluated 17 AL strategies on materials science regression tasks, providing insights directly applicable to chemical property prediction and hit discovery in drug development. [2]
Table 1: Performance Comparison of Active Learning Strategy Types
| Strategy Category | Representative Methods | Early-Stage Performance (Data-Scarce) | Late-Stage Performance (Data-Rich) | Key Characteristics & Best Use Cases |
|---|---|---|---|---|
| Uncertainty-Based | LCMD, Tree-based-R | Superior – Clearly outperforms baseline and geometry methods [2] | Converges with other methods [2] | Selects samples where model predictions are least certain; ideal for rapidly improving model accuracy. |
| Diversity-Based | GSx, EGAL | Lower performance in early stages [2] | Converges with other methods [2] | Selects diverse samples to cover chemical space; best used when model performance is stable. |
| Hybrid (Uncertainty + Diversity) | RD-GS | Superior – Outperforms baseline and geometry methods [2] | Converges with other methods [2] | Balances exploration (diversity) and exploitation (uncertainty); robust for general-purpose use. |
| Random Sampling (Baseline) | Random | Lower performance in early stages [2] | Converges with other methods [2] | Serves as a essential baseline for comparing the added value of intelligent AL strategies. |
Key Benchmarking Insight
A critical finding from recent research is the diminishing returns of AL as the labeled dataset grows. [2] The performance gap between sophisticated strategies and random sampling is most pronounced during the early, data-scarce phase of a campaign. This underscores the paramount importance of strategy selection at the project's inception.
Experimental Protocols for AL Benchmarking
To ensure reliable and reproducible comparison of AL strategies, researchers should adopt a standardized experimental framework. The following protocol, adapted from a rigorous benchmark study, provides a robust methodology. [2]
Workflow for AL Benchmarking
The following diagram illustrates the standardized workflow for a single experimental trial of an Active Learning campaign.
Core Protocol Components
Dataset Partitioning & Initialization: Begin with a fixed dataset. Partition it into a training pool (80%) and a hold-out test set (20%). From the training pool, a very small subset of data points (
n_init) is randomly selected to form the initial labeled setL, while the remainder constitutes the unlabeled poolU. [2]Model Training & Validation: In each AL cycle, a model is trained on the current labeled set
L. The benchmark study highlights the use of Automated Machine Learning (AutoML) to automatically search for the best model architecture and hyperparameters, which is particularly valuable for non-ML-experts and for ensuring fair comparison across strategies. Model validation is typically performed using 5-fold cross-validation. [2]Query Strategy & Annotation: The core of the AL cycle. A predefined query strategy (e.g., uncertainty sampling) selects the most informative candidate
x*from the unlabeled poolU. In a benchmark, the "oracle" (e.g., experimental measurement) is simulated by retrieving the ground-truth labely*. [2]Performance Metrics & Iteration: The newly labeled sample
(x*, y*)is added toLand removed fromU. The updated model's performance is evaluated on the hold-out test set using metrics like Mean Absolute Error (MAE) and the Coefficient of Determination (R²) for regression tasks. This process repeats until a stopping criterion is met (e.g., a predefined budget or performance plateau). [2]
The Scientist's Toolkit: Essential Research Reagents & Solutions
Successfully implementing an AL campaign requires a suite of computational and experimental tools. The table below details key resources mentioned across the analyzed literature.
Table 2: Essential Reagents & Solutions for an AL Campaign
| Tool Category | Specific Examples / Platforms | Function in AL for Hit Discovery |
|---|---|---|
| AI/ML Platforms | Exscientia's Centaur Chemist, Insilico Medicine's Generative AI Platform, Schrödinger's Physics-Enabled Design [4] [65] | Provides end-to-end frameworks for integrating AL into the drug design cycle, from target identification to lead optimization. |
| Automated ML (AutoML) | AutoML frameworks (as featured in benchmark study) [2] | Automates model selection and hyperparameter tuning, reducing manual effort and ensuring robust performance in the AL loop. |
| Virtual Screening Software | Structure-based (e.g., AtomNet) and ligand-based tools [66] [67] | Enables the initial computational screening of ultra-large chemical libraries to create a candidate pool for the AL cycle. |
| Chemical Databases | Large in-house or commercial compound libraries (e.g., ZINC, Enamine) | Serves as the source of unlabeled data (U pool) from which the AL strategy selects compounds for experimental testing. |
| Experimental Assay Systems | High-throughput screening (HTS) assays, phenotypic screens [66] | Acts as the "oracle" within the AL loop, providing the experimental data (labels) for the compounds selected by the AL model. |
Validated Case Study: Active Learning in Action
A compelling example of AL's effectiveness comes from a benchmark where an uncertainty-driven strategy curtailed an experimental campaign in alloy design by more than 60%. [2] Furthermore, a separate study demonstrated that an AL scheme achieved performance parity with full-data baselines while querying only 30% of the data pool, equivalent to a 70-95% savings in computational or labeling resources. [2] This level of efficiency is directly translatable to hit discovery, where each experimental data point carries significant cost.
Another successful application involves benchmark selection for SAT solver development. An active learning approach could predict a new solver's rank with 92% accuracy after using only about 10% of the time it would take to run the solver on the entire benchmark dataset. [68] This demonstrates the power of AL for efficient performance prediction and ranking, a common need when comparing multiple candidate molecules or models.
For researchers embarking on hit discovery campaigns, the evidence is clear: the strategic implementation of active learning can dramatically enhance efficiency and success rates. Benchmarking studies consistently show that uncertainty-based and hybrid AL strategies provide the most significant early-stage advantages when labeled data is scarce and costly. [2]
To maximize the return on investment from an AL campaign, focus on the critical setup phase: establish a robust, AutoML-enabled benchmarking protocol, select a strategy aligned with your project's stage (data-scarce vs. data-rich), and leverage the growing ecosystem of AI-driven discovery platforms. By adopting these evidence-based practices, drug development professionals can position themselves at the forefront of pharmaceutical innovation.
Evaluating Success: Benchmarking Active Learning Performance Against Traditional Methods
In the high-stakes field of drug discovery, hit enrichment—the process of identifying promising chemical compounds with desired biological activity—is a critical early bottleneck. The vastness of chemical space, coupled with the extreme cost and time requirements of physical experiments, makes exhaustive screening impractical. Active Learning (AL) has emerged as a transformative machine learning paradigm that strategically selects the most informative experiments to run, dramatically accelerating the identification of hits and the development of accurate predictive models [13] [38].
This guide provides a comparative analysis of active learning strategies for hit discovery, framing the evaluation within the essential quantitative metrics used to measure success. We define "hit enrichment" as the efficiency of an AL strategy in identifying the highest number of true active compounds with the fewest experiments. Conversely, "model accuracy" refers to the predictive performance of the machine learning model that guides the AL process, which is crucial for its long-term utility [38]. For researchers and scientists, understanding the interplay between these concepts and the metrics that define them is key to selecting and optimizing an AL strategy for their specific drug discovery pipeline.
Core Metrics for Evaluating Performance
Evaluating an Active Learning system requires a dual focus: one set of metrics to assess the quality of the underlying model and another to measure its experimental efficiency.
Model Accuracy Metrics
Model accuracy in classification settings is best understood through the confusion matrix, which breaks down predictions into True Positives (TP), False Positives (FP), True Negatives (TN), and False Negatives (FN) [69]. From this matrix, several key metrics are derived, each with a specific interpretive value.
- Precision answers the question: "Of all the compounds the model predicted as hits, how many are actual hits?" It is calculated as TP / (TP + FP). High precision is critical when the cost of false positives (e.g., pursuing inactive leads) is high [70].
- Recall (Sensitivity) answers: "Of all the actual hits, how many did the model correctly identify?" It is calculated as TP / (TP + FN). High recall is paramount when missing a true positive (e.g., a potential therapeutic) is more costly than following up on a false alarm [70].
- F1 Score is the harmonic mean of precision and recall, providing a single metric to balance the trade-off between the two. It is especially useful for comparing models when you need a straightforward, balanced assessment [69] [70].
- AUC-ROC (Area Under the Receiver Operating Characteristic Curve) measures the model's ability to distinguish between classes (e.g., active vs. inactive) across all possible classification thresholds. An AUC of 1.0 represents perfect separation, while 0.5 represents a model no better than random guessing. It is particularly valuable when the optimal decision threshold is not yet known [69] [70].
Table 1: Key Model Evaluation Metrics for Hit Discovery
| Metric | Definition | Interpretation & When to Use | Ideal Value |
|---|---|---|---|
| Precision | (\frac{\text{True Positives}}{\text{True Positives + False Positives}}) | Measures the model's reliability in flagging true hits. Use when the cost of false positives is high. | Closer to 1.0 |
| Recall (Sensitivity) | (\frac{\text{True Positives}}{\text{True Positives + False Negatives}}) | Measures the model's ability to find all actual hits. Use when missing a hit is unacceptable. | Closer to 1.0 |
| F1 Score | (2 \times \frac{\text{Precision} \times \text{Recall}}{\text{Precision + Recall}}) | A balanced measure of precision and recall. Use for a single metric to compare models. | Closer to 1.0 |
| AUC-ROC | Area under the ROC curve | Measures the model's overall class separation capability, independent of a chosen threshold. | Closer to 1.0 |
| Accuracy | (\frac{\text{Correct Predictions}}{\text{All Predictions}}) | The overall fraction of correct predictions. Can be misleading with imbalanced datasets [71]. | Closer to 1.0 |
Hit Enrichment Metrics
While model metrics are foundational, the ultimate test of an AL strategy in a real-world setting is its experimental efficiency.
- Cumulative Hit Discovery is a fundamental plot that shows the number of true hits identified versus the number of experiments conducted. A superior AL strategy will curve upwards more steeply, finding more hits faster than a random or baseline strategy [38].
- Early Enrichment is a critical concept in hit discovery. The primary goal is often to find a significant portion of the available hits within the first few rounds of experimentation, conserving resources. The performance of strategies is frequently compared by the number of hits found after a fixed, small percentage of the total possible experiments have been run [38].
Comparative Analysis of Active Learning Strategies
Different AL strategies use distinct query functions to select which experiments to run next. The following table summarizes the core strategies investigated in recent anti-cancer drug screening research, providing a direct comparison of their performance and characteristics [38].
Table 2: Comparison of Active Learning Strategies for Anti-Cancer Drug Screening
| AL Strategy | Core Principle | Performance: Hit Discovery | Performance: Model Accuracy | Key Advantage | Key Limitation |
|---|---|---|---|---|---|
| Uncertainty Sampling | Selects data points where the model's prediction is most uncertain (e.g., closest to 0.5 probability). | Fast initial identification of hits, but may plateau. | Rapid initial improvement by resolving ambiguity. | Directly targets the model's knowledge gaps. | Can get stuck exploring local regions of uncertainty. |
| Diversity Sampling | Selects a diverse set of data points to maximize coverage of the chemical space. | Slower start but can find more diverse hits long-term. | Builds a robust, generalizable model foundation. | Explores the search space broadly. | May waste resources on obviously inactive compounds. |
| Greedy Sampling | Selects data points predicted to be hits (e.g., highest probability). | Can find hits quickly if initial model is good. | Model can become biased and overfit to initial predictions. | Maximizes short-term yield of hits. | High risk of getting trapped in local maxima. |
| Hybrid (Uncertainty + Diversity) | Combines uncertainty and diversity criteria to balance exploration and exploitation. | Superior and robust performance in identifying hits efficiently [38]. | Leads to stable and accurate models that generalize well. | Balanced approach mitigates the weaknesses of individual strategies. | More computationally complex to implement. |
The experimental data from a comprehensive investigation on anti-cancer drug screening for 57 drugs revealed that while most AL strategies outperformed random selection, hybrid approaches consistently demonstrated superior efficiency. For instance, a hybrid method could identify the same number of hits as a random strategy using only 20-30% of the experimental budget [38].
Experimental Protocols for Benchmarking AL Strategies
To ensure a fair and reproducible comparison of AL strategies, a standardized experimental protocol is essential. The following workflow, adapted from a study on anti-cancer drug screening, outlines a robust methodology [38].
Detailed Methodology
- Dataset Preparation: A large dataset of compounds (e.g., from a chemical library) is split into a small initial training set (e.g., 1-5% randomly selected and labeled), a large unlabeled pool, and a hold-out test set for final evaluation. Public datasets like the Cancer Cell Line Encyclopedia (CCLE) are often used [38].
- Model Training: An initial predictive model (e.g., Random Forest, Gradient Boosting, or a Neural Network) is trained on the small labeled set. The model's task is to predict a response value, such as IC50 or AUC, for a compound-cell line pair [38].
- Iterative Active Learning Loop: This core loop repeats for a fixed number of iterations or until a performance target is hit.
- Prediction & Strategy: The current model predicts on the entire unlabeled pool. A pre-defined AL strategy (see Table 2) is applied to score and rank all unlabeled compounds.
- Selection & 'Experimentation': The top-k ranked compounds (e.g., 10-100) are selected. In a simulation, their labels are retrieved from the database. In a real-world setting, these would be sent for physical screening.
- Model Update: The newly labeled compounds are added to the training set, and the model is retrained from scratch or fine-tuned.
- Evaluation: Throughout the process and at its conclusion, the strategy's performance is tracked using the metrics outlined in Section 2. This includes plotting cumulative hits and calculating model precision/recall on the hold-out test set.
The Scientist's Toolkit: Research Reagent Solutions
The following table details key computational and data resources essential for conducting AL-driven hit discovery experiments.
Table 3: Essential Research Reagents & Resources for Computational Screening
| Resource / 'Reagent' | Type | Function in the Experiment | Example Sources / Libraries |
|---|---|---|---|
| Chemical Compound Library | Dataset | Provides the vast "search space" of potential hits for the AL algorithm to explore. | ZINC, PubChem, in-house corporate libraries. |
| Bioactivity Data | Dataset | Serves as the initial labeled data for model training; provides ground truth for validation. | ChEMBL, CCLE, GDSC, internal HTS data. |
| Molecular Descriptors/Fingerprints | Computational Representation | Converts chemical structures into numerical vectors that machine learning models can process. | RDKit, Mordred, ECFP fingerprints. |
| Active Learning Framework | Software | Provides the infrastructure and algorithms for implementing various query strategies and workflows. | scikit-learn, modAL, AWSOME-AL [1]. |
| Predictive Model Algorithm | Software | The core machine learning model that learns from data to predict compound activity. | Random Forest (scikit-learn), XGBoost, Deep Neural Networks (PyTorch, TensorFlow). |
Interpretation of Results and Strategic Selection
The choice of the optimal AL strategy is not one-size-fits-all; it depends heavily on the project's primary goal. The following diagram illustrates the logical decision process for selecting a strategy based on the findings from the comparative analysis.
- For Rapid Initial Hit Identification: If the goal is to quickly validate a pipeline or generate a small number of leads for immediate follow-up, a Greedy strategy can be effective, provided the initial model is reasonably accurate.
- For Comprehensive Exploration: If the goal is to thoroughly map a chemical space or build a maximally robust predictive model for future use, a Diversity-based strategy is preferable, despite its potentially slower start.
- For Overall Efficiency and Robustness: In most practical scenarios, where the goal is to efficiently find as many hits as possible without excessive risk of failure, the evidence strongly supports using a Hybrid strategy. Combining uncertainty and diversity sampling balances the need for exploration and exploitation, leading to superior and more reliable outcomes across diverse drug datasets [38].
The systematic quantification of success through well-defined metrics is fundamental to advancing hit discovery research. As demonstrated, Active Learning provides a powerful framework for optimizing this process, but its effectiveness hinges on the careful selection of a query strategy aligned with project objectives. The comparative data presented in this guide underscores that while specialized strategies have their place, hybrid approaches generally offer the most robust and efficient path for both hit enrichment and model accuracy. For researchers, adopting this rigorous, metrics-driven framework is key to accelerating the journey from a vast chemical space to a promising shortlist of therapeutic candidates.
Active learning (AL) strategies demonstrate a significant and consistent advantage over random and greedy screening methods in hit discovery across multiple scientific domains. Quantitative benchmarks from recent large-scale studies in drug discovery and materials science reveal that AL can identify the majority of hits after screening only a small fraction (often 3% or less) of the total experimental space. This data-driven approach achieves hit rates that are 6 to 24 times more efficient than random selection, while also outperforming greedy methods that focus solely on immediate reward. The following guide provides a detailed comparison of their performance, supported by experimental data and methodologies.
Quantitative Performance Benchmarks
The table below summarizes key quantitative findings from recent high-quality studies that directly compare active learning, random, and greedy sampling.
Table 1: Comparative Performance of Active Learning vs. Random and Greedy Screening
| Application Domain | Key Performance Metric | Active Learning (AL) | Random Screening | Greedy Screening | Source & Context |
|---|---|---|---|---|---|
| Preclinical Cancer Drug Screening | Hit Identification Efficiency | Most AL strategies were more efficient than random selection for identifying effective treatments [16]. | Used as a baseline for comparison. | Was outperformed by multiple AL approaches in identifying hits [16]. | Analysis of 57 drugs from the CTRPv2 dataset [16]. |
| Multi-Target Drug Profiling | Hit Discovery Rate | Discovered ~60% of all hits after exploring only 3% of the experimental space [72]. | Required full exploration (100%) to achieve same result. | Not explicitly measured in this study. | Dataset of 177 assays & 20,000 compounds from PubChem [72]. |
| Virtual Screening (Docking) | Top-50,000 Compound Retrieval | Identified 58.97% of top compounds after screening 0.6% of a 99.5M compound library [73]. | Retrieval rate is proportional to fraction screened (e.g., ~0.6% for random). | Performance was surpassed by pretrained AL models [73]. | Benchmark on Enamine REAL library (99.5 million compounds) [73]. |
| Materials Science Regression | Model Accuracy (Early Stage) | Uncertainty & diversity-hybrid strategies clearly outperformed baseline and geometry-only heuristics [2]. | Used as a baseline for comparison. | Geometry-only heuristics (GSx, EGAL) were outperformed [2]. | Benchmark with AutoML on 9 small-sample materials datasets [2]. |
Detailed Experimental Protocols
The superior performance of active learning is demonstrated through rigorous, iterative workflows. The following diagram generalizes the core active learning cycle used across the cited studies.
- Objective: To build drug-specific models predicting the response of cancer cell lines to a specific drug and identify effective treatments (hits).
- Data Source: Cancer Therapeutics Response Portal v2 (CTRP), comprising hundreds of cancer cell lines and drugs [16].
- Workflow:
- Initialization: Start with a small, randomly selected set of experimentally tested cell line-drug pairs.
- Model Training: Train a drug response prediction model (e.g., using random forests or neural networks) on the available data.
- Candidate Selection: Apply an acquisition function to score all untested cell lines for a given drug. The tested strategies included:
- Uncertainty Sampling: Selects cell lines where the model's prediction is most uncertain.
- Diversity Sampling: Selects cell lines that are most dissimilar to those already in the training set.
- Hybrid Approaches: Combine uncertainty and diversity, or greedy and uncertainty.
- Greedy Sampling: Selects cell lines predicted to be the most responsive (highest predicted activity).
- Iteration: The selected top candidates are "tested" (their labels are retrieved from the database), added to the training set, and the cycle repeats.
- Evaluation: Performance was measured by (a) the number of true hits identified over iterations, and (b) the prediction performance of the model trained on the acquired data.
- Objective: To efficiently discover active compound-target pairs (hits) across a vast experimental space.
- Data Source: A dataset constructed from PubChem, containing 177 assays (targets) and 20,000 compounds, totaling 3.54 million possible experiments [72].
- Modeling: A joint model used compound fingerprints (for small molecules) and protein features (for targets) to predict bioactivity scores for any compound-target pair [72].
- Active Learning Loop:
- The model was trained on an initial random batch of experiments.
- The trained model was used to predict scores for all unexplored compound-target pairs.
- The next batch of experiments was selected based on the highest predicted activity (a greedy-like exploitation strategy) or other criteria to improve the model.
- New experimental data was incorporated, and the process repeated.
- Comparison: The hit discovery rate of AL was compared against a simulated random screening campaign that selected batches of experiments at random.
Table 2: Key Resources for Implementing Active Learning in Hit Discovery
| Category | Item / Resource | Function / Description | Example from Literature |
|---|---|---|---|
| Data Resources | Cell Line & Drug Response Database | Provides ground-truth experimental data for training and validating models. | Cancer Therapeutics Response Portal (CTRPv2) [16] |
| PubChem Bioassay Database | A public repository of chemical compounds and their biological activities. | Source for 177 assays and 20,000 compounds [72] | |
| Ultra-Large Compound Libraries | Virtual libraries of synthesizable molecules for virtual screening. | Enamine REAL database (Billions of compounds) [73] | |
| Computational Tools | Surrogate Machine Learning Model | Predicts properties (e.g., docking score, bioactivity) for unexplored candidates. | Graph Neural Networks (GNNs), Random Forests [16] [73] |
| Molecular Docking Software | Computes binding affinity and pose of a ligand to a protein target. | AutoDock Vina [73] | |
| Feature Representation | Converts molecular and protein data into a machine-readable format. | Molecular fingerprints, protein sequence features [72] | |
| Methodological Components | Acquisition Function | The core AL algorithm that selects the most informative candidates for the next experiment. | Uncertainty Sampling, Greedy Selection, Upper Confidence Bound (UCB) [16] [73] |
Critical Analysis and Strategic Recommendations
The accumulated evidence strongly indicates that active learning is not merely an incremental improvement but a paradigm shift for efficient resource allocation in experimental sciences.
- AL vs. Random Screening: The efficiency gain is profound. Discovering 60% of hits from just 3% of the space, as demonstrated in [72], translates to a 20-fold efficiency increase. In ultra-large virtual screening, this reduces computational cost from prohibitive to manageable [73].
- AL vs. Greedy Screening: While greedy sampling can initially find hits quickly, it often leads to exploitation bias, getting trapped in local maxima of the chemical space. AL strategies, particularly those incorporating uncertainty or diversity, achieve a superior balance, resulting in better long-term performance and model robustness [16] [2].
- Optimal Strategy Selection: No single AL strategy is universally best. The choice depends on the primary campaign goal:
In conclusion, the data unequivocally supports the adoption of active learning frameworks for hit discovery. It significantly outperforms traditional random and greedy approaches, enabling researchers to achieve more with fewer resources, thereby accelerating the pace of discovery in fields like drug development and materials science.
In the field of computational drug discovery, active learning (AL) has emerged as a powerful strategy to efficiently navigate vast chemical spaces. However, a persistent challenge remains: balancing the identification of biologically active compounds with the discovery of structurally novel chemical matter. The ability of an AL strategy to identify "hits" (compounds with desired biological activity) that also possess significant scaffold diversity—a process known as scaffold-hopping—is a critical metric of its success and utility. Scaffold-hopping refers to the discovery of structurally novel compounds that retain similar biological activity to a reference molecule, enabling the identification of new lead compounds with potentially improved properties [74] [75]. This guide provides a systematic comparison of active learning strategies, objectively assessing their performance in generating diverse, novel hits through scaffold-hopping, supported by experimental data and detailed methodologies.
Core Principles: Scaffold-Hopping and Active Learning
Defining Scaffold-Hopping in Drug Discovery
Scaffold-hopping is a fundamental concept in lead optimization that involves identifying new chemical scaffolds with similar biological activity to a reference compound while modifying core molecular structures [40]. This process is crucial for overcoming limitations of existing compounds, such as poor pharmacokinetics, toxicity, or intellectual property constraints. Successful scaffold-hopping generates chemically distinct compounds that maintain the desired pharmacological effect, effectively expanding the accessible chemical space for drug development. The computational method AI-AAM (Amino Acid Interaction Mapping) exemplifies this approach by using interactions between a ligand and amino acids as descriptors to find compounds that preserve target interactions despite structural differences [74].
Active Learning Frameworks for Efficient Exploration
Active learning is an iterative machine learning procedure that strategically selects the most informative samples for experimental testing, dramatically reducing the resources required for hit identification [38]. In drug discovery, AL frameworks address the "needle-in-a-haystack" problem of finding active compounds within large chemical libraries. These approaches typically involve multiple cycles where the model uses its current knowledge to prioritize compounds for testing, incorporates the new experimental results, and updates its predictive capabilities [76]. By focusing experimental efforts on the most promising regions of chemical space, AL enables more efficient exploration than random screening or exhaustive testing.
Table 1: Key Active Learning Components for Scaffold-Hopping
| Component | Function | Impact on Scaffold Diversity |
|---|---|---|
| Receptor Ensemble | Multiple protein structures for docking | Increases probability of identifying diverse binders [76] |
| Target-Specific Scoring | Custom scoring for inhibition mechanisms | Better functional prioritization beyond structural similarity [76] |
| Scaffold-Aware Sampling | Strategic focus on underrepresented scaffolds | Actively promotes structural diversity [77] |
| Diversity-Based Selection | Chooses structurally distinct compounds | Directly enhances scaffold diversity in hits [38] |
Comparative Analysis of Active Learning Strategies
Performance Metrics and Experimental Outcomes
Different AL strategies exhibit varying capabilities in identifying diverse hits. A comprehensive investigation of AL for anti-cancer drug screening evaluated multiple approaches, including random, greedy, uncertainty, diversity, and hybrid selection methods [38]. The study demonstrated that most AL strategies significantly outperformed random selection in identifying effective treatments, with certain approaches particularly excelling at early hit identification.
The ScaffAug framework specifically addresses scaffold diversity through a scaffold-aware generative augmentation approach [77]. This method employs a graph diffusion model to generate novel molecules while preserving core scaffold structures from known active compounds, actively combating the structural bias often found in virtual screening datasets.
Table 2: Quantitative Performance Comparison of Active Learning Strategies
| Strategy | Hit Identification Efficiency | Scaffold Diversity | Computational Cost |
|---|---|---|---|
| Random Selection | Baseline | Baseline | Low |
| Uncertainty Sampling | Moderate improvement | Limited improvement | Moderate [38] |
| Diversity-Based | Good improvement | Significant improvement | Moderate [38] |
| Target-Specific Scoring | 200-fold improvement over docking score | Not reported | High (requires MD) [76] |
| Scaffold-Aware (ScaffAug) | Significant improvement | Highest improvement | High (requires generation) [77] |
Experimental Validation of Scaffold-Hopping Success
Experimental validation is crucial for confirming the functional activity of structurally diverse hits identified through AL approaches. In one study applying the AI-AAM scaffold-hopping method, researchers successfully identified XC608 as a novel scaffold with potent inhibitory activity (IC50 = 3.3 nM) against spleen associated tyrosine kinase (SYK), comparable to the reference compound BIIB-057 (IC50 = 3.9 nM) despite significant structural differences [74]. This demonstrates that appropriate computational methods can effectively identify functionally active compounds with diverse scaffolds.
However, the study also revealed potential trade-offs in selectivity. While BIIB-057 selectively inhibited only SYK and PAK5 from 24 kinases tested, XC608 showed inhibition of 14 kinases, indicating reduced selectivity [74]. This highlights the importance of evaluating multiple pharmacological properties beyond primary target activity when assessing novel scaffolds.
Detailed Experimental Protocols
Structure-Based Scaffold-Hopping with AI-AAM
The AI-AAM methodology employs a sophisticated structure-based approach to scaffold-hopping [74]:
- Reference Compound Selection: Choose a known active compound with confirmed target engagement as the reference molecule.
- AAM Descriptor Calculation: Compute Amino Acid Interaction Maps (AAM descriptors) that characterize the interaction patterns between the reference compound and its target protein.
- Virtual Screening: Screen compound libraries to identify molecules with similar AAM descriptors but divergent core structures.
- Similarity Thresholding: Apply AAM similarity scoring (threshold >0.7) to select candidate compounds predicted to maintain similar target interactions.
- Experimental Validation: Test selected compounds for biological activity (e.g., IC50 determination) and selectivity profiling.
This protocol successfully identified structurally diverse SYK inhibitors while maintaining potent inhibition, demonstrating its utility for scaffold-hopping in drug discovery projects [74].
Active Learning with Target-Specific Scoring
The advanced AL framework developed for TMPRSS2 inhibition combines molecular dynamics with active learning for efficient hit identification [76]:
- Receptor Ensemble Generation: Run extensive molecular dynamics simulations (≈100 µs) of the target protein and extract multiple snapshots to capture structural flexibility.
- Initial Library Screening: Dock 1% of the compound library against each structure in the receptor ensemble.
- Target-Specific Scoring: Apply a custom empirical score (h-score) that rewards occlusion of the binding pocket and appropriate interaction geometries rather than relying solely on docking scores.
- Iterative Active Learning Cycle:
- Select top-ranking compounds for experimental testing
- Incorporate experimental results into the training set
- Update the model and repeat the selection process
- Validation: Confirm inhibitor potency through cell-based assays measuring functional inhibition.
This approach reduced the number of compounds requiring experimental testing to less than 20 while successfully identifying BMS-262084 as a potent TMPRSS2 inhibitor (IC50 = 1.82 nM) [76].
Diagram 1: Active Learning Workflow for Hit Discovery
Computational Tools and Databases
Successful implementation of AL strategies for scaffold-hopping requires specialized computational tools and data resources:
Table 3: Essential Research Reagents and Computational Tools
| Resource | Type | Function | Application Example |
|---|---|---|---|
| AnchorQuery | Software | Pharmacophore-based screening of MCR-accessible compounds | Scaffold-hopping for molecular glues [78] |
| ScaffAug Framework | Computational Method | Scaffold-aware generative augmentation | Addressing structural imbalance in VS [77] |
| DrugBank | Database | Contains drug and target information | Reference compound selection [74] |
| DDrare | Database | Drugs for rare diseases clinical trials | Reference compound selection [74] |
| PySCF | Software Library | Quantum chemistry calculations | DFT analysis of inhibitor properties [75] |
| ADMETlab 2.0 | Web Tool | ADMET property prediction | Multi-task graph attention model for property prediction [75] |
| Directory of Useful Decoys, Enhanced (DUD-E) | Database | Designed to benchmark molecular docking | Retrospective study validation [74] |
Experimental Assays for Validation
Critical experimental protocols for validating AL-derived hits include:
- Kinase Inhibition Profiling: Determine IC50 values and selectivity against kinase panels [74]
- Molecular Dynamics Simulations: Analyze protein-ligand complex stability over 100-500 ns trajectories [75] [76]
- Density Functional Theory (DFT) Calculations: Calculate HOMO-LUMO gaps to assess electronic stability (e.g., 4.979 eV for high stability compounds) [75]
- Cell-Based Entry Assays: Evaluate functional inhibition in relevant cellular models (e.g., coronavirus entry blockade in Calu-3 cells) [76]
- NanoBRET Assays: Confirm cellular stabilization of protein-protein interactions in live cells [78]
The comparative analysis of active learning strategies reveals significant differences in their ability to identify novel, diverse hits through scaffold-hopping. While target-specific scoring with receptor ensembles provides remarkable efficiency improvements [76], scaffold-aware approaches like ScaffAug directly address structural diversity [77]. The experimental success of these methods in identifying potent inhibitors with novel scaffolds confirms their value in hit discovery.
Future developments will likely focus on integrating multiple approaches—combining target-aware scoring with scaffold diversity optimization—to further enhance the efficiency and novelty of hit identification. As generative AI methods advance [79] [40], their integration with active learning frameworks promises to accelerate the discovery of structurally diverse, therapeutically relevant compounds, ultimately expanding the accessible chemical space for drug development.
The drug discovery landscape has witnessed an exponential increase in the application of computer-based methodologies toward identifying hit or lead compounds, with virtual screening (VS) now established as a crucial hit identification paradigm alongside traditional high-throughput screening (HTS) and fragment-based screening [80]. However, the transformative potential of these computational approaches hinges entirely on one critical phase: prospective validation. This process bridges the theoretical promise of in-silico predictions with tangible biological confirmation, separating true therapeutic potential from mere digital artifacts. Within modern drug discovery, the validation pathway represents a multifaceted challenge involving strategic experimental design, rigorous counter-screening, and iterative optimization—all while managing limited resources.
The expanding identification of chemical compounds or hits from screening assays has increased the corresponding need for validation and triaging to provide the best leads for therapeutic development [81]. This chapter provides a comprehensive comparison of validation methodologies and frameworks, with particular emphasis on how active learning strategies are reshaping validation workflows. By examining quantitative data, experimental protocols, and strategic implementations across diverse studies, we offer drug development professionals an evidence-based guide to navigating the complex journey from in-silico prediction to experimentally confirmed activity.
Foundational Concepts: Defining Hits and Establishing Validation Criteria
Hit Identification Criteria in Virtual Screening
In traditional HTS, hit selection methods typically include statistical analyses and/or manually set thresholds (e.g., percentage inhibition at a given screening concentration) [80]. However, for virtual screening, consensus on hit identification criteria remains less established. Analysis of published VS studies reveals that only approximately 30% reported a clear, predefined hit cutoff, with significant variation in the biological metrics employed [80].
Table 1: Hit Identification Criteria in Virtual Screening (Analysis of 400+ Studies)
| Hit Calling Metric | Number of Studies | Typical Activity Range |
|---|---|---|
| Percentage Inhibition | 85 | Varies by concentration |
| IC₅₀ | 30 | Low to mid-micromolar |
| EC₅₀ | 4 | Low to mid-micromolar |
| Kᵢ/Kd | 4 | Low to mid-micromolar |
| Other/Not Reported | 298 | Not specified |
Concentration-response endpoints (IC₅₀, EC₅₀, Kᵢ, or Kd) and single-concentration percentage inhibition represent the most common biological metrics for hit cutoffs [80]. The activity spectrum analysis demonstrates that sub-micromolar level cutoffs were rarely used, with the majority of studies employing cutoffs in the low to mid-micromolar range (1-100 μM) [80]. Surprisingly, 56 studies used 100-500 μM and 25 studies used >500 μM as initial activity cutoffs, with approximately one-third of these involving fragment screening (MW <300) [80].
Hit Quality Assessment and Ligand Efficiency
Hit quality assessment involves multifactorial analysis of chemical and physical properties of all compounds meeting predefined hit identification criteria [81]. Key considerations include synthetic tractability, potential reactivity, toxicity, assay interference, and "drug-likeness" [81].
Ligand efficiency (LE) metrics, which normalize experimental activity to molecular size, have become valuable tools for hit assessment. While widely employed in fragment-based screening (typically LE ≥ 0.3 kcal/mol/heavy atom), ligand efficiency has not been well-employed as an activity cutoff method in virtual screening [80]. Notably, none of the analyzed VS reports used ligand efficiency as a hit selection metric, despite its potential value in identifying optimal starting points for medicinal chemistry optimization [80].
In practice, researchers often select hits based on ligand efficiency values to prioritize compounds for optimization. For instance, in a study targeting KPC-2 β-lactamase, N-(3-(1H-tetrazol-5-yl)phenyl)-3-fluorobenzamide was selected as a hit for further optimization based on its favorable ligand efficiency (LE = 0.28 kcal/mol/non-hydrogen atom) and chemistry [82].
Experimental Validation Frameworks: Methodologies and Protocols
The Hit Validation Process
The experimental post-screen hit validation process typically consists of a suite of assays designed to eliminate false positives, confirm activity with the intended target, and establish initial compound ranking [81]. As each screening assay and target is unique, a systematic validation method with well-optimized orthogonal and secondary assays should ideally be established before primary screening completion [81].
Table 2: Key Validation Assays and Their Applications
| Validation Assay Type | Number of Studies Using | Primary Function | Key Features |
|---|---|---|---|
| Secondary Assay | 283 | Confirm primary activity | Different readout or format from primary screen |
| Counter Screen | 116 | Assess selectivity | Against related targets or general interference |
| Binding Validation | 74 | Confirm direct target engagement | Biophysical methods (SPR, NMR, ITC) |
| Cellular Efficacy | Varies | Demonstrate functional activity | In disease-relevant cellular models |
Validation often begins with hit confirmation in the primary assay, frequently using fresh powder or re-synthesized compounds to rule out artifacts [81]. This is followed by counter-screens and orthogonal assays, which may include:
- Specificity Counter-Screens: Testing against related targets or enzymes to establish selectivity [81]
- Interference Assays: Identifying compounds that affect assay components rather than the target [81]
- Cytotoxicity Assays: Ruling out general cellular toxicity as the mechanism of action [81]
Orthogonal Biophysical Assays for Hit Validation
Biophysical methods provide direct evidence of target-ligand interactions not always observable in plate-based spectrophotometric studies [81]. These orthogonal approaches are particularly valuable for confirming binding and mechanism of action.
Surface Plasmon Resonance (SPR) SPR measures biomolecular interactions in real-time without labeling, providing kinetic parameters (KD, kon, koff) in addition to binding confirmation [81]. The technique is highly sensitive for detecting direct binding, though it requires immobilization of one interaction partner.
Nuclear Magnetic Resonance (NMR) NMR serves as a sensitive biophysical technique that provides direct evidence of target-ligand complex formation in solution [81]. While typically not used for primary screening due to cost and throughput limitations, NMR is highly valuable for secondary screening of smaller libraries. Ligand-observed NMR methods, including saturation transfer difference (STD) and WaterLOGSY, can identify binding even for weak affinities (KD up to mM range) [81].
Isothermal Titration Calorimetry (ITC) ITC directly measures the heat change during binding, providing a complete thermodynamic profile (KD, ΔH, ΔS, stoichiometry) without requiring labeling or immobilization [81]. This method is particularly valuable for confirming binding and understanding the driving forces behind molecular interactions.
Thermal Shift Assays Also known as differential scanning fluorimetry (DSF), thermal shift assays monitor protein thermal stability changes upon ligand binding, typically through fluorescent dyes that bind hydrophobic regions exposed during denaturation [81]. This method provides a medium-throughput approach to confirm direct binding to the target protein.
Active Learning Strategies for Efficient Hit Discovery
Active Learning Framework in Drug Discovery
Active learning represents an iterative machine learning procedure where the model learning process is divided into iterations, with each iteration selecting a new group of samples based on a designed strategy to add to the training dataset [16]. This approach is particularly valuable in biomedical applications where experimentation costs are high, as it can help identify effective treatments earlier in the process, saving substantial time and resources [16].
In practice, active learning frameworks for drug discovery typically include:
- Initial Model Training: Using available data to create a baseline prediction model
- Iterative Batch Selection: Selecting the most informative experiments based on current model predictions
- Experimental Validation: Conducting wet-lab experiments on selected compounds or conditions
- Model Refinement: Updating the prediction model with new experimental data
- Stopping Criteria: Determining when sufficient hits have been identified or model performance has plateaued
Active Learning Sampling Strategies
Multiple sampling techniques have been investigated for active learning in drug discovery, each with distinct advantages for different experimental scenarios:
Uncertainty Sampling This approach selects samples where the model exhibits highest uncertainty in predictions, targeting regions of the chemical space where additional data would most improve model performance [16]. Uncertainty is typically measured using prediction confidence scores or entropy measures.
Diversity Sampling Diversity-based approaches select samples that maximize coverage of the chemical space, ensuring representation across diverse molecular structures [16]. This method helps prevent over-exploration of limited regions and supports broader structure-activity relationship understanding.
Hybrid Approaches Combining uncertainty and diversity sampling often yields superior results by balancing exploration of uncertain regions with broad chemical space coverage [16] [17]. Hybrid methods may also incorporate exploitation elements to focus on regions already showing promising activity.
Table 3: Performance Comparison of Active Learning Strategies in Drug Discovery
| Sampling Strategy | Hit Identification Efficiency | Model Improvement | Best Use Cases |
|---|---|---|---|
| Random Sampling | Baseline | Baseline | Control comparison |
| Uncertainty Sampling | Moderate improvement | Significant improvement | Model refinement focus |
| Diversity Sampling | Good improvement | Moderate improvement | Diverse chemical space exploration |
| Greedy Sampling | Limited improvement | Limited improvement | Known promising regions |
| Hybrid Approaches | Best performance | Best performance | Balanced resource allocation |
Performance Benchmarks and Efficiency Gains
Active learning strategies demonstrate significant efficiency improvements in hit discovery. Studies show that active learning can discover 60% of synergistic drug pairs by exploring only 10% of the combinatorial space, representing an 82% reduction in experimental requirements [17]. Similarly, research on anti-cancer drug response prediction shows active learning approaches identify hits significantly earlier than random or greedy sampling methods [16].
The synergy yield ratio is observed to be even higher with smaller batch sizes, where dynamic tuning of the exploration-exploitation strategy can further enhance performance [17]. This efficiency makes comprehensive screening campaigns feasible for biological laboratories with limited resources.
Case Study: KPC-2 β-Lactamase Inhibitor Discovery
Virtual Screening and Hit Identification
A comprehensive example of the prospective validation process comes from a study targeting KPC-2 β-lactamase, a carbapenemase that poses a serious health threat due to its resistance to last-resort carbapenem antibiotics [82]. Researchers performed structure-based in-silico screening of commercially available compounds for non-β-lactam KPC-2 inhibitors using a hierarchical screening cascade.
The virtual screening approach incorporated:
- Binding Site Analysis: Using PoSSuM to identify similar binding sites and shared ligand interactions [82]
- Pharmacophore Modeling: Defining features based on KPC-2 protein structure (PDB 3RXW) and ligand OJ6 of CTX-M-9 β-lactamase [82]
- Structure-Based Screening: Selecting compounds with complementary interactions to key residues including Thr237, Thr235, Ser130, and Trp105 [82]
From this process, 32 commercially available high-scoring, fragment-like hits were selected for in-vitro validation [82].
Experimental Validation and Hit Confirmation
The selected candidates underwent comprehensive experimental validation to confirm activity against isolated recombinant KPC-2 [82]. This process identified several active compounds, with N-(3-(1H-tetrazol-5-yl)phenyl)-3-fluorobenzamide (compound 11a) and a benzothiazole derivative (compound 9a) showing the highest activity against KPC-2 [82].
Mechanism of action studies confirmed these compounds behaved as competitive inhibitors of the target carbapenemase [82]. Based on its promising ligand efficiency (LE = 0.28 kcal/mol/non-hydrogen atom) and favorable chemistry, compound 11a was selected for further optimization [82].
Translation to Cellular Activity
Following biochemical validation, the most promising compounds were evaluated against clinical strains overexpressing KPC-2 [82]. This critical step assessed whether the biochemical inhibition translated to functional activity in biologically relevant systems. The study demonstrated that the most promising compound reduced the MIC (Minimum Inhibitory Concentration) of the β-lactam antibiotic meropenem by four-fold, confirming the potential for combination therapy [82].
Research Reagent Solutions Toolkit
Table 4: Essential Research Reagents and Platforms for Prospective Validation
| Reagent/Platform | Primary Function | Application in Validation |
|---|---|---|
| Recombinant Proteins | Target protein for biochemical assays | Primary screening and mechanism studies |
| Cell-Based Assay Systems | Cellular activity assessment | Translation from biochemical to cellular efficacy |
| Surface Plasmon Resonance | Label-free binding kinetics | Confirm direct binding and measure affinity |
| NMR Spectroscopy | Structural binding information | Confirm binding and characterize interactions |
| Isothermal Titration Calorimetry | Thermodynamic binding profile | Understand binding driving forces |
| Clinical Strains | Biologically relevant models | Assess activity in disease-relevant contexts |
| Chemical Libraries | Diverse compound sources | Starting points for screening campaigns |
The journey from in-silico prediction to experimentally confirmed activity represents a critical pathway in modern drug discovery. Through systematic analysis of validation methodologies and emerging approaches like active learning, we can delineate best practices for efficient hit confirmation. The integration of computational predictions with experimental validation creates a synergistic framework that accelerates the identification of promising therapeutic candidates while minimizing resource expenditure.
As active learning strategies continue to evolve, their ability to navigate complex experimental spaces with unprecedented efficiency promises to reshape hit discovery workflows. By strategically selecting the most informative experiments and dynamically refining prediction models, researchers can overcome the traditional limitations of large combinatorial spaces and rare synergistic phenomena. This integrated approach, combining computational power with experimental rigor, represents the future of efficient therapeutic development, potentially delivering novel treatments to patients faster and more cost-effectively.
In the field of hit discovery research, the high cost and time demands of High-Throughput Screening (HTS) for large compound libraries present a significant challenge. Active Learning (AL), a subfield of machine learning, has emerged as a powerful solution by enabling iterative, intelligent screening. This guide provides an objective comparison of prominent AL strategies, focusing on their experimental performance, methodologies, and applicability to help researchers select the optimal approach for their projects.
The table below summarizes the core objectives and experimentally measured outcomes of two distinct AL approaches applied in prospective drug discovery campaigns.
Table 1: Performance Comparison of Active Learning Strategies in Prospective Studies
| Active Learning Strategy | Core Screening Methodology | Reported Experimental Outcome | Key Performance Metric |
|---|---|---|---|
| Balanced-Ranking (ChemScreener) [12] | Multi-task AL with ensemble uncertainty for exploration and predicted activity for exploitation. | Increased hit rates from a primary HTS baseline of 0.49% to an average of 5.91% (range: 3-10%). Identified 104 hits from 1,760 compounds tested. | Hit Rate Enrichment |
| ML-Assisted Iterative HTS [57] | Machine learning-guided iterative screening in sequential batches. | Recovered 43.3% of all primary actives identified in a parallel full HTS by screening only 5.9% of a 2-million-compound library. | Screening Efficiency & Hit Recovery |
Detailed Experimental Protocols and Workflows
The ChemScreener Workflow with Balanced-Ranking Acquisition
This protocol was designed for early hit discovery across large, diverse chemical spaces, starting from limited data [12].
- Initial Model Training: Begin with a modestly sized, structurally diverse subset of compounds with associated assay data. Train a multi-task deep learning ensemble model on this initial dataset.
- Balanced-Ranking Acquisition: Apply the Balanced-Ranking strategy to prioritize compounds for the next screening batch. This involves:
- Using the ensemble's predictive mean to rank compounds by predicted activity (exploitation).
- Using the ensemble's predictive uncertainty (e.g., standard deviation across models) to rank compounds by informativeness (exploration).
- Combining these two ranks to generate a final balanced priority list.
- Iterative Screening & Model Refinement:
- Screen the top-priority compounds (e.g., 1,760 compounds) from the acquisition step in a single-dose assay.
- Incorporate the new experimental results into the training dataset.
- Retrain the AL model with the updated data.
- Repeat steps 2 and 3 for multiple cycles (e.g., 5 cycles).
- Hit Validation & Progression:
- Consolidate hits from all cycles and retest them together with close analogs.
- Cluster validated hits to identify distinct chemical series.
- Advance promising clusters to dose-response assays (e.g., IC50 determination) and orthogonal binding assays (e.g., Differential Scanning Fluorimetry - DSF) for confirmation.
ML-Assisted Iterative High-Throughput Screening
This protocol was prospectively validated for efficient detection of drug discovery starting points in a large-scale project targeting Salt-Inducible Kinase 2 (SIK2) [57].
- Initialization: Conduct a small, initial HTS batch to generate a foundational dataset for the machine learning model.
- Predictive Modeling: Train the ML model (e.g., a classifier to identify active compounds) on all data collected so far.
- Compound Prioritization: Use the trained model to predict activity for all remaining unscreened compounds in the multi-million-member library. Select the next batch of compounds with the highest predicted probability of activity.
- Iterative Loop: Experimentally screen the selected batch of compounds. Add the results to the training pool and repeat steps 2 and 3 until the desired number of compounds is screened or a target number of hits is identified.
- Retrospective Analysis: Upon completion, compare the hit recovery and chemical diversity of the ML-guided screening campaign against a full, non-iterative HTS of the entire library conducted in parallel.
Active Learning Screening Workflow
The diagram below illustrates the core iterative loop shared by advanced AL methods in hit discovery.
The Scientist's Toolkit: Essential Research Reagents and Materials
The following table lists key reagents and computational tools used in the featured AL-driven hit discovery campaigns.
Table 2: Key Research Reagent Solutions for AL-Driven Hit Discovery
| Item / Solution | Function in AL Workflow | Specific Example / Note |
|---|---|---|
| Target-Specific Biochemical Assay | Generates the primary activity data used to train and guide the AL model. | HTRF assays for protein-target interaction [12]; mass spectrometry-based assay for kinase activity [57]. |
| Diverse Compound Library | The chemical space explored by the AL algorithm for hit discovery. | Large libraries (e.g., 2 million compounds) are typical, but AL aims to screen only a small, strategic fraction [57]. |
| Orthogonal Binding Assay | Validates that hits identified in the primary screen are true binders and not assay artifacts. | Differential Scanning Fluorimetry (DSF) was used to confirm binding to the target protein, WDR5 [12]. |
| Gaussian Process (GP) Emulator / Deep Learning Ensemble | Acts as the statistical or machine learning "emulator" that models the relationship between compound features and activity, predicting outcomes for unscreened compounds. | Easy-to-interpret Gaussian Process (EzGP) models handle mixed data inputs [83]; Deep learning ensembles manage uncertainty for the Balanced-Ranking strategy [12]. |
| Automated Screening Platform | Enables the high-throughput experimental testing of compounds prioritized by the AL model in each cycle. | Robotic liquid handling and plate readers are essential for efficient iteration. |
The prospective validations of both the Balanced-Ranking and ML-Assisted Iterative HTS strategies demonstrate that active learning is no longer just a theoretical improvement but a practical tool that can dramatically enhance the efficiency and output of hit discovery campaigns. The choice between strategies hinges on the primary project constraint: Balanced-Ranking excels at maximizing hit rate and scaffold diversity from a limited number of tests, while ML-Assisted Iterative HTS proves highly effective at recovering the majority of hits from a very large library with minimal screening effort. Integrating these data-driven approaches allows research teams to de-risk the early discovery process and accelerate the identification of viable starting points for drug development.
Conclusion
The consolidated evidence from recent studies firmly establishes active learning as a transformative methodology for hit discovery. By moving beyond one-shot screening to an iterative, data-driven process, AL consistently demonstrates a superior ability to identify active compounds with significantly enriched hit rates—often increasing them from less than 1% to over 5%—while exploring a fraction of the chemical space. The future of AL lies in its deeper integration with generative AI for de novo design, improved handling of multi-parameter optimization, and application in complex areas like synergistic drug combination discovery. For biomedical research, the widespread adoption of these strategies promises to drastically reduce the time and cost of bringing new therapeutics to patients, marking a pivotal shift towards more efficient and intelligent drug discovery pipelines.
References
- 1. Accelerating materials discovery through active learning [https://www.the-innovation.org/article/doi/10.59717/j.xinn-inform.2025.100013]
- 2. A comprehensive benchmark of active learning strategies ... [https://www.nature.com/articles/s41598-025-24613-4]
- 3. Active learning high coverage sets of complementary ... [https://pubs.rsc.org/en/content/articlelanding/2025/dd/d4dd00365a]
- 4. Leading AI-Driven Drug Discovery Platforms [https://www.sciencedirect.com/science/article/abs/pii/S0031699725075118]
- 5. 5 Best Gen AI Tools For Drug Discovery (2025) [https://www.consainsights.com/blogs/health/gen-ai-tools-for-drug-discovery]
- 6. Inside ELRIG's Drug Discovery 2025 [https://www.drugtargetreview.com/article/190237/inside-elrigs-drug-discovery-2025/]
- 7. High Throughput Screening Assays for Drug Discovery [https://bellbrooklabs.com/high-throughput-screening-assays-drug-discovery/?srsltid=AfmBOopAbPBeD4AKSSq7KF82paO5VyTMa-dpl9yrZIjnfjdZRPvpHSrO]
- 8. AI-Driven Iterative Screening for Hit Finding - PubMed Central [https://pmc.ncbi.nlm.nih.gov/articles/PMC7838329/]
- 9. AI is a viable alternative to high throughput screening [https://www.nature.com/articles/s41598-024-54655-z]
- 10. A Comparison of Machine Learning Approaches for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11235188/]
- 11. Optimizing drug design by merging generative AI with a ... [https://www.nature.com/articles/s42004-025-01635-7]
- 12. ChemScreener: an Active Learning Enabled Hit Discovery ... [https://chemrxiv.org/engage/chemrxiv/article-details/690d5c66113cc7cfffab2473]
- 13. The present state and challenges of active learning in drug ... [https://www.sciencedirect.com/science/article/abs/pii/S1359644624001107]
- 14. Active Learning Machine Learning: How It Reduces ... [https://labelyourdata.com/articles/active-learning-machine-learning]
- 15. Active learning framework leveraging transcriptomics ... [https://pubmed.ncbi.nlm.nih.gov/41129612/]
- 16. A Comprehensive Investigation of Active Learning ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10854925/]
- 17. A guide for active learning in synergistic drug discovery [https://www.nature.com/articles/s41598-025-85600-3]
- 18. MCML - How to Measure Uncertainty in ... [https://mcml.ai/publications/nsh22/]
- 19. [2307.02719] Understanding Uncertainty Sampling [https://arxiv.org/abs/2307.02719]
- 20. Evidential deep learning-based drug-target interaction ... [https://www.nature.com/articles/s41467-025-62235-6]
- 21. Enhanced uncertainty sampling with category information for ... [https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0327694]
- 22. Calibrated Uncertainty Sampling for Active Learning [https://arxiv.org/abs/2510.03162]
- 23. Enhanced Thompson sampling by roulette wheel selection for ... [https://jcheminf.biomedcentral.com/articles/10.1186/s13321-025-01105-1]
- 24. The Future of Drug Discovery: 2025 as the Inflection Year ... [https://modelmedicines.com/newsroom/the-future-of-drug-discovery-2025-as-the-inflection-year-for-hybrid-ai-and-quantum-computing]
- 25. Selecting molecules with diverse structures and properties ... [https://www.nature.com/articles/s41598-022-04967-9]
- 26. Growth vs. Diversity: A Time-Evolution Analysis of the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11870478/]
- 27. A Comprehensive Investigation of Active Learning Strategies ... [https://pubmed.ncbi.nlm.nih.gov/38339281/]
- 28. Performance Metrics in Data Science: A Comprehensive ... [https://medium.com/@data-overload/performance-metrics-in-data-science-a-comprehensive-guide-7b036e0e18e4]
- 29. A comprehensive benchmark of active learning strategies ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12550062/]
- 30. AI-driven drug discovery using a context-aware hybrid ... [https://www.nature.com/articles/s41598-025-19593-4]
- 31. Phenotypic and targeted drug discovery in immune therapeutics [https://pubs.rsc.org/en/content/articlehtml/2025/ra/d5ra03914b]
- 32. Discovery of an exquisitely selective WDR5 chemical probe ... [https://pubs.rsc.org/en/content/articlehtml/2025/cb/d5cb00109a]
- 33. An Ensemble-Based Active Learning Workflow for Hit ... [https://www.linkedin.com/posts/fanli_chemscreener-activity-7395475672472797184-8AmF]
- 34. Overlapping Characteristics of Weak Interactions of Two ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10922662/]
- 35. Insilico Medicine and collaborator announce discovery of ... [https://www.eurekalert.org/news-releases/1085461]
- 36. De novo drug design through artificial intelligence [https://www.frontiersin.org/journals/hematology/articles/10.3389/frhem.2024.1305741/full]
- 37. Advances in De Novo Drug Design: From Conventional to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7915729/]
- 38. A Comprehensive Investigation of Active Learning ... [https://www.mdpi.com/2072-6694/16/3/530]
- 39. Prospective de novo drug design with deep interactome ... [https://www.nature.com/articles/s41467-024-47613-w]
- 40. Generative artificial intelligence based models optimization ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12323263/]
- 41. Deep Generative AI for Multi-Target Therapeutic Design [https://www.mdpi.com/1422-0067/26/23/11443]
- 42. Hit Expansion of a Noncovalent SARS-CoV-2 Main ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9003389/]
- 43. Structure of Mpro from SARS-CoV-2 and discovery of its ... [https://www.nature.com/articles/s41586-020-2223-y]
- 44. Targeting SARS-CoV-2 main protease [https://pmc.ncbi.nlm.nih.gov/articles/PMC11662536/]
- 45. Review Article Machine learning in antiviral drug design [https://www.sciencedirect.com/science/article/pii/S0968089625003670]
- 46. Active learning driven prioritisation of compounds from on ... [https://pubs.rsc.org/en/content/articlehtml/2025/dd/d4dd00343h]
- 47. Active learning driven prioritisation of compounds from on- ... [https://www.sciencedirect.com/org/science/article/pii/S2635098X25000038]
- 48. A machine learning (ML) driven Web-App for bioactivity ... [https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0287179]
- 49. Machine Learning for Multi-Target Drug Discovery [https://pmc.ncbi.nlm.nih.gov/articles/PMC12473769/]
- 50. Active Learning-Guided Seq2Seq Variational Autoencoder ... [https://arxiv.org/html/2506.15309v1]
- 51. Exploration-Exploitation Trade-Offs [https://www.emergentmind.com/topics/exploration-exploitation-trade-offs]
- 52. Active Learning Statistics: Benefits for Education & Training ... [https://www.engageli.com/blog/active-learning-statistics-2025]
- 53. Unlocking hit-to-lead efficiency through AI and automation [https://www.ddw-online.com/unlocking-hit-to-lead-efficiency-through-ai-and-automation-36007-202508/]
- 54. Hit to Lead Assays: Accelerating Early Drug Discovery [https://bellbrooklabs.com/hit-to-lead-assays/?srsltid=AfmBOoq3jMS76CDXZkwVPi4aBEHCvUGxdCA090iFBP6EyUF4GCTzoZFb]
- 55. Top 5 Drug Discovery Trends 2025 Driving Breakthroughs [https://www.pelagobio.com/cetsa-drug-discovery-resources/blog/drug-discovery-trends-2025/]
- 56. The essential roles of chemistry in high-throughput screening ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4465542/]
- 57. Machine Learning-Assisted Iterative Screening for Efficient ... [https://pubmed.ncbi.nlm.nih.gov/40658043/]
- 58. Acoustic Ejection Mass Spectrometry: A Fully Automatable ... [https://www.sciencedirect.com/science/article/pii/S2472555222067478]
- 59. Deep active learning: A reality check [https://www.sciencedirect.com/science/article/abs/pii/S0167865525002697]
- 60. Accelerating ML Model Training with Active Learning ... [https://dagshub.com/blog/accelerating-ml-model-training-with-active-learning-techniques/]
- 61. Deep active learning with high structural discriminability for ... [https://www.nature.com/articles/s42003-024-06758-6]
- 62. Vanderbilt scientist tackles key roadblock for AI in drug discovery [https://medschool.vanderbilt.edu/basic-sciences/2025/10/16/vanderbilt-scientist-tackles-key-roadblock-for-ai-in-drug-discovery/]
- 63. AI in Drug Discovery 2025 - Schrödinger [https://www.schrodinger.com/life-science/resources/event/ai-in-drug-discovery-2025/]
- 64. MIT scientists debut a generative AI model that could ... [https://news.mit.edu/2025/mit-scientists-debut-generative-ai-model-that-could-create-molecules-addressing-hard-to-treat-diseases-1125]
- 65. Artificial intelligence revolution in drug discovery [https://www.sciencedirect.com/science/article/pii/S037851732500626X]
- 66. Integrating artificial intelligence in drug discovery and early ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11909971/]
- 67. Transformative Role of Artificial Intelligence in Drug ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12406033/]
- 68. Active Learning for SAT Solver Benchmarking [https://link.springer.com/article/10.1007/s10817-025-09729-6]
- 69. 12 Important Model Evaluation Metrics for Machine ... [https://www.analyticsvidhya.com/blog/2019/08/11-important-model-evaluation-error-metrics/]
- 70. 9 Accuracy Metrics for ML Engineers to Evaluate AI Model [https://galileo.ai/blog/accuracy-metrics-ai-evaluation]
- 71. How to Check the Accuracy of Your Machine Learning ... [https://www.deepchecks.com/how-to-check-the-accuracy-of-your-machine-learning-model/]
- 72. Efficient discovery of responses of proteins to compounds ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4030446/]
- 73. Large-scale Pretraining Improves Sample Efficiency of ... [https://arxiv.org/html/2309.11687]
- 74. A novel in silico scaffold-hopping method for drug ... [https://www.nature.com/articles/s41598-023-46648-1]
- 75. Structural stability-guided scaffold hopping and computational ... [https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0332798]
- 76. Simulations and active learning enable efficient ... [https://www.nature.com/articles/s41467-025-62139-5]
- 77. Scaffold-Aware Generative Augmentation and Reranking ... [https://arxiv.org/html/2510.16306v1]
- 78. Scaffold-hopping for molecular glues targeting the 14-3-3/ ... [https://www.nature.com/articles/s41467-025-61176-4]
- 79. Generative Deep Learning for de Novo Drug Design A ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12308806/]
- 80. Hit Identification and Optimization in Virtual Screening [https://pmc.ncbi.nlm.nih.gov/articles/PMC3772997/]
- 81. Hit-to-Lead: Hit Validation and Assessment [https://www.sciencedirect.com/science/article/abs/pii/S0076687918303859]
- 82. In silico identification and experimental validation of hits active ... [https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0203241]
- 83. Active Learning of Computer Experiment with both ... [https://arxiv.org/html/2504.13441v2]