Generative AI vs. Active Learning: A Strategic Comparison for Accelerating Drug Discovery
This article provides a comprehensive comparison of Generative AI and Active Learning, two pivotal machine learning paradigms transforming pharmaceutical R&D.
Generative AI vs. Active Learning: A Strategic Comparison for Accelerating Drug Discovery
Abstract
This article provides a comprehensive comparison of Generative AI and Active Learning, two pivotal machine learning paradigms transforming pharmaceutical R&D. Tailored for researchers and drug development professionals, it explores the foundational principles of each approach, details their methodological applications in tasks from de novo molecular design to lead optimization, and addresses key implementation challenges. By presenting real-world case studies and validation metrics, it offers a strategic framework for selecting and integrating these technologies to enhance efficiency, reduce costs, and improve success rates in the drug discovery pipeline.
Core Concepts: Demystifying Generative AI and Active Learning in a Pharmaceutical Context
The pursuit of knowledge and innovation is undergoing a fundamental transformation, moving from serendipitous "Discovery by Luck" toward systematic "Discovery by Design." This paradigm shift is largely driven by the emergence of sophisticated computational approaches, particularly generative artificial intelligence (AI) and active learning methodologies. In scientific domains, especially drug development, this transition represents a move away from reliance on chance observations toward engineered, predictable discovery processes powered by algorithms that can explore complex spaces with unprecedented efficiency.
This guide provides an objective comparison of two leading computational approaches—generative AI and active learning—that are enabling this transition. We examine their performance characteristics, experimental protocols, and practical implementations to help researchers, scientists, and drug development professionals make informed decisions about integrating these technologies into their discovery workflows.
Quantitative Performance Comparison
Extensive research has quantified the performance characteristics of both generative AI and active learning approaches across multiple dimensions. The table below summarizes key findings from controlled studies and implementation data.
Table 1: Performance Metrics of Generative AI vs. Active Learning Approaches
| Performance Metric | Generative AI | Active Learning | Data Sources |
|---|---|---|---|
| Learning Gains/Effectiveness | Over double the median learning gains compared to in-class active learning [1] | 54% higher test scores than traditional passive learning [2] [3] | Randomized controlled trials [1] [3] |
| Time Efficiency | Learned significantly more in less time; median 49 minutes vs. 60 minutes for same material [1] | 13 times more learner talk time and 16 times more non-verbal engagement [3] | Time-on-task measurements [1] [3] |
| User Engagement & Motivation | Significantly higher engagement and motivation ratings [1] | 75% of students feel more motivated compared to 30% in traditional environments [2] | Likert-scale surveys and engagement tracking [1] [2] |
| Implementation Scale | Highly scalable; addresses limitations of one-teacher-to-many-students model [1] | 62.7% participation rate vs. 5% in lecture formats [3] | Large-scale educational studies [1] [3] |
| Resource Efficiency | Drastic reduction in inference costs (over 280-fold for equivalent performance) [4] | Enables better model performance with fewer labeled examples [5] | Market analysis and machine learning benchmarks [4] [5] |
| Risks & Limitations | Potential for over-reliance, decreased cognitive engagement, and superficial learning [6] | Requires careful implementation to overcome student perception gaps [3] | Controlled experiments and observational studies [6] [3] |
Experimental Protocols and Methodologies
Generative AI in Educational Intervention
A recent randomized controlled trial at Harvard University provides a robust protocol for evaluating generative AI effectiveness [1].
Population and Setting: The study involved 194 undergraduate students in a physics course, broadly representative of diverse institutional populations.
Experimental Design: A crossover design was implemented where students were divided into two groups. Each group experienced both teaching methodologies in consecutive weeks:
- Week 1: Group 1 used AI-supported lesson at home; Group 2 participated in active learning lesson in class
- Week 2: Conditions were reversed between groups
AI Intervention: The custom AI tutor was designed with specific pedagogical principles:
- Targeted prompt engineering to facilitate active learning
- Cognitive load management techniques
- Growth mindset promotion
- Scaffolding of content with timely, accurate feedback
- Self-paced progression through material
Measurement Instruments:
- Pre-test and post-test assessments for baseline knowledge and content mastery
- 5-point Likert scale surveys measuring engagement, enjoyment, motivation, and growth mindset
- Platform analytics tracking time-on-task for AI group
- Statistical analysis using two-sample rank-sum (Mann-Whitney) tests and linear regression models with multiple controls
Key Findings: The AI group demonstrated significantly higher post-test scores (median = 4.5 vs. 3.5) with less median time on task (49 minutes vs. 60 minutes), while reporting higher engagement and motivation [1].
Active Learning Implementation Framework
Research from Engageli and other institutions establishes a clear protocol for active learning implementation [3].
Setting and Participants: Studies span K-12, higher education, and corporate training environments with diverse participant populations.
Intervention Components:
- Transformation of passive lecture-based learning into participatory experiences
- Interactive techniques including discussions, polls, small-group activities
- Real-time adjustments based on student engagement levels
- Digital and collaborative elements to enhance participation
Measurement Approach:
- Comparison of test scores between active learning and traditional lecture formats
- Tracking of verbal and non-verbal engagement metrics
- Assessment of knowledge retention over time
- Measurement of participation rates and attendance patterns
- Longitudinal tracking of performance in follow-up courses
Key Outcomes: Active learning environments generated 13 times more learner talk time, 16 times higher non-verbal engagement, and 54% higher test scores compared to traditional lectures [3].
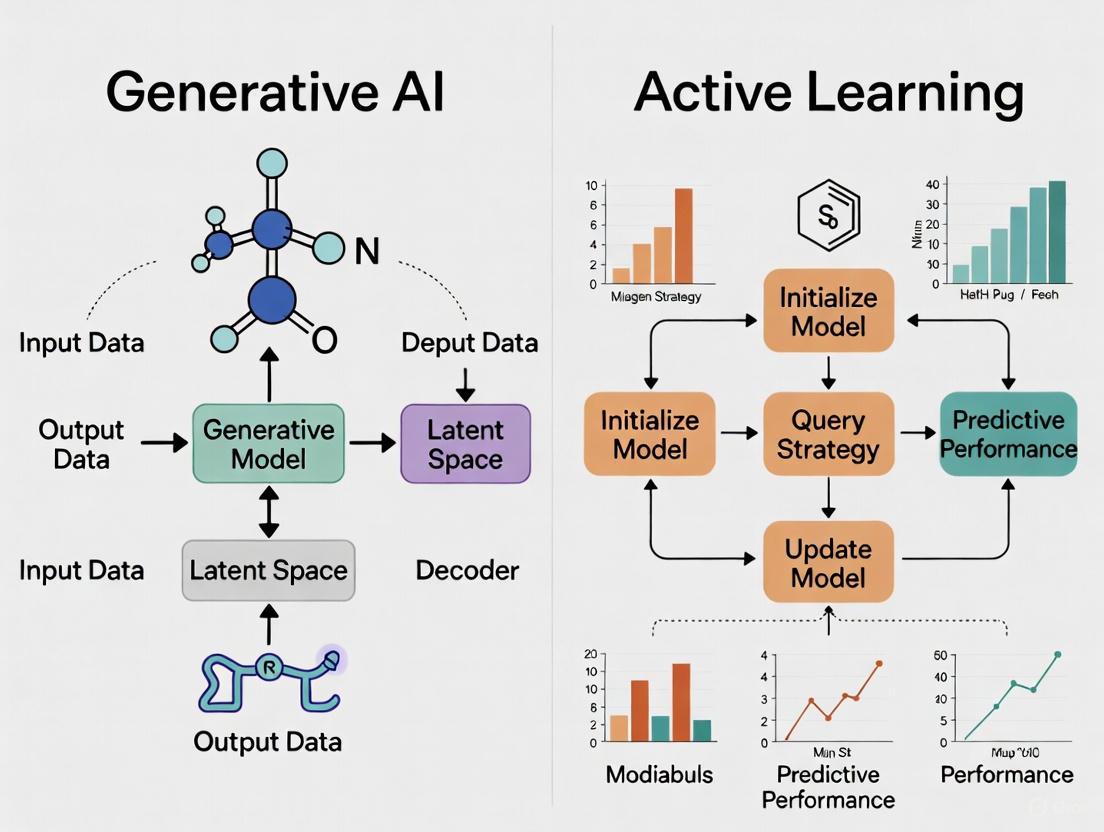
Visualization of Methodological Frameworks
Generative AI Tutoring Workflow
Active Learning Implementation Cycle
The Researcher's Toolkit: Essential Implementation Components
Successful implementation of either generative AI or active learning approaches requires specific "research reagent solutions" – the essential components that enable effective deployment.
Table 2: Essential Research Reagents for AI and Active Learning Implementation
| Component Category | Specific Tools & Solutions | Function & Purpose | Implementation Examples |
|---|---|---|---|
| Computational Infrastructure | GPUs/TPUs, Cloud Computing (AWS, Google Cloud, Azure), High-performance Hardware [7] | Handles large-scale parallel computations for model training and inference | Enables processing of complex AI algorithms and real-time interactions [7] |
| AI Models & Frameworks | Transformer Models (GPT), GANs, VAEs, PyTorch, TensorFlow, Keras [8] [7] | Provides foundation for generative AI capabilities and model development | Custom AI tutors for personalized learning [1] |
| Data Management Systems | Data Collection Tools, Preprocessing Pipelines, Annotation Platforms [7] [5] | Ensures quality, diversity, and appropriate labeling of training data | Critical for both AI training and active learning selection strategies [5] |
| Interaction & Engagement Platforms | Digital Polling, Chat Systems, Collaboration Tools, Video Conferencing with Engagement Features [3] | Facilitates real-time interaction and participation measurement | Enables 13x more learner talk time and 16x more non-verbal engagement [3] |
| Assessment & Analytics | Learning Management Systems, Analytics Dashboards, A/B Testing Frameworks [1] | Tracks effectiveness, engagement metrics, and learning outcomes | Measures learning gains, time-on-task, and knowledge retention [1] |
| Pedagogical Design Components | Prompt Engineering Templates, Cognitive Load Management, Scaffolding Sequences [1] | Ensures educational effectiveness and appropriate challenge levels | Key to successful AI tutor design and active learning activity structure [1] |
Analysis and Interpretation of Findings
The comparative data reveals distinct strengths and applications for each paradigm. Generative AI demonstrates remarkable efficiency in personalized knowledge transfer, enabling students to learn more in less time while providing scalability that addresses fundamental limitations of traditional education models [1]. This comes with the caveat that poorly implemented AI may foster over-reliance and reduce cognitive engagement, potentially undermining long-term knowledge retention [6].
Active learning approaches show consistent advantages in fostering deeper engagement and developing critical thinking skills through social learning and collaboration [3]. The methodology effectively addresses the "perception gap" where students may feel they learn less despite demonstrating significantly better actual retention and understanding [3].
In the context of "Discovery by Design," both approaches offer pathways toward more systematic innovation. Generative AI excels in exploring vast solution spaces and generating novel possibilities, while active learning provides frameworks for collaborative refinement and validation of discoveries. The emerging research suggests that hybrid approaches—leveraging the strengths of both paradigms—may represent the most promising direction for advanced research and development workflows.
The transition from "Discovery by Luck" to "Discovery by Design" is not merely about adopting new technologies, but about fundamentally reengineering how we approach knowledge creation and innovation. Both generative AI and active learning offer powerful methodologies for this transformation, each with distinct performance characteristics and implementation requirements.
Generative AI provides unprecedented scalability and personalization in exploration and content generation, while active learning creates environments conducive to deep engagement and collaborative problem-solving. The experimental evidence suggests context-appropriate application of these approaches—either individually or in combination—can significantly accelerate discovery processes across scientific domains.
For researchers and drug development professionals, the imperative is clear: deliberate design of discovery workflows, informed by robust experimental data and implementing appropriate technological solutions, can systematically enhance innovation outcomes. The paradigm of chance observations is giving way to engineered discovery processes, with generative AI and active learning serving as foundational methodologies in this transformative era.
Eroom's law—the observation that drug discovery is becoming slower and more expensive over time, despite technological improvements—presents a critical economic and innovative challenge for the pharmaceutical industry [9]. This review examines the potential of artificial intelligence (AI) to reverse this trend by comparing two dominant computational approaches: generative AI and active learning. We synthesize current data from 2024-2025, detailing experimental protocols, performance metrics, and practical workflows to guide researchers and drug development professionals in leveraging these technologies to address the productivity crisis.
Understanding Eroom's Law and the "Better than the Beatles" Problem
Eroom's law describes the adverse trend where the inflation-adjusted cost of developing a new drug roughly doubles every nine years [9]. This decline in R&D efficiency is attributed to several factors, including the "better than the Beatles" problem, where new drugs must demonstrate incremental benefit over already highly effective treatments, thereby requiring larger and more expensive clinical trials [9]. Other key causes include increasingly cautious regulators, inefficient resource allocation, and a bias towards basic research methods that often fail in clinical trials due to the complexity of whole organisms [9]. The field of cardiovascular therapeutics (CVD) exemplifies this crisis, with 33% fewer CVD therapeutics approved in the 2000s compared to the previous decade [10].
AI-driven discovery platforms claim to drastically shorten early-stage R&D timelines and cut costs by using machine learning and generative models to accelerate tasks traditionally reliant on cumbersome trial-and-error [11]. This review evaluates the evidence for this claim by comparing the capabilities of generative AI and active learning.
Comparative Analysis: Generative AI vs. Active Learning
The table below summarizes the core characteristics, performance data, and applications of these two approaches based on recent literature.
Table 1: Comparative Analysis of Generative AI and Active Learning in Drug Discovery
| Feature | Generative AI | Active Learning |
|---|---|---|
| Core Paradigm | "Describe first then design": Creates novel molecular structures from scratch [12]. | "Design first then predict": Iteratively selects informative candidates from a library for evaluation [12]. |
| Primary Objective | De novo design of novel, drug-like molecules with optimized properties [11] [12]. | Efficiently navigate vast chemical spaces to identify high-potential hits with minimal resource use [12]. |
| Key Strengths | - Explores novel chemical space beyond known scaffolds.- Can generate molecules tailored to specific target profiles.- High speed in ideation [13] [12]. | - High efficiency in resource-constrained settings.- Reduces number of costly assays or simulations.- Improves predictive model accuracy with each cycle [12]. |
| Reported Performance | - ISM001-055: From target to Phase I in 18 months (Insilico Medicine) [11].- CDK2 Inhibitors: 8 out of 9 synthesized molecules showed in vitro activity [12].- Exscientia: ~70% faster design cycles with 10x fewer synthesized compounds [11]. | - Achieves 5–10x higher hit rates than random selection in drug combination searches [12].- Significantly reduces number of docking or ADMET assays needed to identify top candidates [12]. |
| Institutional Examples | Insilico Medicine, Exscientia, Recursion, BenevolentAI, Schrödinger, MIT (BoltzGen) [11] [13]. | Commonly integrated into molecular modeling pipelines and QSAR/QSPR model development [12]. |
Experimental Protocols and Workflows
Protocol 1: A Generative AI Model for Novel Protein Binders
Recent work from MIT on the model BoltzGen provides a protocol for generating novel protein binders for "undruggable" targets [13].
- Aim: To generate novel, functional protein binders from scratch for challenging biological targets.
- Key Innovations:
- Unified Architecture: The model performs both protein design and structure prediction, maintaining state-of-the-art performance across tasks [13].
- Physics-Aware Constraints: Built-in constraints, designed with wet-lab collaborator feedback, ensure generated proteins are functional and physically plausible [13].
- Rigorous Validation: The model was tested on 26 therapeutically relevant targets, including those explicitly chosen for their dissimilarity to training data. Validation was conducted in eight independent wet labs [13].
- Outcome: BoltzGen successfully generated novel protein binders ready to enter the drug discovery pipeline, demonstrating the ability to address previously undruggable targets [13].
Protocol 2: A Hybrid Workflow Merging Generative AI with Active Learning
A 2025 study published in Communications Chemistry detailed a hybrid workflow that nests a generative model within an active learning framework to overcome the limitations of either method used in isolation [12].
- Aim: To generate diverse, drug-like molecules with high predicted affinity and synthesis accessibility for specific targets.
- Workflow Overview:
Diagram 1: VAE-Active Learning Workflow
- Key Steps & Methodology:
- Data Representation & Initial Training: A Variational Autoencoder (VAE) is initially trained on a general set of molecules, then fine-tuned on a target-specific set [12].
- Nested Active Learning Cycles:
- Inner Cycle (Chemical Optimization): The VAE generates molecules, which are evaluated by chemoinformatic oracles for drug-likeness, synthetic accessibility (SA), and novelty. Molecules passing these filters are used to fine-tune the VAE, creating a self-improving cycle focused on chemical desirability [12].
- Outer Cycle (Affinity Optimization): After several inner cycles, accumulated molecules undergo physics-based evaluation (e.g., molecular docking). Those with excellent docking scores are added to a permanent set used for further VAE fine-tuning, shifting the focus towards high-affinity binders [12].
- Candidate Selection: Promising candidates from the permanent set undergo more rigorous molecular modeling simulations (e.g., PELE, Absolute Binding Free Energy) before final selection for synthesis and in vitro testing [12].
- Experimental Validation:
The Scientist's Toolkit: Essential Research Reagents & Solutions
The following table details key computational and experimental resources integral to implementing the described AI-driven discovery workflows.
Table 2: Key Research Reagent Solutions for AI-Driven Drug Discovery
| Item / Solution | Function / Description | Example Use Case |
|---|---|---|
| Generative AI Platforms (e.g., Exscientia, Insilico, BoltzGen) | Software that uses AI to design novel molecular structures from scratch based on desired properties [11] [13]. | De novo design of protein binders (BoltzGen) [13] or small-molecule inhibitors (Insilico's ISM001-055) [11]. |
| Active Learning (AL) Framework | An iterative computational protocol that selects the most informative data points for evaluation to maximize learning efficiency [12]. | Prioritizing compounds for docking studies or bioassays to rapidly identify hits with minimal resource expenditure [12]. |
| Variational Autoencoder (VAE) | A type of generative model that learns a compressed, continuous representation (latent space) of molecular structures, enabling smooth interpolation and generation [12]. | Core component of the hybrid workflow for generating novel molecules; its latent space is iteratively refined via AL cycles [12]. |
| Physics-Based Oracle (e.g., Molecular Docking, PELE, ABFE) | Computational methods that use physical principles to predict the binding affinity and pose of a molecule to a target protein [12]. | Used in the outer AL cycle to evaluate and filter generated molecules for their predicted binding energy and mode of action [12]. |
| Chemoinformatic Oracle | Algorithms that predict chemical properties such as drug-likeness, synthetic accessibility (SA), and novelty [12]. | Used in the inner AL cycle to filter out generated molecules that are not synthesizable or do not adhere to drug-like criteria [12]. |
Integrated Workflow for Target Selection and Candidate Generation
For a research team aiming to initiate a new project, the following diagram outlines a logical decision pathway for selecting and applying these AI methodologies.
Diagram 2: Target Strategy Selection
The data and experimental protocols synthesized here demonstrate that AI methodologies, particularly generative AI and active learning, are transitioning from theoretical promise to tangible utility in combating Eroom's law. While no AI-discovered drug has yet gained full approval, the acceleration of candidates into clinical trials and the enhanced efficiency in pre-clinical stages provide compelling evidence of a paradigm shift [11]. The most powerful approach may not be a choice between generative AI or active learning, but their strategic integration. The hybrid VAE-AL workflow, which leverages the creative power of generative models guided by the efficient, physics-informed prioritization of active learning, offers a robust framework for generating high-quality, novel drug candidates [12]. For researchers and drug development professionals, mastering these tools and workflows is no longer a niche specialty but an economic and scientific imperative to ensure a future pipeline of innovative and accessible therapeutics.
The process of drug discovery has historically been characterized by high costs, extensive timelines, and low success rates. Traditional methods, which often rely on the exhaustive evaluation of molecular libraries, fundamentally limit the exploration of vast and diverse chemical spaces [12]. Generative Artificial Intelligence (GenAI) represents a disruptive paradigm shift, moving from a "design first, then predict" approach to an inverse "describe first, then design" methodology [12]. This allows researchers to algorithmically navigate the estimated 10^23 to 10^60 drug-like molecules in the chemical universe to create novel biological compounds from scratch [14] [15]. By learning the underlying patterns and rules of chemical and biological data, generative models can produce previously unseen molecular structures with tailored properties, dramatically accelerating the identification of promising therapeutic candidates [16] [15] [17].
This review positions generative AI as the creative engine for de novo molecular design, objectively comparing its performance and methodologies against and in conjunction with other computational approaches, particularly active learning (AL). Active learning is a specific instance of sequential experimental design that uses machine learning to intelligently choose the next batch of molecular structures for evaluation, closely mimicking the iterative design-make-test-analyze cycles of laboratory experiments [18]. We will explore how these approaches individually and synergistically address the core challenges of modern drug discovery.
Comparative Analysis of Generative AI Architectures
Various generative AI architectures have been developed, each with distinct strengths, limitations, and optimal applications. The table below provides a structured comparison of the primary model types used in de novo molecular design.
Table 1: Comparison of Key Generative AI Architectures for Molecular Design
| Model Architecture | Core Operating Principle | Key Advantages | Inherent Challenges | Exemplary Applications |
|---|---|---|---|---|
| Variational Autoencoders (VAEs) [18] [14] | Encodes input into a probabilistic latent space; decodes sampled points to generate new data [17]. | Continuous, interpretable latent space enabling smooth interpolation; robust and scalable training; fast parallelizable sampling [12]. | Can generate blurry or invalid structures; prior distribution may over-simplify complex data [14]. | Integration with active learning cycles; efficient exploration of chemical space [12]. |
| Generative Adversarial Networks (GANs) [14] [17] | Two neural networks (generator & discriminator) are trained adversarially [17]. | Capable of producing high yields of chemically valid molecules [12]. | Training instability and "mode collapse" (limited diversity) [12]. | Image-driven molecular design; creative content generation [17]. |
| Autoregressive Transformers [12] [17] | Models sequence data (e.g., SMILES) by predicting the next token based on all previous ones [17]. | Captures long-range dependencies in data; leverages powerful pre-trained chemical language models [12]. | Sequential decoding can make training and sampling slower [12]. | Goal-directed generation using large chemical corpora [19]. |
| Diffusion Models [14] [19] | Iteratively denoises random noise into valid molecular structures through a reversal process [12] [17]. | High sample quality and diversity; state-of-the-art performance in structured output generation [12] [19]. | Computationally intensive due to many sampling steps [12]. | 3D molecular structure generation [19]; high-fidelity inverse design [17]. |
Performance Benchmarking: Generative Models and Hybrid Workflows
The true test of these technologies lies in their ability to produce novel, valid, and effective molecular structures. The following table summarizes quantitative performance data from recent studies and workflows, highlighting the synergy between generative AI and active learning.
Table 2: Experimental Performance of Generative AI and Active Learning Workflows
| Study / Workflow | Core Methodology | Target(s) | Key Experimental Results & Performance Metrics |
|---|---|---|---|
| VAE with Nested Active Learning [12] | VAE integrated with inner (chemoinformatics) and outer (molecular modeling) active learning cycles. | CDK2, KRAS | CDK2: 9 molecules synthesized, 8 showed in vitro activity (1 with nanomolar potency). KRAS: 4 molecules identified with potential activity. Generated novel, diverse scaffolds with high predicted affinity and synthesis accessibility. |
| REINVENT + Free Energy Simulations [18] | Generative AI (REINVENT) combined with precise absolute binding free energy (ABFE) simulations in an active learning protocol. | 3CLpro, TNKS2 | Discovered ligands with higher scores than a baseline surrogate model for 3CLpro and compounds with experimentally determined affinities for TNKS2. Achieved high chemical diversity, exploring a different chemical space than the baseline. |
| Property-Guided Diffusion (GaUDI) [17] | Equivariant graph neural network for property prediction combined with a generative diffusion model. | Organic Electronic Materials | Achieved 100% validity in generated molecular structures while optimizing for single and multiple objectives. |
| Graph Convolutional Policy Network (GCPN) [17] | Reinforcement learning (RL) model that sequentially adds atoms and bonds to construct molecules. | General Molecular Properties | Demonstrated capability to generate molecules with desired chemical properties while ensuring high chemical validity. |
| GraphAF [17] | Autoregressive flow-based model fine-tuned with reinforcement learning. | General Molecular Properties | Combined efficient sampling from a learned distribution with targeted optimization towards desired molecular properties. |
Detailed Experimental Protocols
To ensure reproducibility and provide a clear understanding of the methodological rigour behind these approaches, we detail two of the most effective protocols from the benchmarked studies.
Protocol 1: VAE with Nested Active Learning Cycles
This workflow, which yielded experimentally validated hits for CDK2 and KRAS, integrates generative and discriminative models within an iterative refinement framework [12].
Data Representation and Initial Training:
- Representation: Training molecules are represented as SMILES strings, which are tokenized and converted into one-hot encoding vectors.
- Training: A VAE is first pre-trained on a general molecular dataset to learn the fundamental rules of chemical viability. It is then fine-tuned on a target-specific training set to bias the generation towards structures with increased target engagement.
Molecule Generation and Inner AL Cycle (Cheminformatics Oracle):
- Generation: The fine-tuned VAE is sampled to yield a batch of new molecular structures.
- Evaluation: Generated molecules are evaluated by cheminformatics "oracles" for key properties including:
- Drug-likeness: Adherence to rules like Lipinski's Rule of Five.
- Synthetic Accessibility (SA): Estimated ease of chemical synthesis.
- Novelty/Dissimilarity: Measured against the current training set to ensure exploration.
- Fine-tuning: Molecules meeting predefined thresholds are added to a "temporal-specific set," which is used to further fine-tune the VAE, creating a feedback loop that prioritizes desired chemical properties.
Outer AL Cycle (Physics-Based Affinity Oracle):
- After several inner cycles, accumulated molecules in the temporal-specific set undergo more computationally intensive evaluation via molecular docking simulations, which serve as a physics-based affinity oracle.
- Molecules with favorable docking scores are transferred to a "permanent-specific set," which is used for the next round of VAE fine-tuning, directly steering the generation toward structures with high predicted binding affinity.
Candidate Selection and Validation:
- The most promising candidates from the permanent-specific set undergo rigorous filtration and advanced molecular modeling, such as Protein Energy Landscape Exploration (PELE) simulations, for an in-depth assessment of binding interactions and stability [12].
- Top-ranked compounds are then selected for synthesis and in vitro bioassays for experimental validation.
The following workflow diagram illustrates this nested active learning process:
Protocol 2: REINVENT with Binding Free Energy Ranking
This GAL (Generative Active Learning) protocol demonstrates the powerful combination of AI-driven generation with high-accuracy physics-based simulations on high-performance computing systems [18].
Molecular Generation with REINVENT: The REINVENT algorithm, a specialized generative model for molecular design, is used to propose a large initial batch of candidate molecules conditioned on the target protein.
Surrogate Model Pre-screening: A faster, surrogate machine learning model (e.g., a QSAR or docking model) is used to screen the large generated library and select a smaller, top-ranking batch of molecules for more precise evaluation. This step optimizes computational efficiency.
Precise Affinity Ranking via Free Energy Simulations: The selected batch of molecules undergoes rigorous binding affinity assessment using Absolute Binding Free Energy (ABFE) calculations, specifically the ESMACS (Enhanced Sampling of Molecular dynamics with Approximation of Continuum Solvent) protocol. These physics-based molecular dynamics simulations provide a highly accurate ranking of candidates, far surpassing the precision of docking scores.
Active Learning Feedback Loop: The results from the ABFE calculations are fed back into the REINVENT model. This feedback informs and guides the subsequent generation cycle, creating a closed-loop system that iteratively improves the quality of the generated molecules toward higher-affinity ligands.
The Scientist's Toolkit: Essential Research Reagents and Solutions
The experimental protocols outlined above rely on a suite of computational tools and resources. The following table details these key "research reagents" for implementing generative AI and active learning in molecular design.
Table 3: Essential Research Reagents and Computational Tools for AI-Driven Molecular Design
| Tool / Resource | Type | Primary Function in Workflows | Exemplary Use Case |
|---|---|---|---|
| SMILES/SELFIES [15] | Molecular Representation | String-based representations that encode molecular structure; the "language" for many generative models. | SMILES strings are tokenized and used as input for VAEs and Transformer models [12] [15]. |
| VAE (Variational Autoencoder) [12] | Generative Model Architecture | Learns a continuous latent representation of molecules; enables generation and interpolation in chemical space. | Core generator in the nested AL workflow for CDK2/KRAS [12]. |
| REINVENT [18] | Generative AI Software | A generative model specifically designed for de novo molecular design and optimization. | Used in the GAL protocol for generating ligands for 3CLpro and TNKS2 [18]. |
| Molecular Docking [12] | Physics-Based Simulation | Predicts the preferred orientation and preliminary binding affinity of a small molecule to a protein target. | Serves as the "affinity oracle" in the outer active learning cycle [12]. |
| ABFE (Absolute Binding Free Energy) [18] | Physics-Based Simulation | Provides highly accurate calculation of binding affinity using molecular dynamics; used for precise ranking. | Final rigorous assessment in the REINVENT GAL protocol [18]. |
| PELE (Protein Energy Landscape Exploration) [12] | Advanced Sampling Algorithm | Models protein-ligand binding pathways and induced-fit conformational changes for in-depth pose analysis. | Used for candidate refinement and selection after docking in the VAE-AL workflow [12]. |
Integrated Workflow Logic
The most successful strategies merge generative AI's creative power with active learning's strategic guidance. The following diagram synthesizes the core logical relationship between these components into a unified, iterative workflow for modern, data-driven drug discovery.
The empirical data and experimental protocols presented in this review compellingly argue that generative AI serves as the indispensable creative engine for de novo molecular design. However, its full potential is unlocked when coupled with the strategic, iterative refinement provided by active learning and robust physics-based validation [12] [18]. While standalone generative models can rapidly explore chemical space, they can struggle with challenges such as target engagement, synthetic accessibility, and the generalization beyond their training data [12]. Active learning frameworks directly address these limitations by embedding generative models within a closed-loop feedback system, leveraging both cheminformatics and physics-based oracles to steer the generation toward drug-like, synthesizable, and high-affinity ligands [12].
The synergy between these approaches is evident in the reported results. The VAE-AL workflow successfully generated novel scaffolds for CDK2 and KRAS, leading to experimentally confirmed active compounds, including a nanomolar-potency inhibitor [12]. Similarly, the GAL protocol combining REINVENT with free energy simulations discovered ligands with higher scores than baseline models and accessed diverse, unexplored regions of chemical space [18]. These successes demonstrate a powerful convergence of data-driven AI and physics-based modeling, creating a new paradigm for molecular design.
Future directions in this field point towards even greater integration and automation. This includes the convergence of generative models with Bayesian retrosynthesis planners, self-supervised pre-training on ultra-large chemical corpora, and the multimodal integration of omics-derived features for precision therapeutics [19]. The synthesis of generative AI, closed-loop automation, and advanced computing is paving the way for fully autonomous molecular design ecosystems, poised to radically accelerate the journey from concept to viable therapeutic candidate [19].
In the fields of materials science and drug development, researchers face a fundamental challenge: exploring vast experimental design spaces with limited time and financial resources. Exhaustive trial-and-error approaches are often impractical, creating a critical need for strategies that can maximize information gain from a minimal number of experiments. Active Learning (AL) has emerged as a powerful solution to this problem. AL is a subfield of machine learning that studies algorithms designed to select the most informative data points to improve their own models, forming an iterative refinement loop [20]. This guide provides an objective comparison of traditional AL models against a emerging paradigm: generative AI and Large Language Model-based Active Learning (LLM-AL). Benchmarked across diverse scientific domains, these approaches demonstrate how strategic data selection can dramatically accelerate discovery, potentially reducing the number of experiments needed by over 70% [21].
Foundational Active Learning Protocols and Performance
The Core Active Learning Workflow
The standard AL process is an iterative cycle comprising three critical stages, as established in computational biology and materials science reviews [20]:
- Model Training: An initial model is learned from a (often small) starting set of experimental data.
- Query Selection: The model generates hypotheses to propose the most informative subsequent experiments. The selection is based on criteria designed to reduce model uncertainty or maximize expected improvement.
- Model Update: New data from the performed experiment is used to update the model. This cycle repeats until a performance target is met or resources are exhausted.
This workflow can be visualized as a continuous loop of learning and experimentation.
Traditional Machine Learning Models in AL
Traditional AL relies on well-established machine learning models as its core "brain" for experiment selection. The table below summarizes four common models and their typical application in AL pipelines.
Table 1: Key Traditional Machine Learning Models for Active Learning
| Model | Primary Function in AL | Key Characteristic | Noted Challenge |
|---|---|---|---|
| Gaussian Process Regressor (GPR) | Models a distribution over functions to make predictions. | Provides native uncertainty quantification, crucial for query selection. | Hyperparameter tuning is brittle with scarce data [21]. |
| Random Forest Regressor (RFR) | Ensemble model using multiple decision trees. | Robust to outliers and handles mixed data types. | Lacks inherent, well-calibrated uncertainty estimates. |
| Bayesian Neural Network (BNN) | Neural network with probability distributions over weights. | Combines flexibility of NNs with Bayesian uncertainty. | Computationally intensive and complex to train. |
| eXtreme Gradient Boosting (XGBoost) | Optimized gradient-boosting library. | High predictive accuracy and execution speed. | Not inherently designed for uncertainty-aware query. |
Documented Performance of Traditional AL
When integrated into automated, closed-loop systems, traditional AL has demonstrated significant value. Studies and industrial applications highlight its impact on experimental efficiency:
- Order-of-Magnitude Efficiency Gains: AL implementations using Bayesian optimization have repeatedly demonstrated identifying optimal material candidates with up to 10 to 20 times fewer experimental iterations compared to unguided or random screening [21].
- Enhanced Learning Outcomes: In educational settings, which serve as a proxy for training efficiency, AI-enhanced active learning programs have shown a 54% increase in test scores and generate 10 times more engagement than traditional passive methods [2].
- Corporate Training Efficiency: In corporate settings, AI-powered training, which often leverages AL principles, leads to a 57% increase in learning efficiency, with employees completing training faster while demonstrating superior mastery and retention [2].
The Emerging Paradigm: LLM-Based Active Learning
The LLM-AL Framework and Benchmarking
A 2025 study introduced a training-free LLM-based Active Learning framework (LLM-AL) that operates in an iterative few-shot setting [21]. This approach leverages the pretrained knowledge and universal token-based representations of LLMs to propose experiments directly from text-based descriptions of experimental conditions and results. The researchers benchmarked LLM-AL against conventional ML models (GPR, BNN, RFR, XGBoost) across four diverse materials science datasets: matbench_steels (alloy design), P3HT/CNT (polymer nanocomposites), Perovskite, and Membrane optimization [21].
The study explored two prompting strategies:
- Parameter-Format: Inputs are structured as concise feature-value pairs.
- Report-Format: Inputs are rewritten into expanded descriptive text to provide additional experimental context.
Comparative Performance Data
The performance of LLM-AL and traditional ML models was measured by their efficiency in converging on optimal candidates within each dataset. The results demonstrate a strong advantage for the LLM-based approach.
Table 2: Experimental Efficiency: LLM-AL vs. Traditional ML Models
| Dataset | Primary Domain | Top Performing Model(s) | Key Performance Metric |
|---|---|---|---|
| matbench_steels | Alloy Design | LLM-AL (Parameter-Format) | Consistently reached optima using <30% of data. |
| P3HT/CNT | Polymer Nanocomposites | LLM-AL | Outperformed all traditional ML models. |
| Perovskite | Photovoltaic Materials | LLM-AL | Consistently reached optima using <30% of data. |
| Membrane | Membrane Optimization | LLM-AL (Report-Format) | Most notable improvement with descriptive prompts. |
| Across all datasets | Multiple | LLM-AL | >70% reduction in experiments needed to find top candidates [21]. |
The LLM-AL Experimental Workflow
The LLM-AL framework modifies the traditional AL loop by using a Large Language Model as the surrogate model for experiment selection. The process begins with a text-based prompt that contains prior experimental results and context, from which the LLM suggests the next most informative experiment.
Critical Analysis: LLM-AL vs. Traditional Machine Learning
Performance and Stability
The benchmark study yielded several critical findings for researchers considering these approaches [21]:
- Performance: LLM-AL consistently converged to optimal targets using less than 30% of the available data across most dataset and prompt format combinations. It consistently outperformed traditional ML models, requiring fewer iterations to reach the same performance level.
- Stability: Despite the inherent non-determinism of LLMs, the performance variability of LLM-AL across repeated runs was found to be broadly consistent and within the variability range typically observed for traditional ML approaches.
- Prompting Strategy: The optimal prompting format is context-dependent. The parameter-format (concise) was superior for datasets with many independent variables (e.g., compositions), while the report-format (descriptive) excelled for datasets with procedural descriptors, where added context revealed hidden relationships.
Advantages and Limitations
Table 3: Functional Comparison: LLM-AL vs. Traditional AL
| Feature | LLM-AL | Traditional AL |
|---|---|---|
| Generalizability | High; operates in universal token space, transferable across domains [21]. | Low; often requires problem-specific feature engineering [21]. |
| Cold-Start Performance | Strong; leverages pretrained knowledge to guide exploration with sparse data [21]. | Weak; suffers from the "cold-start" problem with low predictive accuracy initially [21]. |
| Input Representation | Flexible text-based inputs (descriptive or structured). | Rigid, fixed-length numerical feature vectors. |
| Interpretability | Potential for human-readable reasoning (e.g., via chain-of-thought). | Often a "black box"; decisions can be hard to interpret. |
| Computational Cost | Higher per-query cost due to model size. | Lower per-query cost. |
| Primary Bottleneck | Prompt design and context management. | Feature engineering and hyperparameter tuning. |
The Scientist's Toolkit: Essential Research Reagents
Implementing an effective AL pipeline, whether traditional or LLM-based, requires a suite of computational "reagents." The following tools are essential for conducting modern, data-efficient research.
Table 4: Key Research Reagent Solutions for Active Learning
| Reagent / Tool | Function | Relevance to AL |
|---|---|---|
| Large Language Model (e.g., GPT, Cohere) | Core surrogate model for experiment suggestion. | Serves as the "brain" in LLM-AL, interpreting prompts and proposing experiments based on learned knowledge [21]. |
| Traditional ML Libraries (e.g., Scikit-learn, XGBoost) | Provides algorithms for GPR, RFR, XGBoost, etc. | Forms the backbone of traditional AL pipelines for model training and prediction [21]. |
| Benchmark Datasets | Standardized data for model training and validation. | Critical for benchmarking AL performance across different strategies (e.g., matbench_steels, Perovskite) [21]. |
| Interactive Visualization Tools | Elucidates the model training and query selection process. | Helps researchers understand when and how AL works by tracking prediction changes across query stages [22]. |
| High-Contrast Accessibility Tools | Ensures software and visualizations are accessible. | Crucial for inclusive tool development, testing rendering in Windows High Contrast Mode, etc. [23] [24]. |
The empirical evidence clearly positions Active Learning as a transformative strategy for optimizing data efficiency in scientific discovery. The comparison between traditional ML and the emerging LLM-AL paradigm reveals a shifting landscape. While traditional models like GPR and BNN remain powerful, they are often constrained by their lack of generalizability and reliance on feature engineering. The LLM-AL framework demonstrates that leveraging the broad, pretrained knowledge of large language models can mitigate the cold-start problem and provide a more flexible, generalizable tool for guiding experimental design across diverse domains [21]. For researchers and drug development professionals, the choice of strategy will depend on the specific problem structure, data availability, and computational resources. However, the overarching conclusion is that integrating strategic data selection via Active Learning is no longer optional but is becoming essential infrastructure for efficient and accelerated research.
The research and development (R&D) landscape is being transformed by two powerful computational approaches: generative artificial intelligence (Gen-AI) and active learning. While often discussed separately, their combined potential within scientific workflows represents a frontier of innovation. Generative AI, a subset of artificial intelligence that utilizes machine learning models to create new, original content—from molecular structures to predictive text—operates by learning patterns and structures from existing data [25]. In contrast, active learning refers to AI systems that engage in an iterative process of selecting the most informative data points for human labeling or experimental validation, thereby maximizing learning efficiency from limited data [2]. Within the context of scientific R&D, particularly in fast-moving fields like drug development, understanding the distinct strengths, limitations, and synergistic potential of these approaches is critical for accelerating discovery. This guide provides an objective comparison of their performance, supported by experimental data and concrete protocols for integration.
Performance Comparison: Quantitative Analysis of Approaches
The table below summarizes key performance metrics for Generative AI and Active Learning, synthesized from recent studies and meta-analyses. This data provides a foundation for understanding their complementary roles.
Table 1: Comparative Performance of Generative AI and Active Learning in R&D Contexts
| Performance Metric | Generative AI | Active Learning | Comparative Insights |
|---|---|---|---|
| Learning Efficiency / Score Improvement | 30% improvement in student performance [26]; improves outcomes by up to 30% [2] | 54% higher test scores in AI-enhanced active learning environments [2] | Active learning demonstrates a significantly larger effect size for knowledge acquisition and retention. |
| Effect on Innovation & Creativity | 64% of data leaders say AI enables innovation [27]; 64% of organizations report AI enables innovation [28] | Generates 10x more engagement than passive learning [2] | Gen-AI is a direct catalyst for novel idea generation, while active learning sustains deep engagement necessary for innovation. |
| Impact on Cognitive Engagement | Can lower cognitive effort and "cognitive debt"; associated with less brain activity in writing tasks [25] | High cognitive engagement through iterative querying and problem-solving [2] | A key differentiator; active learning promotes deeper cognitive processing, while Gen-AI risks cognitive offloading. |
| Intervention Duration for Efficacy | Medium-term interventions (4–12 weeks) yielded higher effect sizes [26] | Effective in short-term, focused training initiatives [29] | Gen-AI may require longer integration to show stable benefits, while active learning can produce rapid gains. |
| Domain Specificity & Accuracy | May be less accurate for highly technical or niche tasks without fine-tuning [27] | Excels at identifying and addressing specific knowledge gaps [2] | Active learning is inherently designed to navigate complex, specific problem spaces efficiently. |
Experimental Protocols: Methodologies for Validation
To objectively evaluate these approaches, researchers have employed rigorous experimental designs. Below are detailed protocols from key studies.
Protocol 1: Randomized Controlled Trial on AI Reliance
- Objective: To measure the impact of unrestricted Gen-AI use on learning outcomes and student engagement in a technical subject.
- Source: Corvinus University of Budapest experiment on operations research students [6].
- Methodology:
- Population: Students enrolled in an operations research class.
- Randomization: Participants were randomly assigned to one of two groups using a random algorithm to prevent self-selection bias.
- Intervention Group: Permitted to use generative AI tools (e.g., ChatGPT) during classes and examinations.
- Control Group: Required to complete all work without the use of AI tools.
- Fairness Control: A point compensation mechanism was applied post-test to equalize the average performance between the two groups, ensuring neither was disadvantaged by the group assignment.
- Metrics: Measured understanding of material, engagement levels, and final exam performance.
- Key Findings: The experiment, though revised due to student complaints, indicated that uncontrolled use of AI tools led to disengaged students and a lower understanding of the material. The strong student reaction to the removal of AI access was itself evidence of deep reliance [6].
Protocol 2: Meta-Analysis on Generative AI Efficacy
- Objective: To comprehensively synthesize the effect of Generative AI on university students' learning outcomes across diverse domains.
- Source: Large-scale meta-analysis published in ScienceDirect [26].
- Methodology:
- Literature Search: Systematic search across multiple academic databases following PRISMA guidelines.
- Study Selection: Incorporated 57 qualified studies with 97 effect size estimations. Studies were required to control for baseline differences between experimental and control groups.
- Framework: Analysis was grounded in the Activity Theory-Mobile Computer-Supported Collaborative Learning (AT-MCSCL) framework, categorizing moderators into learner features, tools, roles, rules, and context.
- Outcome Measures: Academic achievement, affective-motivational status, higher-order thinking, language skills, and metacognition.
- Key Findings: Gen-AI produced a large overall positive effect on learning outcomes. The largest effect sizes were observed in language skills, followed by academic achievement, affective-motivational status, and higher-order thinking. The effect on metacognition was not statistically significant [26].
Synergistic Workflow Integration
The true power of Gen-AI and active learning emerges when they are integrated into a cohesive R&D workflow. The following diagram visualizes this synergistic cycle, where Gen-AI acts as a generator of possibilities and active learning as a mechanism for targeted validation and refinement.
This workflow can be operationalized in a drug discovery pipeline as follows:
- Generative AI for Candidate Generation: AI models like specialized LLMs (e.g., Merlyn Mind) or molecular structure generators propose initial compound structures or biological targets based on vast scientific literature and existing drug databases [30].
- Active Learning for Prioritization: The list of generated candidates is too large for experimental validation. An active learning algorithm queries a human expert (or a high-fidelity simulation) to label the most promising or diverse candidates, dramatically narrowing the list to a high-priority subset [2].
- Generative AI for Protocol Design: For the prioritized candidates, Gen-AI assists in drafting initial experimental protocols or suggesting necessary research reagents [25].
- Active Learning for Experimental Optimization: The experimental design is refined through an active learning loop that seeks to minimize the number of wet-lab experiments needed to reach a conclusion, efficiently exploring parameter spaces like concentration, temperature, and timing.
- Iterative Validation and Re-generation: Results from physical experiments are fed back into the generative model, fine-tuning it to produce more accurate and viable candidates in the next cycle, thus closing the loop [27].
The Scientist's Toolkit: Essential Research Reagents & Solutions
For researchers aiming to implement the hybrid workflow described above, the following table details key "reagents" — both computational and physical — that are essential for conducting experiments.
Table 2: Key Research Reagent Solutions for AI-Enhanced R&D
| Research Reagent / Tool | Type | Primary Function in Workflow |
|---|---|---|
| Large Language Models (e.g., GPT-4, Claude, Gemini) | Computational | Serves as a core Generative AI engine for brainstorming, literature synthesis, hypothesis generation, and initial code or protocol drafting [27] [30]. |
| Specialized AI Tutors (e.g., LearnLM, Physics Wallah's Model) | Computational | Provides domain-specific knowledge support and guided problem-solving, acting as an on-demand expert in fields like STEM [30]. |
| Active Learning Query Strategy Algorithms (e.g., Uncertainty Sampling, Diversity Sampling) | Computational | The core "logic" that decides which data points or experiments are most informative to perform next, optimizing resource allocation [2]. |
| Synthetic Data Generators | Computational | Creates statistically realistic datasets to augment small experimental datasets, used for training robust machine learning models without privacy concerns or high initial costs [27]. |
| High-Throughput Screening Assays | Wet-lab / Physical | Provides the rapid, automated experimental data generation required to feed and validate the active learning cycle, especially in biology and chemistry [28]. |
| Automated Lab Equipment & Lab Information Management Systems (LIMS) | Physical / Digital Infrastructure | Executes designed experiments and seamlessly logs structured, high-quality data back into the digital workflow, creating a closed-loop system [28]. |
Generative AI and active learning are not competing technologies but complementary forces in the modern R&D toolkit. The evidence shows that Generative AI excels as a force multiplier for creativity and content generation, while active learning provides the strategic focus and cognitive engagement needed for deep, efficient knowledge acquisition. The risks of over-reliance on Gen-AI, such as cognitive offloading and superficial understanding, can be mitigated by integrating it within an active learning framework that demands continuous validation and critical thinking. For research organizations, the imperative is to move beyond siloed experiments and strategically design workflows that harness this synergy. By doing so, they can unlock unprecedented acceleration in innovation, from the initial spark of an idea to its rigorous and efficient validation.
From Theory to Bench: Practical Applications of Generative AI and Active Learning in R&D
The pharmaceutical industry is undergoing a significant transformation through the integration of artificial intelligence (AI) into traditional drug discovery workflows. This evolution represents not a replacement of established approaches but rather the development of complementary tools that augment human expertise and computational chemistry methods refined over decades [31]. Two prominent AI paradigms have emerged: generative AI, which creates novel molecular structures from scratch, and active learning, which strategically selects experiments to maximize learning and optimize compounds. While both approaches leverage machine learning, they operate on fundamentally different principles and excel in distinct aspects of the drug discovery pipeline.
Generative AI involves algorithms that create new data based on learned patterns, with models like variational autoencoders (VAEs) and generative adversarial networks (GANs) being trained on chemical and biological datasets to propose novel molecules [32]. In contrast, active learning represents an iterative framework where the AI algorithm selectively identifies the most informative experiments to perform, thereby maximizing knowledge gain while minimizing resource expenditure [33]. This comparative guide examines the performance, applications, and practical implementation of these two approaches within modern pharmaceutical research and development.
Comparative Performance Analysis
The table below summarizes the key performance characteristics of generative AI versus active learning approaches across critical metrics relevant to drug discovery professionals.
Table 1: Performance Comparison of Generative AI and Active Learning in Drug Discovery
| Performance Metric | Generative AI | Active Learning |
|---|---|---|
| Primary Application | De novo molecular design; creating novel chemical entities [32] | Lead optimization; refining existing compounds [34] [33] |
| Data Efficiency | Requires large initial training datasets (~104-106 compounds) [31] | Highly efficient in low-data regimes; optimal for ~102 initially known compounds [33] |
| Success Rate | Can generate synthesizable candidates with drug-like properties (>70% success in some studies) [32] | Discovers 60% of synergistic drug pairs by exploring only 10% of combinatorial space [33] |
| Time Acceleration | Reduced target-to-candidate timeline from years to months (e.g., 18 months for INS018_055) [31] | Reduces experimental burden by 82% for synergy identification [33] |
| Key Advantage | Exploration of novel chemical space beyond human bias | Cost-effective exploitation of known chemical space |
| Clinical Validation | Multiple candidates in trials (e.g., rentosertib - Phase II; ISM-6631 - Phase I) [35] | Extensive retrospective validation; emerging prospective applications [34] |
Experimental Protocols and Workflows
Generative AI Protocol for De Novo Drug Design
The experimental workflow for generative AI in de novo drug design follows a multi-stage process that integrates deep learning with experimental validation [32] [31]:
Data Curation and Preprocessing: Collect and curate large-scale chemical databases (e.g., ChEMBL, ZINC) containing molecular structures and associated biological activities. Apply standardization, normalization, and chemical representation techniques (e.g., SMILES, molecular graphs).
Model Training: Implement and train generative models such as:
- Variational Autoencoders (VAEs): Learn continuous latent representations of molecular structures enabling interpolation and novel compound generation.
- Generative Adversarial Networks (GANs): Employ generator and discriminator networks in adversarial training to produce realistic molecular structures.
- Transformers: Utilize self-attention mechanisms to model complex molecular patterns and generate novel structures autoregressively.
Molecular Generation and Optimization: Generate novel compounds conditioned on specific target properties (e.g., high binding affinity, optimal physicochemical properties, selectivity). Apply transfer learning and reinforcement learning to fine-tune models for specific target classes.
In Silico Validation: Screen generated molecules using predictive models for key parameters including:
- Drug-target interaction predictions using deep learning classifiers
- ADMET (Absorption, Distribution, Metabolism, Excretion, Toxicity) profiling
- Synthetic accessibility scoring
Experimental Validation: Synthesize top-ranking candidates and validate through in vitro assays (binding affinity, functional activity) and in vivo studies for promising leads.
Active Learning Protocol for Lead Optimization
The active learning framework for lead optimization employs an iterative, closed-loop design that efficiently guides experimental efforts [34] [33]:
Initial Model Training: Develop a preliminary machine learning model (e.g., random forest, neural network) using initially available compound activity data. Molecular representations may include Morgan fingerprints, MAP4 fingerprints, or graph-based embeddings.
Acquisition Function Design: Implement selection strategies to identify the most informative compounds for experimental testing, including:
- Exploration: Selecting compounds where the model is most uncertain (e.g., high prediction variance)
- Exploitation: Selecting compounds predicted to have high potency or desirable properties
- Hybrid Approaches: Balancing exploration and exploitation using methods like upper confidence bound
Iterative Experimentation Cycle:
- Batch Selection: Choose a limited set of compounds (batch size typically 10-100) based on the acquisition function
- Experimental Testing: Synthesize and evaluate selected compounds for target activity and properties
- Model Retraining: Update the predictive model with new experimental results
- Convergence Check: Repeat until performance criteria met or budget exhausted
Validation and Model Interpretation: Validate final model performance on hold-out test sets and apply explainable AI techniques to identify structural features driving compound activity.
AI Drug Discovery Workflow
The Scientist's Toolkit: Essential Research Reagents and Platforms
Successful implementation of AI-driven drug discovery requires specific computational tools, datasets, and experimental resources. The following table details key components of the modern AI drug discovery toolkit.
Table 2: Essential Research Reagents and Platforms for AI-Driven Drug Discovery
| Tool Category | Specific Examples | Function & Application |
|---|---|---|
| Generative AI Platforms | Insilico Medicine, Exscientia, Relay Therapeutics | End-to-end platforms for de novo molecular design and optimization [32] [35] |
| Active Learning Frameworks | RECOVER, DeepSynergy, Custom implementations | Iterative experimental design for lead optimization and synergy prediction [33] |
| Molecular Representations | Morgan Fingerprints, MAP4, Graph Neural Networks | Convert chemical structures into numerical features for machine learning [33] |
| Cellular Context Features | GDSC gene expression, CCLE omics data | Incorporate cellular environment into prediction models [33] |
| Benchmark Datasets | Oneil, ALMANAC, DrugComb | Curated drug combination screening data for training and validation [33] |
| Validation Assays | High-throughput screening, Binding assays, ADMET profiling | Experimental verification of AI-generated compounds [31] |
Signaling Pathways and Experimental Design
AI Method Selection Guide
Discussion and Future Perspectives
The integration of both generative AI and active learning into pharmaceutical R&D represents a paradigm shift in how drug discovery is conducted. Generative AI excels in the early discovery phase by exploring vast chemical spaces and generating novel molecular entities, while active learning provides superior efficiency in lead optimization by strategically guiding experimental resources [32] [33]. The most successful implementations increasingly leverage hybrid approaches that combine the creative capacity of generative models with the resource efficiency of active learning.
Current challenges include data quality and availability, model interpretability, regulatory acceptance, and integration with traditional medicinal chemistry expertise [31]. However, the field is advancing rapidly, with the 2025 FDA draft guidance establishing a risk-based credibility assessment framework for AI applications that complements existing regulatory frameworks [31]. As these technologies mature, they promise to significantly reduce the time and cost of drug development while increasing success rates, ultimately accelerating the delivery of innovative therapies to patients.
The future of AI in drug discovery lies in the development of more sophisticated agentic AI systems that can autonomously navigate discovery pipelines, the integration of multi-modal data (genomic, proteomic, clinical), and the creation of more accurate predictive models through advances in foundation models specifically trained on chemical and biological data [31]. For researchers and drug development professionals, understanding the complementary strengths of generative AI and active learning is crucial for selecting the appropriate tool for each stage of the drug discovery process.
The exploration of chemical space for novel drug candidates represents a monumental challenge in pharmaceutical research, with the space of synthesizable small molecules estimated to exceed 10^33 compounds [36]. Generative artificial intelligence (AI) has emerged as a transformative force in this domain, enabling researchers to design molecules with tailored properties rather than relying solely on exhaustive screening [37] [17]. Among the various architectures employed, Variational Autoencoders (VAEs), Generative Adversarial Networks (GANs), and Transformer-based models have demonstrated particular promise for molecular generation tasks. These architectures differ significantly in their theoretical foundations, operational mechanisms, and performance characteristics, making the choice between them critical for successful drug discovery applications.
When framed within broader research on generative AI combined with active learning approaches, these models take on enhanced significance. Active learning creates iterative feedback loops where models are refined based on computational or experimental evaluations, progressively improving the quality of generated molecules [38] [12]. This synergy addresses a key limitation of standalone generative models: poor generalization to new chemical spaces beyond the training data distribution. As the field advances toward increasingly automated drug discovery pipelines, understanding the comparative strengths and limitations of these architectures becomes essential for researchers and drug development professionals.
Architectural Frameworks and Mechanisms
Variational Autoencoders (VAEs)
VAEs are probabilistic generative models that learn to encode input molecules into a lower-dimensional latent space and then decode samples from this space to generate novel molecular structures [39] [37]. The architecture consists of two primary components: an encoder that maps input data to a probability distribution in latent space (typically characterized by mean and standard deviation parameters), and a decoder that reconstructs molecules from points sampled from this distribution [40]. The training objective combines reconstruction loss (ensuring input molecules can be accurately reconstructed) with a KL-divergence term that regularizes the latent space to approximate a standard normal distribution [40].
Modern VAE implementations for molecular generation have evolved significantly from early approaches. The STAR-VAE (Selfies-encoded, Transformer-based, AutoRegressive Variational Auto Encoder) framework exemplifies this evolution, incorporating a bi-directional Transformer encoder and an autoregressive Transformer decoder [36]. This architecture is trained on large-scale molecular datasets (e.g., 79 million drug-like molecules from PubChem) and uses SELFIES representations to guarantee 100% syntactic validity of generated molecules [36]. The latent-variable formulation provides a principled foundation for conditional generation, where property predictors supply conditioning signals that consistently shape the latent prior, inference network, and decoder [36].
Another innovative approach, the Transformer Graph Variational Autoencoder (TGVAE), employs molecular graphs as input data to capture complex structural relationships more effectively than string-based representations [41]. This model addresses common issues like over-smoothing in graph neural networks and posterior collapse in VAEs to ensure robust training and improve the generation of chemically valid and diverse molecular structures [41].
Generative Adversarial Networks (GANs)
GANs operate on a fundamentally different principle, framing molecular generation as an adversarial game between two competing neural networks: a generator that creates synthetic molecules from random noise, and a discriminator that distinguishes between real molecules from the training data and fake ones produced by the generator [39] [40]. Through this adversarial training process, the generator progressively improves its ability to produce realistic molecular structures that can fool the discriminator [40].
Despite their potential, GANs face significant challenges when applied to discrete molecular representations like SMILES strings, as the discrete nature of the data disrupts gradient-based optimization essential for GAN training [42]. Several architectures have been developed to address these limitations. RL-MolGAN introduces a novel Transformer-based discrete GAN framework that utilizes a first-decoder-then-encoder structure, diverging from traditional Transformer designs [42]. This framework integrates reinforcement learning (RL) with Monte Carlo Tree Search (MCTS) to stabilize GAN training and optimize the chemical properties of generated molecules [42]. An extended version, RL-MolWGAN, incorporates Wasserstein distance and mini-batch discrimination to further enhance training stability [42].
Another approach to addressing GAN limitations involves hybrid architectures. The LM-GAN framework combines a masked language model with a GAN, leveraging the language model's ability to learn common subsequences from training data and apply them as automated, generalized mutation operators [43]. This hybrid approach demonstrates superior performance over standalone masked language models, particularly for smaller population sizes [43].
Transformer-Based Models
Transformer architectures have revolutionized molecular generation by leveraging self-attention mechanisms to capture long-range dependencies in molecular representations [39]. Unlike sequential models that process tokens one by one, Transformers process all parts of an input sequence simultaneously, making them particularly effective at addressing the sensitivity of SMILES representations to small perturbations [42].
Transformer-based molecular generators are typically organized into decoder-only and encoder-decoder families [36]. Decoder-only models like MolGPT adapt the GPT-style autoregressive Transformer for SMILES, generating molecules token by token with high validity and support for property- and scaffold-conditioned sampling [36]. However, they lack an explicit encoder to structure latent representations, which limits controllable exploration of molecular space [36]. Encoder-decoder models like Chemformer and SELFIES-TED provide richer conditioning interfaces than decoder-only systems but often function as deterministic transducers rather than probabilistic latent-variable generators [36].
The STAR-VAE framework represents a hybrid approach that combines Transformer architectures with latent-variable modeling, unifying broad distribution learning with controllable conditional generation [36]. This model demonstrates how modernized, scale-appropriate VAEs with Transformer components remain competitive for molecular generation when paired with principled conditioning and parameter-efficient finetuning [36].
Performance Comparison and Experimental Data
Quantitative Performance Metrics
Table 1: Benchmark Performance of Molecular Generation Architectures
| Architecture | Model Name | Validity (%) | Uniqueness (%) | Novelty (%) | Diversity | Property Optimization |
|---|---|---|---|---|---|---|
| VAE | STAR-VAE [36] | >99% (SELFIES) | High | High | 0.83 (GuacaMol) | Strong (Tartarus) |
| VAE | TGVAE [41] | High | High | High | Superior to baselines | Effective |
| GAN | RL-MolGAN [42] | High | - | - | - | Effective for target properties |
| GAN | LM-GAN [43] | - | - | - | Superior for small populations | Enhanced efficiency |
| Transformer | MolGPT [36] | High | High | Lower than other ML frameworks | - | Supports conditioned generation |
| Transformer | STAR-VAE [36] | >99% | - | - | - | Shifts docking score distributions |
Table 2: Conditional Generation Performance on Protein Targets
| Architecture | Model | Target Protein | Performance | Experimental Validation |
|---|---|---|---|---|
| VAE with AL | VAE-AL GM [12] | CDK2 | 8/9 synthesized molecules showed in vitro activity, 1 with nanomolar potency | Synthesized and tested |
| VAE with AL | VAE-AL GM [12] | KRAS | 4 molecules with predicted activity | In silico validation |
| Conditional VAE | STAR-VAE [36] | 1SYH, 6Y2F | Docking score distribution statistically stronger than baseline | In silico docking |
Analysis of Comparative Performance
The experimental data reveals distinct performance patterns across architectural families. VAE-based models consistently demonstrate strong performance across multiple benchmarks, with STAR-VAE matching or exceeding baselines on GuacaMol and MOSES benchmarks under comparable computational budgets [36]. The TGVAE similarly outperforms existing approaches, generating a larger collection of diverse molecules and discovering previously unexplored structures [41]. Notably, VAEs show particular strength in conditional generation tasks, with STAR-VAE successfully shifting docking-score distributions toward stronger predicted binding for specific protein targets [36].
GAN-based approaches show promising results in property optimization but face challenges in training stability and diversity. The RL-MolGAN framework demonstrates the ability to generate molecules with desired chemical properties by incorporating reinforcement learning and MCTS [42]. The LM-GAN hybrid architecture shows particular advantage in scenarios with smaller population sizes, addressing the mode collapse problem that often plagues traditional GANs [43].
Transformer-based models excel in capturing long-range dependencies in molecular sequences but may exhibit limitations in novelty compared to other approaches [36]. MolGPT demonstrates high validity and uniqueness but lower novelty scores compared to various modern machine learning frameworks [36]. However, Transformer architectures integrated with latent-variable formulations, as in STAR-VAE, overcome these limitations while maintaining the benefits of attention mechanisms [36].
Experimental Protocols and Methodologies
Benchmarking Standards
Experimental evaluation of molecular generative models typically follows established benchmarking protocols using standardized datasets and metrics. Common benchmarks include GuacaMol [36], MOSES [36], and Tartarus [36], which provide standardized frameworks for evaluating model performance across multiple dimensions including validity, uniqueness, novelty, and diversity.
The GuacaMol benchmark employs a suite of tasks designed to evaluate various aspects of generative model performance, including the ability to generate molecules with specific property profiles [36]. The MOSES benchmark provides standardized metrics and baselines to ensure fair comparison between different generative models [36]. The Tartarus benchmark specifically focuses on protein-ligand design, evaluating a model's ability to generate molecules with strong predicted binding affinities for specific protein targets [36].
Active Learning Integration Protocols
The integration of generative models with active learning frameworks follows specific experimental protocols that create iterative feedback loops. The VAE-AL GM workflow provides a representative example, featuring a VAE with two nested active learning cycles that iteratively refine predictions using chemoinformatics and molecular modeling predictors [12].
Table 3: Active Learning Cycle Components in Molecular Generation
| Cycle Type | Evaluation Oracles | Filtering Criteria | Output |
|---|---|---|---|
| Inner AL Cycle [12] | Chemoinformatic predictors (drug-likeness, synthetic accessibility) | Druggability, SA, similarity thresholds | Temporal-specific set |
| Outer AL Cycle [12] | Molecular modeling (docking simulations) | Docking score thresholds | Permanent-specific set |
| Full Pipeline [12] | Molecular dynamics (PELE), binding free energy simulations | Stringent filtration for candidate selection | Molecules for synthesis |
The experimental protocol typically involves these key steps:
- Initial Training: The generative model is initially trained on a general training set to learn fundamental chemical principles, then fine-tuned on a target-specific training set to enhance target engagement [12].
- Molecule Generation: The trained model samples new molecules from the learned chemical space [12].
- Inner AL Cycles: Generated molecules are evaluated using chemoinformatic oracles for drug-likeness, synthetic accessibility, and similarity to known actives. Molecules meeting threshold criteria are used to fine-tune the model in subsequent iterations [12].
- Outer AL Cycles: Accumulated molecules from inner cycles undergo more computationally intensive evaluations (e.g., docking simulations). High-performing molecules are transferred to a permanent-specific set for model fine-tuning [12].
- Candidate Selection: After multiple AL cycles, the most promising candidates undergo stringent filtration and selection processes, potentially including advanced molecular modeling and experimental validation [12].
This protocol demonstrates how active learning enables generative models to extrapolate beyond their training data, with one study reporting generated molecule properties reaching 0.44 standard deviations beyond the training data range [38].
Research Reagent Solutions
Table 4: Essential Research Resources for Molecular Generation Experiments
| Resource Category | Specific Examples | Function in Research | Key Features |
|---|---|---|---|
| Chemical Databases [37] | PubChem [36], ZINC [37], ChEMBL [37] | Training data for generative models | Millions to billions of drug-like molecules |
| Molecular Representations | SMILES [42], SELFIES [36], Molecular Graphs [41] | Input formats for generative models | Balance between validity and expressiveness |
| Benchmarking Platforms | GuacaMol [36], MOSES [36], Tartarus [36] | Standardized model evaluation | Comparative performance assessment |
| Property Predictors | Docking simulations [12], Quantum chemical calculations [38] | Evaluation of generated molecules | Physics-based property estimation |
| Experimental Validation | Synthesis & bioassays [12] | Confirmatory testing | Ground truth measurement of activity |
Architectural Workflows
The workflow for VAE-based molecular generation illustrates the sequential process from data preparation to molecule generation:
The GAN-based molecular generation workflow demonstrates the adversarial training process:
The active learning integration workflow demonstrates how generative models improve through iterative refinement:
The comparative analysis of VAEs, GANs, and Transformers for molecular generation reveals a complex landscape where each architectural family offers distinct advantages and limitations. VAE-based approaches provide stable training, interpretable latent spaces, and strong performance in conditional generation tasks, making them particularly suitable for integration with active learning frameworks. GAN-based models can produce high-quality molecular structures but face challenges with training stability and diversity, though hybrid approaches and reinforcement learning integration show promise in addressing these limitations. Transformer architectures excel at capturing long-range dependencies in molecular sequences but may benefit from integration with latent-variable formulations to enhance controllability and novelty.
When combined with active learning paradigms, all three architectures demonstrate enhanced ability to explore novel chemical spaces and generate molecules with optimized properties. The iterative feedback provided by active learning cycles addresses a fundamental limitation of standalone generative models: the poor generalization to chemical spaces beyond the training distribution. As the field progresses, the integration of these architectural approaches with increasingly sophisticated active learning frameworks, multi-objective optimization strategies, and experimental validation pipelines will likely accelerate the discovery of novel therapeutic compounds with tailored properties.
Active Learning (AL) is a machine learning paradigm designed to minimize the cost of data annotation by iteratively selecting the most informative unlabeled data points for expert labeling [44] [45]. In supervised learning, the performance of a classifier is heavily dependent on the quality and quantity of labeled data [46]. However, obtaining labeled samples is often difficult, expensive, and time-consuming, especially in specialized fields like drug discovery [12]. Active learning addresses this challenge by enabling a model to interactively query an oracle (e.g., a human expert) to label new data points with the most valuable true labels [45]. The core component of any active learning system is its query strategy or acquisition function—a method for scoring unlabeled instances based on their potential informativeness to the model [45] [47].
The following diagram illustrates the generic, iterative workflow of an active learning process, common to most query strategies.
This guide provides an objective comparison of the three predominant active learning query strategies—Uncertainty Sampling, Diversity Sampling, and Committee Sampling (Query-by-Committee)—within the context of modern research, particularly focusing on applications in scientific domains such as drug development. We synthesize experimental data and methodologies from recent literature to offer a clear analysis of their performance, strengths, and limitations.
Comparative Analysis of Core Query Strategies
The effectiveness of an active learning query strategy hinges on its ability to accurately identify "informative" data points. The table below summarizes the core principles, metrics, and comparative advantages of the three main strategies.
Table 1: Comparison of Core Active Learning Query Strategies
| Strategy | Core Principle | Key Metrics/Measures | Advantages | Challenges & Limitations | |||||
|---|---|---|---|---|---|---|---|---|---|
| Uncertainty Sampling [44] [45] [47] | Selects data points where the model's prediction is least confident. | - Least Confidence: `1 - P(ŷ | x) [45] [47]<br>- Margin:P(ŷ₁ |
x) - P(ŷ₂ | x) [45] [47]<br>- Entropy:-Σ P(yᵢ |
x) log P(yᵢ | x)` [45] [47] | - Simple and computationally efficient [45]- Directly targets decision boundaries [47] | - Prone to selecting outliers [46]- Can lead to mode collapse (e.g., over-sampling one class like "8" in MNIST) [44]- Ignores data distribution [46] |
| Diversity Sampling [44] [48] | Selects data that are representative of the overall unlabeled data distribution. | - Clustering (e.g., Kernel K-means) [46]- Core-Set Approach [44]- Density-Based Measures [44] | - Improves model generalization [48]- Avoids redundant samples [46]- Explores entire feature space [48] | - May select many already well-understood examples [48]- Requires a good initial data representation [48] | |||||
| Committee Sampling (QBC) [44] [45] [47] | Maintains a committee of models; selects points where committee members disagree the most. | - Vote Entropy [47]- Consensus Entropy [47]- KL Divergence between members [47] | - Reduces individual model bias [48]- Provides a robust measure of uncertainty [47] | - Computationally expensive to train multiple models [48] [47]- Requires maintaining diversity among committee members [47] |
Experimental Protocols and Performance Data
Empirical Results on Standard Benchmarks
Extensive experimental studies have been conducted to evaluate the relative performance of these strategies. A large-scale hyperparameter survey involving over 4.6 million combinations highlighted that the specific implementation of an AL strategy and its hyperparameters significantly impacts performance and reproducibility [49].
A study focused on overcoming the limitations of individual strategies proposed a hybrid framework combining uncertainty, representativeness, and diversity. The experimental protocol on benchmark datasets was as follows:
- Uncertainty Measure: Used the Best-versus-Second-Best (BvSB) approach [46].
- Representativeness Measure: Used Gaussian Processes to evaluate how well a sample represents others in the unlabeled pool [46].
- Information Content: Combined uncertainty and representativity in a weighted product form,
Infor(x_i) = α * Uncertainty(x_i) * Rep(x_i)[46]. - Diversity Measure: Applied kernel k-means clustering to the high-information-content set and selected cluster centers to ensure diversity and minimize redundancy [46].
This hybrid approach demonstrated superior performance compared to state-of-the-art methods that used only one or two criteria, showcasing the power of integrated strategies [46].
Application in Drug Discovery: A Case Study
A compelling application of active learning is in generative AI workflows for drug design, where labeling (e.g., molecular docking or synthesis) is extremely costly. A 2025 study tested a generative model (Variational Autoencoder) with nested active learning cycles on two drug targets, CDK2 and KRAS [12].
Table 2: Experimental Results from the Drug Design Study [12]
| Target | Key Challenge | AL-Generative AI Workflow Outcome | Experimental Validation |
|---|---|---|---|
| CDK2 | Densely populated patent space; need for novel, selective inhibitors. | Generated novel, diverse scaffolds with high predicted affinity and synthesis accessibility. | Of 9 molecules synthesized, 8 showed in vitro activity, with 1 exhibiting nanomolar potency. |
| KRAS | Sparsely populated chemical space; historically difficult to target. | Successfully explored novel chemical space, generating viable molecules distinct from known scaffolds. | Identified 4 molecules with potential activity through in silico methods validated by the CDK2 assay results. |
Detailed Experimental Protocol [12]:
- Model & Representation: A Variational Autoencoder (VAE) was trained on molecules represented as tokenized SMILES strings.
- Nested Active Learning Cycles:
- Inner AL Cycle: Generated molecules were evaluated by chemoinformatic oracles (drug-likeness, synthetic accessibility). Molecules meeting thresholds were used to fine-tune the VAE.
- Outer AL Cycle: After several inner cycles, molecules were evaluated by a physics-based oracle (molecular docking scores). High-scoring molecules were added to a permanent set for VAE fine-tuning.
- Candidate Selection: Promising molecules underwent advanced molecular modeling simulations (e.g., PELE, Absolute Binding Free Energy calculations) before final selection for synthesis and in vitro testing.
This protocol demonstrates a hybrid strategy in practice, using AL to guide a generative model towards molecules that are not only novel and synthesizable (diversity) but also have high predicted target affinity (a specialized form of uncertainty).
The Scientist's Toolkit: Research Reagents & Solutions
The following table details key computational tools and methodologies referenced in the featured experiments, essential for researchers aiming to implement these strategies.
Table 3: Essential Research Reagents and Computational Solutions
| Item / Solution | Function / Description | Relevant Context |
|---|---|---|
| Generative Model (VAE) [12] | A variational autoencoder that learns a continuous latent representation of molecules to generate novel chemical structures. | Core of the drug design workflow; enables exploration of chemical space. |
| Chemoinformatic Oracles [12] | Computational predictors that evaluate generated molecules for drug-likeness, synthetic accessibility, and similarity to known compounds. | Acts as a filter in the inner AL cycle, promoting practical and novel candidates. |
| Physics-Based Oracles (Docking) [12] | Molecular docking simulations that predict the binding affinity and pose of a molecule to a target protein. | Acts as an affinity oracle in the outer AL cycle; provides a more reliable, physics-based estimate of activity. |
| Monte Carlo (MC) Dropout [48] [47] | A technique that uses dropout during inference to simulate an ensemble of models from a single network, providing uncertainty estimates. | A computationally efficient approximation of Bayesian neural networks for uncertainty estimation. |
| Kernel K-means Clustering [46] | A clustering algorithm used to group data points in a high-dimensional feature space, ensuring diversity in selected samples. | Used in hybrid strategies to select diverse, non-redundant samples from a high-information set. |
Hybrid and Advanced Strategies: Integrating Strengths
Given the complementary strengths and weaknesses of the core strategies, modern research heavily favors hybrid approaches [44] [46] [48]. The danger of using only uncertainty sampling, for instance, is the loss of diversity, which can lead to sampling bias and mode collapse, as witnessed in a study on MNIST where the model over-sampled the digit "8" [44]. Similarly, using only diversity sampling may select many uninformative samples from dense regions of already-learned data.
The logical relationship and integration points of these strategies within a hybrid framework are shown below.
The most effective hybrid strategies, as seen in the drug design case study, often combine uncertainty (or a related performance metric like expected loss or docking score) with diversity to select samples that are both challenging for the model and representative of the broader data distribution [44] [12]. For deep learning models, Batch Mode Deep Active Learning (BMDAL) is crucial, as it uses hybrid strategies to select an optimal batch of samples in each cycle, balancing uncertainty and diversity to avoid selecting similar, redundant data points [44].
In summary, no single query strategy is universally superior. The choice depends on the specific application, computational budget, and data characteristics. Uncertainty Sampling is a strong, efficient baseline but risks sampling bias. Diversity Sampling ensures broad coverage but may be inefficient. Committee Sampling offers robust uncertainty estimation at a higher computational cost. Current research and high-impact applications, particularly in demanding fields like drug discovery, demonstrate that hybrid strategies—which systematically combine the principles of uncertainty, diversity, and representativeness—deliver the most robust and effective results, enabling generative AI and active learning to jointly push the boundaries of scientific discovery.
The field of artificial intelligence is undergoing a fundamental transformation, moving beyond static models toward dynamic, self-improving systems. This evolution is characterized by the convergence of two powerful paradigms: architectural breakthroughs in continual learning and pedagogical frameworks for human-AI collaboration. On the architectural front, nested learning has emerged as a revolutionary approach that redefines machine learning models as systems of interconnected, multi-level optimization problems that operate simultaneously at different frequencies [50]. This paradigm directly addresses the critical limitation of catastrophic forgetting that has plagued conventional large language models (LLMs), where learning new tasks sacrifices proficiency on previously acquired knowledge [51].
Parallel to these architectural advances, research in active learning has demonstrated how generative AI tools can function as cognitive partners within structured educational frameworks, particularly in resource-constrained environments [52]. The integration of these approaches—nesting active learning cycles within continually optimizing AI architectures—creates a powerful framework for guiding generative AI in complex domains like drug discovery and clinical pharmacy. This synthesis represents more than incremental improvement; it constitutes a fundamental shift from models as frozen repositories of knowledge to dynamic systems capable of continuous, self-directed improvement through structured interaction [50] [52] [51].
Performance Comparison: Evaluating AI Systems Across Domains
Clinical Pharmacy Competency Assessment
Recent research has quantitatively evaluated the performance of mainstream generative AI systems across core clinical pharmacy scenarios using a multidimensional framework. A 2025 study tested eight AI systems using 48 clinically validated questions across medication consultation, medication education, prescription review, and case analysis with pharmaceutical care scenarios [53]. The evaluation employed six clinical pharmacists with ≥5 years of experience who conducted double-blind scoring across six dimensions: accuracy, rigor, applicability, logical coherence, conciseness, and universality, with scores from 0-10 based on predefined criteria [53].
Table 1: Clinical Pharmacy Performance Across AI Systems
| AI System | Medication Consultation | Medication Education | Prescription Review | Case Analysis | Overall Composite |
|---|---|---|---|---|---|
| DeepSeek-R1 | 9.4 (SD 1.0) | 9.2 (SD 1.1) | 9.3 (SD 1.0) | 9.3 (SD 1.0) | 9.3 |
| Claude-3.5-Sonnet | 8.7 (SD 1.2) | 8.5 (SD 1.3) | 8.8 (SD 1.2) | 8.6 (SD 1.3) | 8.7 |
| GPT-4o | 8.5 (SD 1.3) | 8.3 (SD 1.4) | 8.4 (SD 1.3) | 8.2 (SD 1.4) | 8.4 |
| Gemini-1.5-Pro | 8.4 (SD 1.3) | 8.2 (SD 1.4) | 8.3 (SD 1.3) | 8.1 (SD 1.4) | 8.3 |
| Kimi | 7.9 (SD 1.5) | 7.7 (SD 1.6) | 7.8 (SD 1.5) | 7.6 (SD 1.6) | 7.8 |
| Qwen | 7.7 (SD 1.6) | 7.5 (SD 1.7) | 7.6 (SD 1.6) | 7.4 (SD 1.7) | 7.6 |
| Doubao | 7.3 (SD 1.7) | 7.1 (SD 1.8) | 7.2 (SD 1.7) | 7.0 (SD 1.8) | 7.2 |
| ERNIE Bot | 6.9 (SD 1.8) | 6.7 (SD 1.9) | 6.8 (SD 1.8) | 6.8 (SD 1.5) | 6.8 |
DeepSeek-R1 achieved the highest overall performance, significantly outperforming other models in complex clinical tasks (P<.05) [53]. The study revealed critical limitations across models, including high-risk decision errors where 75% of systems omitted critical contraindications and 90% erroneously recommended macrolides for drug-resistant Mycoplasma pneumoniae in China's high-resistance setting, demonstrating inadequate localization of knowledge [53].
Educational Outcomes with Active Learning Integration
In educational contexts, the integration of generative AI within active learning frameworks has demonstrated significant quantitative benefits. A 15-week quasi-experimental study with 148 undergraduate students compared an experimental group using AI tools within a cognitive partnership model against a control group receiving traditional teacher-centered instruction [52].
Table 2: Educational Outcomes with AI-Augmented Active Learning
| Performance Metric | Experimental Group (AI + Active Learning) | Control Group (Traditional Instruction) | Statistical Significance |
|---|---|---|---|
| Writing Performance Improvement | Z = -6.325, p < .001 | Z = -2.128, p = 0.033 | P < .001 |
| Skill Progression Timeline | 6-8 weeks for notable emergence | 12+ weeks for comparable gains | N/A |
| Positive Perception of Learning Tool | 79.7% of participants | 42% of participants | P < .001 |
| Intentions for Continued Use | 86% of participants | 51% of participants | P < .001 |
| Development of Ethical Awareness | 72% demonstrated significant growth | 38% demonstrated significant growth | P < .001 |
The quantitative results clearly show that the experimental group achieved statistically significant improvements in writing performance compared to modest gains in the control group, with notable skill progression emerging after 6-8 weeks of intervention [52]. Qualitative analysis revealed that AI tools successfully functioned as cognitive partners, metacognitive mirrors, and equity tools in these resource-constrained contexts [52].
Experimental Protocols and Methodologies
Nested Learning Architecture Implementation
Google's Nested Learning paradigm represents a fundamental rethinking of machine learning architecture, treating models not as monolithic entities but as systems of interconnected, multi-level optimization problems [50]. The proof-of-concept implementation, known as the "Hope" architecture, serves as a self-modifying recurrent architecture that leverages continuum memory systems (CMS) to scale to larger context windows [50].
Core Experimental Protocol:
Architecture Design: Hope is implemented as a variant of the Titans architecture with augmented CMS blocks to enable unbounded levels of in-context learning [50]. The system employs multi-time-scale updates where different components update at carefully controlled frequencies, creating a spectrum of memory modules from short-term to long-term retention [50].
Continuum Memory System: The CMS establishes a memory spectrum where each module updates at a specific frequency rate, creating a much richer and more effective memory system for continual learning compared to standard Transformers, where sequence models act as short-term memory and feedforward networks as long-term memory [50].
Deep Optimizers: The nested learning perspective reformulates optimizers as associative memory modules, changing the underlying objective from simple dot-product similarity to standard loss metrics like L2 regression loss, making them more resilient to imperfect data [50].
Validation Framework: Experiments evaluated the architecture on language modeling, long-context reasoning, continual learning, and knowledge incorporation tasks, using common benchmarks and public language modeling tasks to assess perplexity and accuracy against modern recurrent models and standard transformers [50].
Clinical Pharmacy Evaluation Methodology
The comparative analysis of generative AI systems in clinical pharmacy employed a rigorous methodological framework [53]:
Experimental Design:
- Question Selection: 48 clinically validated questions selected via stratified sampling from real-world sources including hospital consultations, clinical case banks, and national pharmacist training databases [53].
- Stratified Sampling: Questions covered medication consultation (20 questions), medication education (10 questions), prescription review (10 questions), and case analysis with pharmaceutical care (8 questions) [53].
- AI Systems Tested: Eight different generative AI systems (ERNIE Bot, Doubao, Kimi, Qwen, GPT-4o, Gemini-1.5-Pro, Claude-3.5-Sonnet, and DeepSeek-R1) were tested using standardized prompts within a single day (February 20, 2025) to ensure consistency [53].
- Evaluation Framework: Six experienced clinical pharmacists (≥5 years experience) conducted double-blind scoring across six dimensions: accuracy, rigor, applicability, logical coherence, conciseness, and universality, scored 0-10 per predefined criteria [53].
Standardized Prompting Instructions: All questions were input to models using a standardized format with the core instruction template: "Act in the role of a clinical pharmacist. Based on the latest clinical guidelines and evidence-based principles, answer the following question." For prescription review tasks, additional emphasis was added: "Determine whether this prescription contains errors and provide your rationale." For case analysis, the prompt specified: "Analyze the pharmacotherapy plan for this case and develop a pharmaceutical care plan addressing Indication, Efficacy, Safety, and Adherence" [53].
Active Learning-GenAI Synergy Framework
The educational implementation followed a carefully structured protocol [52]:
Research Design:
- Participants: 148 undergraduate students from Bahauddin Zakariya University, Pakistan, divided into experimental and control groups.
- Intervention Duration: 15-week structured intervention.
- Experimental Group Protocol: Utilized AI tools (ChatGPT, Claude AI, Meta AI, and Canva) within a cognitive partnership model grounded in Kolb's experiential learning theory and active learning principles [52].
- Control Group Protocol: Received traditional teacher-centered instruction without AI integration.
- Data Collection: Quantitative data on writing performance was supplemented by qualitative data from semi-structured interviews to capture student experiences and ethical awareness development [52].
Theoretical Framework: The study operationalized active learning through Bonwell and Eison's framework, emphasizing that students must engage in "meaningful learning activities and think about what they are doing" [52]. This was implemented through Kolb's experiential learning cycle: (1) draft with AI (concrete experience), (2) critique AI outputs (reflective observation), (3) extract principles (abstract conceptualization), and (4) revise iteratively (active experimentation) [52].
Visualization: Nested Active Learning Architecture
The following diagram illustrates the integrated architecture of nested learning cycles combined with active learning principles, showing how different components interact across multiple time scales and learning dimensions.
Nested Active Learning Architecture
This architecture demonstrates how the Hope model implements a self-modifying system with infinite, looped learning levels [50] [51]. The fast-updating external layer handles immediate user interactions, the intermediate layer processes and integrates knowledge, while the slow-updating core architecture layer performs meta-learning and long-term consolidation, effectively preventing catastrophic forgetting through multi-time-scale optimization [50].
The Scientist's Toolkit: Research Reagent Solutions
Implementing nested active learning cycles for generative AI requires specific methodological tools and frameworks. The following table details essential research reagents and their functions in experimental protocols.
Table 3: Essential Research Reagents for Nested Active Learning Experiments
| Research Reagent | Function | Example Implementation | Experimental Purpose |
|---|---|---|---|
| Continuum Memory System (CMS) | Creates spectrum of memory modules updating at different frequencies | Hope architecture variant of Titans [50] | Prevents catastrophic forgetting; enables multi-timescale learning |
| Deep Optimizers | Reformulates optimizers as associative memory modules | L2 regression loss instead of dot-product similarity [50] | Increased resilience to imperfect data |
| Standardized Clinical Evaluation Matrix | Six-dimension scoring framework for clinical competency | Accuracy, rigor, applicability, logical coherence, conciseness, universality (0-10 scale) [53] | Quantitative comparison of AI system performance in clinical contexts |
| Cognitive Partnership Model | Structured framework for human-AI collaboration in learning | Kolb's experiential learning cycle with AI tools [52] | Implements active learning principles with AI as metacognitive mirror |
| Multi-Time-Scale Update Mechanism | Controls learning rates across different architectural components | Fast outer layers, slow inner layers [50] [51] | Enables continual learning without catastrophic forgetting |
| Double-Blind Scoring Protocol | Independent evaluation by domain experts | Six clinical pharmacists scoring AI responses [53] | Ensures objective assessment of AI performance |
| Active Learning-GenAI Synergy Framework | Pedagogical structure for ethical AI integration | 15-week intervention with cognitive partnership model [52] | Promotes critical engagement and reflective practice |
| Self-Modifying Architecture | Enables recursive meta-learning | Hope model with infinite, looped learning levels [50] [51] | Implements learning-to-learn capabilities |
Discussion: Implications for Drug Discovery and Clinical Applications
The integration of nested learning architectures with active learning cycles has profound implications for high-stakes domains like drug discovery and clinical decision support. In pharmaceutical research, generative AI is already demonstrating remarkable efficiency gains, reducing early drug discovery timelines from 4-7 years to 13-18 months in cases like Insilico Medicine's AI-driven pipeline [54]. The nested learning approach addresses critical limitations in current AI systems identified in clinical evaluations, including the handling of complex reasoning scenarios and localization of medical knowledge [53].
The continuum memory system inherent in nested learning architectures directly tackles the challenge of knowledge currency in clinical applications. Unlike current LLMs with static long-term knowledge and active short-term context but no intermediate learning capability [51], nested systems maintain a spectrum of memory modules that update at different frequencies. This enables appropriate knowledge retention and updating—a critical capability for integrating the latest clinical guidelines and research findings into AI-assisted decision support systems.
Furthermore, the active learning component ensures that human expertise remains integral to the system, addressing the identified limitations in complex clinical reasoning where even advanced systems like Claude-3.5-Sonnet struggled with detecting contradictions like gender-diagnosis mismatches (e.g., prostatic hyperplasia in female patients) [53]. The nested active learning framework creates a structured approach for human-AI collaboration that leverages the strengths of both human clinical expertise and AI's computational capabilities.
The convergence of nested learning architectures and active learning frameworks represents a significant advancement in artificial intelligence, particularly for scientific and clinical applications. The experimental data demonstrates that neither architectural innovation nor pedagogical structure alone is sufficient; rather, their integration creates systems capable of continuous improvement while maintaining alignment with human expertise and ethical frameworks.
The performance comparisons reveal substantial variation in current AI capabilities across clinical domains, with DeepSeek-R1 achieving superior performance in complex clinical tasks while other systems demonstrated significant limitations in safety-critical areas like contraindication identification [53]. These findings underscore the importance of rigorous, domain-specific evaluation alongside architectural innovation.
As the field progresses, the integration of nested learning principles with active learning cycles provides a roadmap for developing AI systems that can adapt to new information while preserving existing knowledge, engage in meaningful collaboration with human experts, and ultimately transform fields like drug discovery and clinical medicine through continuous, guided improvement. The future of generative AI lies not in increasingly larger static models, but in dynamically learning systems that grow through structured interaction with both data and human expertise.
The integration of artificial intelligence (AI) into pharmaceutical research represents a paradigm shift from traditional, labor-intensive drug discovery toward data-driven, automated approaches. This transformation is marked by the emergence of distinct technological philosophies, exemplified by Insilico Medicine and Exscientia. While both companies leverage AI to dramatically compress development timelines and reduce costs, their core methodologies differ significantly. Insilico Medicine has pioneered an end-to-end generative AI approach, using deep learning to simultaneously invent novel biological targets and therapeutic molecules. By contrast, Exscientia's platform embodies an active learning paradigm, creating a tight, iterative "Design-Make-Test-Analyze" loop that augments human expertise with algorithmic precision. This case study provides a structured comparison of their platforms, technologies, and validated performance metrics, offering researchers a clear perspective on how these leading AI strategies are reshaping the drug discovery landscape.
Platform Architecture & Core Technologies
The fundamental architectures of Insilico Medicine's and Exscientia's platforms reveal their distinct approaches to integrating AI into the drug discovery workflow.
Insilico Medicine's Pharma.AI: An End-to-End Generative Engine
Insilico's platform operates as a connected, generative system that covers the entire preclinical journey from target identification to candidate nomination [55] [56].
- PandaOmics: This target discovery engine utilizes natural language processing (NLP) to analyze millions of data files, including research publications, grants, patents, and clinical trials. It combines this with multi-omics data analysis (transcriptomics, proteomics, epigenetics) and a disease-focused knowledge graph to identify and prioritize novel drug targets. It provides AI-driven scores for target confidence, commercial tractability, druggability, and mechanism clarity [57] [55].
- Chemistry42: As the cornerstone of Insilico's generative chemistry effort, this platform employs an ensemble of generative models—including generative adversarial networks (GANs) and reinforcement learning—for de novo molecular design. It is notable for integrating these AI models with physics-based methods for binding affinity prediction and synthetic accessibility scoring. The platform can generate thousands of novel molecule candidates in a matter of days [57] [55] [56].
- InClinico: This module is designed to predict the probability of success for clinical trials, completing the end-to-end pipeline by informing go/no-go decisions before significant clinical investment [58].
Exscientia's Centaur Platform: Active Learning in a Closed Loop
Exscientia’s platform is built around the "Centaur" model, which strategically combines human expertise with AI to drive an iterative design cycle [11].
- Precision Design: Exscientia works backward from patient needs to define a Target Product Profile (TPP) that outlines the complex properties required for an effective medicine. Its AI algorithms then generate panels of potential drug candidates that meet these precise criteria [59].
- Automated Experimentation: A key differentiator is the integration of AI-driven design with wet-lab automation. The company's "AutomationStudio" uses robotic systems to synthesize and test AI-designed compounds, enabling 24/7 operation with minimal human intervention. This creates a high-throughput, data-generating feedback loop [59] [11].
- Active Learning: The platform uses active learning algorithms to help expert designers select the most informative compounds for synthesis. These compounds are chosen specifically because their results will refine the AI models for future design cycles, creating a continuous learning system that becomes more efficient with each iteration [59].
Table 1: Core Technology Comparison
| Feature | Insilico Medicine | Exscientia |
|---|---|---|
| Core AI Philosophy | End-to-end generative AI | Active learning & human-AI collaboration ("Centaur") |
| Target Discovery | PandaOmics: AI-driven from multi-omics & text | Patient-first biology; TPP-driven design |
| Molecule Design | Chemistry42: Generative ensemble (GANs, RL) | Deep learning models trained on chemical libraries |
| Key Integration | Connects biology (PandaOmics) & chemistry (Chemistry42) | Integrates AI design with automated robotics |
| Automation Focus | Computational generation and prediction | Automated compound synthesis & testing |
Visualizing the Core AI Workflows
The distinct approaches of the two platforms can be visualized as two different workflows. Insilico's is a sequential, generative flow, while Exscientia's is a tight, iterative cycle.
Clinical Validation & Performance Metrics
The ultimate validation of any drug discovery platform is its ability to produce viable clinical candidates. Both companies have demonstrated this capability, achieving significant reductions in time and cost compared to industry averages.
Insilico Medicine's Anti-Fibrotic Program
Insilico's most celebrated success is the development of ISM001-055, a potential treatment for Idiopathic Pulmonary Fibrosis (IPF). This program serves as a landmark proof-of-concept for end-to-end AI-driven discovery [55] [56].
- Timeline and Cost: The program advanced from novel target discovery to Phase I clinical trials in approximately 30 months, at a cost of around $2.6 million for the preclinical phase. This compares to an industry standard of 3-6 years and costs that can exceed $400 million for preclinical development alone [55] [56].
- Experimental Protocol:
- Target Hypothesis: PandaOmics was trained on omics and clinical datasets related to tissue fibrosis and aging. Using the iPANDA algorithm, it performed gene and pathway scoring, deep feature synthesis, and causality inference to reveal 20 potential targets. One novel intracellular target was prioritized [55].
- Molecule Generation: Chemistry42 was used to design a library of small molecules targeting the novel protein. The generative adversarial network (GAN) created drug-like structures, which were then optimized for target inhibition, solubility, ADME properties, and CYP inhibition profile [55].
- Validation: The lead molecule series showed nanomolar (nM) IC50 values in target inhibition assays. In a Bleomycin-induced mouse lung fibrosis model, the compounds demonstrated significant improvement in fibrosis and lung function. A favorable safety profile was confirmed in a 14-day repeated dose range-finding study in mice [55].
Exscientia's Clinical-Stage Candidates
Exscientia has advanced multiple compounds into clinical trials, both in-house and through partnerships, demonstrating the broad applicability of its active learning platform [59] [11].
- DSP-1181: Developed in collaboration with Sumitomo Dainippon Pharma, this molecule for obsessive-compulsive disorder was the first AI-designed drug candidate to enter Phase I trials. It was designed in less than 12 months, a fraction of the typical time [11] [60].
- Efficiency Metrics: Exscientia reports that its platform achieves 70% faster drug design cycles and requires the synthesis of 10 times fewer compounds than the industry average to identify a viable clinical candidate. This drastic reduction in experimental overhead translates directly into cost savings and speed [59].
- Pipeline: The company has advanced eight molecules into clinical trials. Its internal focus includes a CDK7 inhibitor (GTAEXS-617) for solid tumors and an LSD1 inhibitor (EXS-74539), for which an IND was approved and a Phase I trial initiated in early 2024 [11].
Table 2: Quantitative Performance Benchmarks
| Metric | Industry Standard | Insilico Medicine | Exscientia |
|---|---|---|---|
| Preclinical Timeline | 3 - 6 years [55] | ~18-30 months [55] [56] | Up to 70% acceleration [59] |
| Preclinical Cost | ~$430M (out-of-pocket) [55] | ~$2.6M (preclinical) [55] | Not explicitly stated, but significantly reduced |
| Compounds Synthesized | Thousands to millions | 80 molecules for IPF candidate [56] | 10x fewer than industry average [59] |
| Clinical-Stage Molecules | N/A | 6 molecules in trials [59] | 8 molecules in trials [11] |
Experimental & Research Protocols
For researchers seeking to understand or implement similar approaches, the following details the key methodologies and reagents intrinsic to these platforms.
Detailed Experimental Workflows
Insilico's Generative Workflow for ISM001-055 [55]:
- Target Discovery with PandaOmics:
- Input Data: Collated multi-omics datasets (e.g., transcriptomics, proteomics) from fibrotic tissue, annotated by age and sex.
- AI Analysis: Executed the iPANDA algorithm for pathway activity analysis. Applied deep feature synthesis and causality inference to identify critical disease drivers.
- Novelty Assessment: Used an NLP engine to analyze millions of patents, publications, and grants, ranking targets based on novelty and disease association.
- Output: A shortlist of 20 prioritized targets, from which a novel intracellular target was selected.
- Molecule Generation with Chemistry42:
- Generative Process: The GAN-based ensemble generated novel molecular structures in SMILES format, optimized against the target binding pocket.
- Multi-parameter Optimization: Molecules were scored and filtered based on predicted potency (IC50), solubility, ADMET properties, and synthetic accessibility using the ReRSA score.
- Output: A batch of ~80 molecules was synthesized and tested, leading to the identification of the ISM001 series.
Exscientia's Active Learning Cycle [59] [11]:
- Precision Design:
- Input: A precisely defined Target Product Profile (TPP) specifying potency, selectivity, and ADME requirements.
- AI Design: Deep learning models propose candidate structures that satisfy the TPP constraints.
- Automated Synthesis & Testing:
- Make: Approved compound designs are sent to the AutomationStudio, where robotic systems execute the chemical synthesis.
- Test: The synthesized compounds are automatically subjected to high-throughput biological assays (e.g., binding affinity, cellular activity) and phenotypic screening on patient-derived samples.
- Learn & Iterate:
- Data Analysis: All experimental results are fed back into the AI models.
- Active Learning: Algorithms identify which compound designs will provide the most informative data to refine the model, guiding the next design cycle.
The Scientist's Toolkit: Key Research Reagents & Solutions
The following table details essential components used in the featured experiments and their functions.
Table 3: Essential Research Reagents and Materials
| Reagent / Solution | Function in the Protocol | Context of Use |
|---|---|---|
| Multi-omics Datasets (Transcriptomics, Proteomics) | Provides the biological data foundation for AI-driven target identification and hypothesis generation. | Used by Insilico's PandaOmics to train models and identify disease-relevant pathways [57] [55]. |
| Patient-Derived Tissue Samples | Enables ex vivo testing of drug candidates in a more clinically relevant human model. | Used by Exscientia (via Allcyte acquisition) for high-content phenotypic screening of AI-designed compounds [11]. |
| Bleomycin | An agent used to induce pulmonary fibrosis in mouse models, creating a representative in vivo system for testing drug efficacy. | Used by Insilico in the Bleomycin-induced mouse lung fibrosis model to validate the anti-fibrotic effect of ISM001-055 [55]. |
| Curated Chemical Building Block Libraries | Provides a set of readily available, synthesizable chemical fragments for the AI to use in constructing novel molecules. | Integral to both platforms; Exscientia's retrosynthesis AI uses them for route planning, and Insilico's Chemistry42 uses them for synthetic accessibility scoring [59] [57]. |
| High-Throughput Screening Assays | Automated biological tests that rapidly measure the activity of thousands of compounds against a target or phenotype. | Used in Exscientia's automated testing loop and for validating hits generated by platforms like Atomwise [61] [11]. |
Discussion: Generative AI vs. Active Learning in Context
The successes of Insilico Medicine and Exscientia provide a robust framework for comparing generative AI and active learning approaches within the broader thesis of AI-driven drug discovery.
Generative AI (Insilico): This approach excels at exploration and novelty. It is designed to venture into vast, uncharted chemical and biological spaces to invent completely new targets and molecular structures from scratch. Its strength lies in its ability to form deep, non-intuitive connections across disparate data types (biology and chemistry), making it particularly powerful for tackling diseases with poorly understood mechanisms or where no known targets exist. The primary validation of this approach is the creation of a novel target and a novel molecule that successfully enters clinical trials [55] [56].
Active Learning (Exscientia): This paradigm excels at optimization and efficiency. It is engineered to make the most intelligent use of experimental resources by iteratively refining a search towards a predefined goal (the TPP). The "Centaur" model leverages the pattern-finding power of AI while retaining the strategic oversight of human experts. This is highly effective for optimizing known target classes, improving drug properties (e.g., selectivity, pharmacokinetics), and systematically reducing uncertainty with each experimental cycle. Its success is validated by its consistent ability to accelerate the design process and reduce the number of compounds needed to reach a candidate [59] [11].
In conclusion, the choice between these two powerful AI philosophies is not a matter of superiority, but of strategic alignment with research goals. Insilico Medicine's generative approach offers a path to groundbreaking, first-in-class therapies for complex diseases, while Exscientia's active learning platform provides a robust engine for efficiently creating best-in-class drugs and optimizing known modalities. As the field evolves, the integration of these complementary paradigms—generative exploration followed by active learning optimization—may well represent the future of AI-accelerated drug development.
The integration of artificial intelligence (AI) is fundamentally reshaping the pharmaceutical research and development landscape. Two particularly powerful approaches, generative AI and active learning, are demonstrating significant potential to accelerate discovery timelines, reduce costs, and increase the probability of success [62] [63]. While both are machine learning techniques, they serve distinct and complementary roles. Generative AI specializes in the de novo creation of novel molecular structures and the prediction of complex properties [64] [65]. In contrast, active learning is an iterative, data-efficient framework that intelligently selects the most informative experiments to perform, thereby optimizing resource allocation in both virtual and physical screening campaigns [33].
The application of these technologies spans the critical early stages of drug discovery. In target identification, AI models mine vast genomic and multi-omic datasets to pinpoint novel, druggable disease targets [62] [65]. For virtual screening, they enable the rapid evaluation of millions of compounds, far surpassing the throughput of traditional physical methods [66] [63]. In drug repurposing, AI algorithms find new therapeutic uses for existing medicines by analyzing molecular mechanisms, disease biology, and clinical outcomes [62]. This guide provides a comparative analysis of generative AI and active learning across these domains, supported by performance data and experimental protocols.
Performance Comparison: Quantitative Data
The quantitative impact of AI-driven approaches is evident across key performance metrics, from clinical success rates to the efficiency of screening campaigns. The table below summarizes comparative performance data.
Table 1: Performance Metrics of AI-Driven Drug Discovery
| Metric | Traditional Approach | AI-Driven Approach | Data Source / Context |
|---|---|---|---|
| Phase I Trial Success Rate | 40-65% | 80-90% | AI-designed small molecules [62] [65] |
| Discovery Timeline (Target to Candidate) | ~5 years | 1.5 - 2.5 years | e.g., Insilico Medicine's ISM001-055 [11] [62] |
| Active Learning Screening Efficiency | Exhaustive search required | Discovers 60% of synergistic pairs with only 10% of experiments | Drug combination screening [33] |
| Lead Optimization Cycles | 4-6 years | 1-2 years | AI-powered predictive modeling and virtual screening [65] |
| AI-Designed Molecules to Clinical Stages | Nearly 0 (pre-2020) | >75 molecules by end of 2024 | Clinical-stage AI candidates [11] |
Clinical Pipeline and Success Rates
AI-designed drug candidates are progressing through clinical trials with notable efficiency and success. As of 2024, over 75 AI-derived molecules had reached clinical stages, a remarkable leap from nearly zero just a few years prior [11]. An industry analysis noted that these AI-assisted candidates achieve Phase I success rates of 80-90%, substantially higher than the industry average of 40-65% [62] [65]. This high success rate is attributed to better candidate selection through predictive analytics and rigorous in silico validation [62]. Notable breakthroughs include Insilico Medicine's ISM001-055 for idiopathic pulmonary fibrosis, which advanced from target to Phase I trials in approximately 18 months, and Schrödinger's TYK2 inhibitor, zasocitinib, which has progressed to Phase III trials [11] [62].
Screening and Experimental Efficiency
Active learning dramatically improves the efficiency of resource-intensive screening processes. In the context of synergistic drug combination screening—where positive hits are rare—active learning can identify 60% of all synergistic drug pairs by testing only 10% of the total combinatorial space [33]. This represents an 82% reduction in the experimental burden (from 8,253 measurements to 1,488) to find 300 synergistic combinations [33]. This data-efficient approach is crucial for navigating large and complex search spaces where exhaustive screening is prohibitively expensive and time-consuming.
Experimental Protocols and Methodologies
Protocol for Active Learning in Drug Synergy Screening
This protocol, based on the work of [33], outlines the steps for implementing an active learning cycle to discover synergistic drug combinations.
Table 2: Key Research Reagents for Active Learning in Drug Synergy
| Research Reagent / Tool | Function in the Protocol |
|---|---|
| Pre-existing Synergy Dataset (e.g., O'Neil, ALMANAC) | Provides initial data for pre-training the AI model. Contains drug pairs, cell lines, and measured synergy scores. |
| AI Prediction Model (e.g., MLP, GCN, Transformer) | The core algorithm that predicts synergy scores for untested drug pairs based on molecular and cellular features. |
| Molecular Features (e.g., Morgan Fingerprints, MAP4) | Numerical representations of the chemical structure of each drug, used as input for the AI model. |
| Cellular Features (e.g., Gene Expression Profiles from GDSC) | Numerical representations of the genomic context of the target cell line, used as input for the AI model. |
| Selection Criteria / Acquisition Function | The algorithm that prioritizes which drug pairs to test next based on the model's predictions (e.g., highest predicted synergy, greatest uncertainty). |
| High-Throughput Screening Platform | Automated laboratory system to physically test the selected drug combinations and generate new ground-truth synergy data. |
Workflow Steps:
- Initialization: Pre-train a synergy prediction model (e.g., a Multi-Layer Perceptron or MLP) on a publicly available dataset like O'Neil. The model uses molecular features (Morgan fingerprints) and cellular features (gene expression profiles) as input.
- First Batch Selection: Use the pre-trained model to predict synergy for all untested drug pairs in your library. The acquisition function selects the first batch (e.g., 96 or 384) of combinations with the highest predicted synergy.
- Experimental Testing: The selected drug combinations are tested experimentally using a high-throughput screening platform to obtain actual synergy scores (e.g., LOEWE score).
- Model Retraining: The new experimental data is added to the training set, and the AI model is retrained (fine-tuned) on this expanded dataset.
- Iteration: Steps 2-4 are repeated for multiple batches. With each iteration, the model becomes more accurate for the specific experimental context, allowing it to better guide the search towards the most promising regions of the chemical and biological space.
- Key Parameters: Batch size is a critical parameter; smaller batches allow for more dynamic exploration and can yield a higher synergy hit rate [33].
Protocol for Generative AI inDe NovoMolecule Design
This protocol describes the process of using generative AI for designing novel drug candidates against a validated target.
Workflow Steps:
- Target Specification and Property Profiling: Define the target product profile, including the 3D structure of the protein target (potentially from AlphaFold 3), desired binding affinity, selectivity, and optimal ADMET (Absorption, Distribution, Metabolism, Excretion, Toxicity) properties [62] [63].
- Model Sampling and Generation: A generative model, such as a Generative Adversarial Network (GAN) or a language model trained on chemical structures (e.g., SMILES), is used to generate millions of novel molecular structures in silico [64] [65].
- Virtual Screening and Multi-Property Optimization: The generated library is virtually screened against the target. AI models predict the binding affinity, synthesizability, and key ADMET properties for each molecule, filtering out unsuitable candidates [62] [63].
- Candidate Selection and Synthesis: A shortlist of the most promising candidates, often just a few dozen compounds, is selected for synthesis. Companies like Exscientia report requiring 10 times fewer synthesized compounds than industry norms using this approach [11] [62].
- Experimental Validation: The synthesized compounds undergo in vitro testing to confirm biological activity and desired properties, creating a closed feedback loop to further refine the generative models.
Diagram 1: Generative AI de novo design workflow.
Comparative Analysis: Application Spectrum
Target Identification
- Generative AI employs large-scale knowledge graphs that integrate disparate data from genomics, proteomics, and clinical records to uncover novel disease-associated targets and biological pathways that are difficult to identify through manual research [62]. For example, researchers at the Oxford Drug Discovery Institute used AI to evaluate 54 immune-related genes as potential Alzheimer's disease targets in days, a process that traditionally took weeks [62].
- Active Learning is less directly applied to initial target discovery. Its role becomes critical in the subsequent target validation phase, where it can optimize experimental designs to efficiently confirm the biological and therapeutic relevance of AI-prioritized targets with minimal resource expenditure.
Virtual Screening
- Generative AI acts as a powerful engine for creating massive virtual compound libraries from scratch. It can also power the scoring functions used in virtual screening to predict ligand-protein binding affinity, though these functions still face challenges with accuracy and false positives [66] [63].
- Active Learning transforms virtual screening from a static, one-step process into a dynamic, iterative cycle. It addresses the limitations of static scoring functions by sequentially selecting the most informative compounds for more accurate (and often more costly) simulation methods or for experimental testing, thereby maximizing the value of each simulation or experiment [33].
Diagram 2: Active learning iterative screening cycle.
Drug Repurposing
- Generative AI excels in drug repurposing by using large-scale models to integrate molecular mechanism data, disease biology, and clinical outcomes. This allows it to match existing compounds to new therapeutic indications that may have been overlooked during their initial development [62]. Startups in this space aim to reduce repurposing timelines to under two years and costs below $1 million [62].
- Active Learning has a more niche application in repurposing. It could be deployed to optimize the experimental screening of existing drug libraries against a new disease model, efficiently identifying the most promising repurposing candidates for further validation.
Integrated Workflow and Future Outlook
The most powerful applications of AI in drug discovery emerge when generative AI and active learning are used in tandem. A synergistic workflow can be established where generative AI designs a vast and diverse set of novel candidates, and active learning guides their experimental validation in an efficient, iterative loop. This creates a closed "design-make-test-analyze" cycle that continuously improves the AI models and accelerates the discovery process [11] [33].
The future of AI in drug discovery will see broader adoption of foundation models trained on multimodal biomedical data and the development of more autonomous AI-driven laboratory systems [62]. However, challenges remain, including the need for high-quality, unbiased data, the "black box" nature of some complex models, and the evolution of regulatory frameworks to assess AI-generated therapeutics [62] [63]. Despite these hurdles, the evidence is clear: AI has moved from a speculative tool to a core technology that is fundamentally reshaping the speed, cost, and success of bringing new medicines to patients.
Navigating Challenges: Data, Translation, and Implementation Hurdles
The integration of artificial intelligence (AI) into drug discovery represents a paradigm shift from traditional serendipitous methods toward engineered therapeutic design. Despite accelerated in-silico success, a persistent translation gap remains between computational promise and demonstrated clinical efficacy. This gap manifests when AI-designed molecules with excellent predicted properties fail in biological assays or human trials due to unanticipated complexities of human physiology, disease heterogeneity, or insufficient pharmacokinetic profiles. The pharmaceutical industry faces Eroom's Law (Moore's Law backward), where drug development becomes slower and more expensive over time despite technological advances, with a 90% failure rate once candidates enter clinical trials and costs exceeding $2 billion per approved drug [67].
Two dominant computational approaches have emerged to address this challenge: generative AI, which creates novel molecular structures with desired properties, and active learning (AL), which strategically selects experiments to maximize knowledge gain. This guide provides a comparative analysis of these methodologies, examining their respective capabilities to generate clinically viable drug candidates through examination of experimental protocols, performance metrics, and clinical translation success rates. Understanding the relative strengths and limitations of each approach enables researchers to make informed decisions about platform selection and implementation strategy for bridging the critical translation gap.
Comparative Framework: Generative AI vs. Active Learning
Core Definitions and Methodological Differences
Generative AI in drug discovery involves machine learning models that learn the underlying patterns and relationships in molecular data to generate novel chemical structures with optimized properties. These models include variational autoencoders (VAEs), generative adversarial networks (GANs), diffusion models, and transformer-based architectures. They operate on the "describe first then design" inverse paradigm, creating molecules tailored to specific target profiles rather than screening existing libraries [12]. These systems can design novel molecular scaffolds unseen in training data, significantly expanding explorable chemical space.
Active Learning represents a strategic framework where algorithms selectively choose the most informative data points for experimental validation, creating an iterative feedback loop that improves model performance with minimal resource expenditure. Unlike traditional approaches that test the most promising candidates in each round, AL prioritizes samples based on their potential to improve model understanding, often focusing on regions of high uncertainty or diversity [68]. This approach is particularly valuable in drug discovery where experimental resources are limited and the chemical space is enormous.
Key Technical Implementations
Generative AI Platforms have demonstrated substantial clinical progress. Leading platforms include:
- Insilico Medicine's Chemistry42: A generative chemistry engine that combines multiple AI approaches for de novo molecular design, responsible for ISM001-055, the first AI-generated drug to show positive Phase 2a results for idiopathic pulmonary fibrosis [67].
- Exscientia's Centaur Chemist: An end-to-end platform that integrates algorithmic design with human expertise, claiming design cycles ~70% faster and requiring 10× fewer synthesized compounds than industry norms [11].
- Schrödinger's Physics-Enabled Platform: Combines machine learning with physics-based computational methods for molecular modeling, advancing the TYK2 inhibitor zasocitinib into Phase III clinical trials [11].
Active Learning Methodologies have evolved to address specific drug discovery challenges:
- Deep Batch Active Learning: Novel methods like COVDROP and COVLAP use Monte Carlo dropout and Laplace approximation to quantify model uncertainty and select batches that maximize joint entropy, considering both uncertainty and diversity of selected molecules [68].
- VAE with Nested AL Cycles: Integrates a variational autoencoder with two nested active learning cycles that iteratively refine predictions using chemoinformatics and molecular modeling predictors [12].
- Synergistic Drug Combination AL: Frameworks like RECOVER employ active learning to efficiently discover synergistic drug pairs with 5–10× higher hit rates than random selection by dynamically guiding experimental campaigns [33].
Table 1: Core Methodological Differences Between Generative AI and Active Learning Approaches
| Feature | Generative AI | Active Learning |
|---|---|---|
| Primary Objective | Create novel molecular structures with desired properties | Optimize experimental selection to maximize knowledge gain |
| Core Paradigm | "Describe first then design" [12] | Iterative "design-make-test-learn" cycles [68] |
| Key Strength | Exploration of novel chemical space | Efficient resource utilization |
| Data Dependency | Requires large training datasets | Functions effectively in low-data regimes |
| Clinical Validation | Multiple candidates in clinical trials (e.g., Insilico, Exscientia) [11] | Extensive retrospective validation; emerging clinical translation |
Experimental Protocols and Workflows
Generative AI Workflow with Nested Active Learning
A sophisticated implementation combining both approaches integrates a variational autoencoder with nested active learning cycles [12]. This hybrid architecture aims to leverage the strengths of both methodologies while mitigating their individual limitations.
Protocol Steps:
- Data Representation and Initial Training: Represent training molecules as tokenized SMILES strings converted into one-hot encoding vectors. Initially train the VAE on a general molecular dataset to learn viable chemical structures, then fine-tune on a target-specific training set to enhance target engagement.
- Molecule Generation and Inner AL Cycles: Sample the VAE to generate new molecules. Subject chemically valid generated molecules to evaluation by chemoinformatic predictors (drug-likeness, synthetic accessibility, similarity filters). Molecules meeting threshold criteria are added to a temporal-specific set used to fine-tune the VAE in subsequent training iterations.
- Outer AL Cycle with Physics-Based Validation: After set inner cycles, accumulated molecules in the temporal-specific set undergo docking simulations as an affinity oracle. Molecules meeting docking score thresholds transfer to a permanent-specific set for VAE fine-tuning.
- Candidate Selection and Refinement: Apply stringent filtration and selection processes to identify promising candidates from the permanent-specific set. Use intensive molecular modeling simulations (e.g., PELE) for in-depth evaluation of binding interactions and stability within protein-ligand complexes [12].
Diagram 1: Generative AI with nested AL cycles workflow. (Title: AI Drug Discovery Workflow)
Deep Batch Active Learning Protocol for Molecular Optimization
This protocol focuses on optimizing molecular properties through strategic batch selection, particularly valuable for ADMET (Absorption, Distribution, Metabolism, Excretion, Toxicity) and affinity property optimization [68].
Protocol Steps:
- Initial Model Training: Train initial neural network models (typically graph neural networks) on available labeled data for the target property.
- Uncertainty Quantification: Employ multiple methods (MC dropout, Laplace approximation) to compute a covariance matrix between predictions on unlabeled samples, quantifying both uncertainty and diversity.
- Batch Selection: Use a greedy iterative approach to select a submatrix of size B×B from the covariance matrix with maximal determinant, ensuring selected batches maximize information content through diversity and uncertainty.
- Experimental Testing and Model Retraining: Synthesize and test selected batches experimentally, then use the newly acquired data to retrain models, creating an iterative feedback loop that progressively improves model accuracy with minimal experimental effort.
Active Learning for Synergistic Drug Combination Discovery
This specialized protocol addresses the challenge of identifying rare synergistic drug pairs within large combinatorial spaces [33].
Protocol Steps:
- Algorithm Pre-training: Pre-train deep learning models (e.g., RECOVER) on existing drug combination datasets (e.g., Oneil, ALMANAC) using molecular features (Morgan fingerprints) and cellular context features (gene expression profiles).
- Iterative Batch Selection and Testing: Divide the experimental campaign into sequential batches rather than conducting all measurements simultaneously. Use the AI algorithm to select the most informative drug combinations for each batch based on exploration-exploitation strategies.
- Dynamic Model Refinement: Incorporate data from each experimental batch to continuously refine algorithm parameters, enhancing predictive accuracy for subsequent selections.
- Validation and Hit Confirmation: Experimentally confirm predicted synergistic pairs through secondary assays, focusing on pairs with the highest predicted synergy scores and clinical relevance.
Performance Metrics and Comparative Data
Quantitative Performance Comparison
Table 2: Performance Metrics of Generative AI and Active Learning Approaches
| Metric | Generative AI | Active Learning | Hybrid Approaches |
|---|---|---|---|
| Discovery Speed | 18-30 months to Phase 1 (vs. 5-year average) [11] | 5-10× higher hit rates than random selection [33] | 8 out of 9 synthesized molecules showed in vitro activity [12] |
| Success Rates | 80-90% Phase I success rate (vs. ~50% historical) [69] | Discovers 60% of synergistic pairs with 10% combinatorial exploration [33] | Novel scaffolds with high predicted affinity and synthesis accessibility [12] |
| Resource Efficiency | ~70% faster design cycles; 10× fewer synthesized compounds [11] | 82% reduction in experimental materials and time [33] | Significant potential saving in number of experiments needed [68] |
| Clinical Progress | 75+ AI-derived molecules in clinical stages by end of 2024 [11] | Limited direct clinical translation data | 1 molecule with nanomolar potency in CDK2; 4 with potential KRAS activity [12] |
| Limitations | Mixed Phase II results; biology translation challenges [67] | Primarily validated retrospectively; requires initial data | Computational intensity; integration complexity |
Clinical Translation Case Studies
Generative AI Success: Insilico Medicine's ISM001-055, a TNIK inhibitor for idiopathic pulmonary fibrosis, represents the first generative AI-designed drug with positive Phase 2a results. The program progressed from target discovery to Phase 1 in approximately 30 months - roughly half the industry average. In the Phase 2a trial with 71 patients, the high dose (60 mg QD) showed a mean improvement of 98.4 mL in Forced Vital Capacity compared to a decline of -62.3 mL in the placebo group [67].
Generative AI Setback: Recursion Pharmaceuticals' REC-994 for Cerebral Cavernous Malformation was discontinued after long-term extension data failed to show sustained improvements. While the AI correctly identified biological activity in cellular assays, translation to human efficacy in the complex neurological disease proved elusive, highlighting the "translation gap" between cellular models and human physiology [67].
Active Learning Success: For synergistic drug combination discovery, active learning frameworks have demonstrated the ability to identify 300 out of 500 synergistic combinations with only 1,488 measurements - an 82% reduction in experimental time and materials compared to the 8,253 measurements required without strategic selection [33].
Hybrid Approach Success: The VAE with nested AL cycles approach generated novel scaffolds for CDK2 and KRAS targets. For CDK2, 9 molecules were synthesized with 8 showing in vitro activity, including one with nanomolar potency. The approach successfully explored novel chemical spaces while maintaining synthetic accessibility and predicted affinity [12].
The Scientist's Toolkit: Essential Research Reagents and Platforms
Table 3: Key Research Reagent Solutions for AI-Driven Drug Discovery
| Category | Specific Tools/Platforms | Function | Representative Providers |
|---|---|---|---|
| Generative Models | Chemistry42, Centaur Chemist, Molecular Language Models | De novo molecular design with optimized properties | Insilico Medicine, Exscientia, BioGPT |
| Active Learning Frameworks | COVDROP, COVLAP, BAIT, GeneDisco | Batch selection for optimal experiment planning | DeepChem, Custom implementations |
| Property Prediction | Deep neural networks, Graph neural networks, PBPK models | Predict ADMET, affinity, and physicochemical properties | Schrödinger, Atomwise, Insitro |
| Experimental Validation | High-throughput screening, Synthesis automation, Patient-derived models | Biological validation of AI-predicted candidates | Allcyte (patient-derived models), Automated robotics |
| Data Resources | ChEMBL, DrugComb, GDSC, UK Biobank | Training data for AI algorithms with chemical/biological context | Public databases, Proprietary collections |
Integrated Workflow for Optimal Translation
Diagram 2: Integrated AI approach for translation. (Title: Integrated AI Drug Discovery)
The most promising path for bridging the translation gap involves integrating generative AI and active learning into a cohesive workflow that leverages their complementary strengths. This integrated approach follows a sequential process: generative AI creates diverse molecular candidates with desired properties; active learning prioritizes the most informative candidates for synthesis and testing; multi-modal validation incorporates physics-based simulations (docking, PBPK) and complex biological models (organoids, patient-derived samples); and clinical trial simulation uses virtual patient cohorts and in-silico trials to optimize trial design and predict human efficacy [70].
This integrated framework addresses key limitations of either approach used in isolation. While generative AI can rapidly explore vast chemical spaces, it benefits from active learning's strategic guidance for experimental validation. Conversely, active learning's efficiency is enhanced when applied to the novel chemical spaces identified by generative models rather than being constrained to existing compound libraries. The combination creates a virtuous cycle where generative exploration and strategic validation progressively refine candidates toward clinical viability.
The comparative analysis of generative AI and active learning approaches reveals distinct but complementary roles in addressing the translation gap in drug discovery. Generative AI excels at exploring novel chemical space and designing molecules with optimized properties, while active learning provides strategic efficiency in experimental validation. The emerging evidence suggests that integrated approaches leveraging both methodologies show particular promise for generating clinically viable candidates.
Current performance data indicates that AI-discovered molecules show substantially higher Phase I success rates (80-90%) compared to historical averages (~50%), suggesting improved ability to design molecules with drug-like properties [69]. However, Phase II success rates (~40%) remain comparable to industry averages, highlighting the persistent challenges of translating computational predictions to human efficacy.
Future advancements will likely focus on improving biological relevance through incorporation of patient-derived data, enhanced disease models, and more sophisticated simulation of human physiology. As these technologies mature and more clinical data becomes available, the integration of generative AI and active learning represents the most promising path toward fundamentally restructuring drug discovery from an empirical screening process to an engineered therapeutic design discipline, ultimately bridging the translation gap between in-silico promise and clinical reality.
The rapid adoption of artificial intelligence (AI) in scientific research, particularly in drug development, hinges on the ability to create reliable, unbiased, and broadly applicable models. The core challenge lies in the data that fuels these systems: its quality, quantity, and inherent biases directly dictate the utility and trustworthiness of the resulting AI. This guide objectively compares two dominant paradigms for leveraging data in AI-driven science: Generative AI and Active Learning. We frame this comparison within broader research on their respective capabilities and limitations, providing researchers with a data-driven foundation for selecting and implementing these approaches. The performance of any AI model is intrinsically linked to its training data. Biases in data collection, annotation, or selection can produce models that perpetuate stereotypes, yield inaccurate scientific predictions, and fail to generalize beyond their initial training domain. This analysis delves into the methodologies to overcome these challenges, comparing the ability of generative AI and active learning to expand applicability domains while ensuring robustness and fairness.
Performance Comparison: Generative AI vs. Active Learning
A direct comparison of Generative AI and Active Learning requires examining their performance across key metrics relevant to scientific discovery, such as data efficiency, robustness to bias, and capability in expanding to new domains. The following table synthesizes findings from recent benchmarks and research publications to provide a quantitative and qualitative comparison. It is critical to note that "Active Learning" in this context often refers to systems that use these strategies to guide data acquisition or model training, which may themselves incorporate generative components.
Table 1: Performance Comparison of Generative AI and Active Learning Approaches
| Performance Metric | Generative AI | Active Learning |
|---|---|---|
| Data Efficiency | Often requires massive datasets (e.g., GPT-4 trained on ~13T tokens) [71]. | Designed for high data efficiency; reduces labeling burden by up to 30% by prioritizing informative samples [2]. |
| Bias Mitigation Capability | Can amplify biases in training data (e.g., demographic, historical); requires explicit strategies like adversarial learning or synthetic data oversampling [72]. | Can potentially compound initial biases if the sampling strategy is not carefully designed; less inherent mitigation than targeted generative techniques. |
| Domain Expansion | Excels at generating data for novel domains via fine-tuning or prompt-based steering; enables exploration of chemical spaces beyond training data [30]. | Systematically expands the domain by querying the most uncertain or model-informative regions of the input space. |
| Benchmark Performance (MMLU) | State-of-the-art models (e.g., Gemini 2.5) achieve scores upwards of 89% on specific tasks [71]. | Performance is highly dependent on the base model and query strategy; can achieve superior performance with fewer data points compared to passive learning. |
| Real-World Task Success | Success rates vary; e.g., on real freelance coding tasks (SWE-Lancer), top models succeed ~26.2% of the time, highlighting the benchmark-to-reality gap [71]. | In educational settings, AI-enhanced active learning programs can improve student test scores by 54% compared to traditional methods [2]. |
| Computational Cost | Very high training and inference costs; cloud compute can run into tens of thousands of dollars monthly [73]. | Lower overall data labeling costs, but requires iterative model retraining and a robust querying infrastructure. |
Detailed Experimental Protocols
To validate and compare the approaches outlined above, researchers employ rigorous experimental protocols. The following sections detail the methodologies for two key types of experiments cited in the performance comparison.
Protocol for Bias Mitigation in Generative AI
This protocol is based on established methods from the literature for identifying and countering biases in generative models [72].
1. Hypothesis: Implementing a Fairness-Aware Adversarial Perturbation (FAAP) framework will significantly reduce the detectability of protected attributes (e.g., race, gender) in a generative model's latent representations, thereby mitigating bias in its outputs.
2. Materials & Setup:
- Model: A deployed generative model (e.g., a text or image generator) where internal parameters are inaccessible (black-box).
- Datasets: Standard benchmarks known to contain societal biases (e.g., CVs with gendered names, image datasets with occupational stereotypes).
- Infrastructure: High-performance computing cluster with multiple GPUs for adversarial training.
3. Procedure:
- Step 1 - Bias Audit: First, a comprehensive audit is conducted to establish a baseline. The model generates content based on neutral prompts related to sensitive concepts (e.g., "a CEO," "a nurse"). The outputs are analyzed for skewed distributions of protected attributes using predefined metrics.
- Step 2 - Adversarial Framework Setup: The FAAP framework is implemented. It consists of two components:
- A Generator: This network learns to create minimal perturbations that are added to the input data.
- A Discriminator: This network is trained to identify protected attributes from the model's internal latent representations after the perturbed input is processed.
- Step 3 - Adversarial Training: The generator and discriminator are trained in a competitive loop. The generator's objective is to produce perturbations that "fool" the discriminator, making the protected attribute undetectable. The discriminator continuously improves its detection capability. This process shapes the model's latent space to be invariant to the protected attributes.
- Step 4 - Evaluation: The model's outputs generated from perturbed inputs are compared against the baseline audit. Reduction in stereotype reinforcement and equalization of output distributions across demographic groups are measured using quantitative fairness metrics (e.g., demographic parity, equality of opportunity).
Protocol for Active Learning in a Scientific Domain
This protocol outlines a general approach for using active learning to efficiently build a predictive model in a data-scarce environment, such as predicting compound activity in drug discovery.
1. Hypothesis: An active learning strategy, which strategically selects data points for experimental validation, will achieve a target model performance with significantly fewer labeled examples than a model trained on a randomly selected dataset of the same size.
2. Materials & Setup:
- Initial Dataset: A small "seed" library of compounds with experimentally validated activity labels.
- Unlabeled Pool: A large, diverse library of compounds (e.g., from a virtual chemical space) without activity labels.
- Model: A base predictive model, such as a Random Forest or Graph Neural Network.
- Oracle: An experimental assay (e.g., high-throughput screening) capable of providing the ground-truth label for a compound selected by the active learning system.
3. Procedure:
- Step 1 - Initial Model Training: The base model is trained on the initial seed dataset.
- Step 2 - Iterative Active Learning Loop: The following cycle is repeated until a predefined performance threshold or labeling budget is exhausted:
- a. Prediction & Uncertainty Estimation: The trained model predicts the activity for all compounds in the unlabeled pool. An uncertainty metric (e.g., entropy, least confidence, or Bayesian uncertainty) is calculated for each prediction.
- b. Query Strategy: A query strategy is applied to select the most informative compounds. A common strategy is "uncertainty sampling," where the compounds with the highest prediction uncertainty are selected. More advanced strategies like "query-by-committee" or "expected model change" may also be used.
- c. Oracle Labeling: The top k selected compounds are sent for experimental validation via the "oracle" (the assay), and their true activity labels are obtained.
- d. Dataset Update & Retraining: The newly labeled compounds are removed from the unlabeled pool and added to the training set. The predictive model is then retrained on this augmented dataset.
- Step 3 - Performance Comparison: The performance (e.g., AUC-ROC, precision) of the active learning-trained model is compared against a control model trained on a dataset of the same size but selected at random. The reduction in data required to achieve the same performance level is the key metric of success.
Workflow and Pathway Visualizations
The following diagrams illustrate the core logical workflows for the bias mitigation and active learning protocols described above.
Bias Mitigation with Adversarial Learning
Active Learning Cycle for Scientific Discovery
The Scientist's Toolkit: Research Reagent Solutions
Implementing the experimental protocols for AI comparison requires a suite of computational and physical tools. The following table details key resources essential for this field of research.
Table 2: Essential Research Reagents and Tools for AI Comparison Studies
| Item Name | Function/Brief Explanation | Example Use Case |
|---|---|---|
| Bias Audit Benchmarks | Standardized datasets and metrics (e.g., BOLD, BBQ) designed to systematically evaluate model outputs for stereotypes and discriminatory biases. | Quantifying the baseline level of gender or racial bias in a generative language model before and after applying a mitigation technique. |
| Uncertainty Quantification Library | A software library (e.g., uncertainty-toolbox, laplace-redux) that implements various methods for estimating predictive uncertainty in machine learning models. |
Calculating entropy or confidence intervals for predictions in the active learning loop to identify the most uncertain data points. |
| Chemical Compound Library | A large, diverse collection of chemical structures (e.g., ZINC, Enamine REAL) used as the unlabeled data pool in virtual screening. | Serving as the source of candidate molecules for the active learning protocol in a drug discovery project. |
| High-Throughput Screening Assay | An automated experimental system capable of rapidly testing the biological activity of thousands of compounds. | Acting as the "oracle" in the active learning cycle to provide ground-truth labels for selected compounds. |
| Adversarial Training Framework | A deep learning framework (e.g., PyTorch, TensorFlow) configured with custom loss functions to train competing generator and discriminator networks. | Implementing the Fairness-Aware Adversarial Perturbation (FAAP) protocol for bias mitigation. |
| Cloud Compute Platform | On-demand, scalable computing resources (e.g., AWS, GCP, Azure) essential for training large generative models and running complex simulations. | Providing the GPU clusters needed for training state-of-the-art generative AI models and running large-scale active learning simulations. |
Ensuring Synthetic Accessibility and Drug-Likeness in AI-Generated Molecules
Artificial intelligence (AI) has emerged as a transformative force in small-molecule drug discovery, capable of generating billions of novel molecular structures in silico [74] [75]. However, a significant challenge known as the "generation-synthesis gap" persists: the majority of AI-proposed molecules cannot be practically synthesized in laboratory settings or fail to exhibit necessary drug-like properties [76] [77]. This limitation has prompted the development of sophisticated computational strategies to ensure that AI-generated candidates are both synthetically accessible and possess drug-like characteristics.
The fundamental challenge stems from the fact that generative AI models often prioritize target affinity and potency without inherent knowledge of synthetic chemistry constraints [78]. Consequently, molecules may contain structurally complex or unstable fragments, require impractical multi-step synthetic routes, or exhibit poor pharmacokinetic profiles [79] [77]. Addressing this challenge requires integrating specialized assessment methodologies throughout the molecular generation workflow, blending generative AI with active learning approaches that iteratively refine candidates based on synthetic feasibility and drug-likeness metrics [12].
This comparison guide examines the leading computational frameworks and scoring methodologies designed to bridge this gap, providing researchers with objective performance data and experimental protocols for implementing these approaches in drug discovery pipelines.
Defining the Key Assessment Metrics
Synthetic Accessibility (SA) Scoring
Synthetic Accessibility (SA) quantifies how easily a molecule can be synthesized in the laboratory, considering available building blocks, reaction types, stereochemical complexity, and scaffold feasibility [77]. Multiple computational approaches have been developed to estimate SA:
- Fragment-Based Scoring: Models such as SAscore and SYBA evaluate molecular fragments against frequencies in known chemical databases, assigning penalties for rare substructures [79] [77].
- Complexity-Based Scoring: These methods impose penalties for molecular features that increase synthetic difficulty, including large ring systems, multiple stereocenters, bridged or spiro atoms, and unusual bond types [77].
- Retrosynthesis-Based Scoring: More advanced approaches like SCScore and RAscore employ computer-aided synthesis planning (CASP) to evaluate synthetic routes, though these are computationally intensive for high-throughput screening [79] [76].
- Hybrid Models: Tools such as SynFrag combine fragment assembly with autoregressive generation to learn stepwise molecular construction patterns, offering rapid SA assessment with consideration of synthetic logic [76].
Table 1: Comparison of Synthetic Accessibility Scoring Methods
| Method | Approach | Scaling System | Speed | Key Advantages |
|---|---|---|---|---|
| SAscore [77] | Fragment-based + complexity penalties | 1 (easy) to 10 (hard) | Fast | Fast calculation, easily interpretable |
| SYBA [79] | Bayesian classification of fragments | Easy vs. hard classification | Fast | High accuracy for drug-like molecules |
| SCScore [79] | Retrosynthetic complexity | 1-5 scale | Moderate | Correlates with number of synthetic steps |
| SynFrag [76] | Fragment assembly generation | Continuous score | Fast | Captures synthesis logic, interpretable attention mechanisms |
| Makya (Iktos) [78] | Reaction-based generation | Binary (synthesizable/not) | Moderate | Guaranteed synthetic routes using real starting materials |
Drug-Likeness Evaluation
Drug-likeness encompasses molecular properties that determine suitability as oral therapeutics, typically assessed through established rules and quantitative metrics:
- Lipinski's Rule of Five: Evaluates molecular weight ≤500, LogP ≤5, hydrogen bond donors ≤5, and hydrogen bond acceptors ≤10 [75].
- Quantitative Estimate of Drug-likeness (QED): Combines multiple physicochemical properties into a unified score ranging from 0 (undrug-like) to 1 (highly drug-like) [75].
- ADMET Prediction: Machine learning models predict absorption, distribution, metabolism, excretion, and toxicity properties using structural fingerprints and molecular descriptors [74] [75].
Comparative Analysis of AI Platforms and Approaches
Generative AI with Embedded Chemistry Constraints
Leading AI drug discovery platforms have developed distinctive approaches to integrate synthetic accessibility and drug-likeness directly into the generation process:
Iktos's Makya Platform employs a "chemistry-first" approach that builds molecules via sequences of feasible chemical reactions using real commercially available starting materials [78]. This method guarantees synthetic accessibility by construction rather than post-hoc filtering. In benchmark comparisons, Makya outperformed open-source approaches like REINVENT 4, producing a larger proportion of compounds with viable synthetic routes and greater scaffold diversity [78].
Variational Autoencoder with Active Learning (VAE-AL) frameworks integrate generative AI with iterative refinement cycles that optimize for both synthetic accessibility and target affinity [12]. The system employs nested active learning loops where generated molecules are evaluated using chemoinformatic predictors (SA scores, drug-likeness filters) and molecular modeling oracles (docking scores). Molecules meeting threshold criteria are used to fine-tune the generative model, progressively improving output quality.
Schrödinger's Physics-Based Platform combines generative AI with physics-based molecular modeling and machine learning [11]. This approach has demonstrated clinical validation, advancing the TYK2 inhibitor zasocitinib into Phase III trials. The platform optimizes for drug-likeness throughout the design process, balancing potency with physicochemical properties [11].
Performance Comparison of Integrated Platforms
Table 2: Platform Performance in Experimental Validations
| Platform/Approach | Target(s) | Molecules Synthesized | Experimental Hit Rate | Notable Outcomes |
|---|---|---|---|---|
| VAE-AL Workflow [12] | CDK2, KRAS | 9 molecules for CDK2 | 8/9 active (89%); 1 nanomolar potency | Novel scaffolds with high predicted SA |
| Makya (Iktos) [78] | Multiple undisclosed targets | Not specified | Higher % of synthesizable candidates vs. REINVENT 4 | Greater scaffold diversity with guaranteed routes |
| Exscientia [11] | Multiple clinical programs | 10x fewer compounds than industry norms | 70% faster design cycles | First AI-designed drug to Phase I (DSP-1181) |
| Insilico Medicine [11] | TNIK (ISM001-055) | Not specified | Phase IIa positive results | Target-to-clinical in 18 months |
Experimental Protocols and Workflows
VAE-AL Framework Methodology
The VAE-AL framework represents a sophisticated integration of generative AI with active learning, specifically designed to address synthetic accessibility and drug-likeness [12]. The protocol involves these key stages:
- Initial Training: A variational autoencoder is initially trained on a broad chemical database (e.g., ChEMBL, ZINC) to learn general chemical space.
- Target-Specific Fine-tuning: The model is fine-tuned on target-specific active compounds to bias generation toward relevant chemotypes.
- Nested Active Learning Cycles:
- Inner Cycle: Generated molecules are evaluated using fast chemoinformatic filters (SA score, QED, similarity to known actives). Candidates passing thresholds are added to a temporal set used for model fine-tuning.
- Outer Cycle: After multiple inner cycles, accumulated molecules undergo molecular docking. High-scoring candidates advance to a permanent set for further model refinement.
- Candidate Selection: Final candidates are selected based on synthesis feasibility, novelty, and binding free energy calculations.
This workflow was validated on CDK2 and KRAS targets, successfully generating novel, synthesizable scaffolds with high predicted affinity [12]. For CDK2, the approach yielded 8 active compounds from 9 synthesized, including one with nanomolar potency.
VAE-AL Framework Workflow
Chemistry-Aware Generation Protocol
Iktos's Makya platform implements a fundamentally different approach that guarantees synthetic feasibility by construction [78]:
- Reaction-Based Generation: Instead of generating molecular strings, the system selects real commercially available building blocks from curated catalogs.
- Stepwise Assembly: Molecules are constructed through sequences of known chemical reactions, respecting reaction rules and compatibility.
- Constraint Application: Users can impose synthetic constraints including maximum step count, specific reaction types, or preferred building block sources.
- Multi-Objective Optimization: The platform simultaneously optimizes for target engagement, drug-likeness, and synthetic feasibility while maintaining chemical diversity.
This methodology ensures that all proposed molecules have realistic synthetic routes, addressing the core limitation of string-based generative approaches [78].
Essential Research Reagents and Computational Tools
Successful implementation of synthetic accessibility and drug-likeness assessment requires specialized computational tools and databases. The table below catalogues essential resources referenced in the experimental protocols.
Table 3: Research Reagent Solutions for SA and Drug-Likeness Assessment
| Tool/Resource | Type | Primary Function | Application in Workflows |
|---|---|---|---|
| RDKit SA_Score [77] | Software Library | Synthetic accessibility scoring based on fragment frequency and complexity | Fast SA screening in high-throughput generative workflows |
| SynFrag [76] | Web Platform / Code | SA prediction via fragment assembly generation | Interpretable SA assessment with attention mechanisms |
| eTox (Neurosnap) [77] | Predictive Service | Simultaneous toxicity and SA prediction (1-10 scale) | Integrated toxicity and SA screening in candidate prioritization |
| Mordred Descriptors [77] | Descriptor Calculator | 1,614 molecular descriptors for QSAR modeling | Comprehensive molecular profiling for drug-likeness assessment |
| Commercial Building Block Databases [78] | Chemical Database | Curated available chemical starting materials | Chemistry-aware generation in platforms like Makya |
| ChEMBL / ZINC [12] | Chemical Database | Annotated bioactive molecules and commercially available compounds | Training data for generative models and benchmark comparisons |
The integration of robust synthetic accessibility and drug-likeness assessment into AI-driven molecular discovery represents a critical advancement toward practical pharmaceutical applications. The comparative analysis reveals that while generative AI alone can propose novel structures, the combination with active learning frameworks or chemistry-aware generation significantly enhances the feasibility and efficiency of drug discovery pipelines.
Platforms that embed synthetic feasibility directly into the generation process, such as Iktos's Makya and VAE-AL frameworks, demonstrate superior performance in producing synthesizable candidates with maintained diversity and target engagement [78] [12]. The experimental success of these approaches—evidenced by high synthesis success rates and nanomolar potency in validated targets—underscores the importance of addressing the generation-synthesis gap at the architectural level rather than through post-hoc filtering.
As AI-designed molecules continue to advance through clinical trials, the systematic integration of these methodologies will be essential for realizing the full potential of AI-driven drug discovery, transforming computational innovation into tangible therapeutic benefits.
Computational and Expert Resource Constraints in Active Learning Loops
In the rapidly evolving field of artificial intelligence, two distinct paradigms have emerged as powerful tools for scientific discovery: active learning and generative AI. While generative models like large language models (LLMs) have demonstrated remarkable capabilities in creating novel content, active learning provides a framework for data-efficient model training by strategically selecting the most informative samples for expert annotation. This comparison guide examines these approaches through the critical lens of computational and expert resource constraints, particularly within research domains such as drug development where labeled data is scarce and expensive to acquire.
Active learning operates through an iterative human-in-the-loop process where machine learning models selectively query human experts to label new data points. This approach is specifically designed to maximize model performance while minimizing labeling costs by focusing expert attention on the most valuable data instances. In contrast, generative AI typically requires massive pre-training datasets but can subsequently generate synthetic data or provide zero-shot predictions. Understanding the trade-offs between these approaches enables researchers to make informed decisions about which methodology best suits their specific resource constraints and research objectives.
Defining the Approaches
Active Learning: Principles and Workflow
Active learning is a specialized machine learning framework that addresses the fundamental challenge of limited labeled data by implementing intelligent data selection strategies. The core premise involves iteratively selecting the most informative unlabeled samples for expert annotation, thereby reducing the overall labeling burden while maintaining or improving model performance [80]. This approach is particularly valuable in domains like materials science and drug development where experimental synthesis and characterization require expert knowledge, expensive equipment, and time-consuming procedures [81].
The typical active learning workflow operates through a structured cycle. Initialization begins with a small labeled dataset to train a baseline model. Through iterative sampling, the system then selects the most promising unlabeled instances based on specific query strategies, which are subsequently sent for human annotation by domain experts. The newly labeled data is incorporated into the training set, leading to model retraining and refinement. This cycle continues until performance plateaus or resources are exhausted, representing a strategic balance between computational efficiency and expert involvement [80] [81].
Generative AI: Capabilities and Resource Demands
Generative AI refers to a class of machine learning models capable of creating new content—including text, images, or molecular structures—based on patterns learned from large datasets. As noted by MIT experts, "Generative AI goes beyond capturing complex correlations and patterns in existing data to creating new content" [27]. These models, particularly large language models like GPT-4 and similar architectures, have demonstrated remarkable capabilities in both content generation and, increasingly, in predictive tasks that were traditionally the domain of specialized machine learning models.
The resource profile of generative AI differs significantly from active learning. These models typically require substantial pre-training resources, with advanced GPUs like NVIDIA's A100 or H100 and extensive memory (often 128GB or more) needed for the initial training phase [82] [83]. The computational demands are driven by the scale of parameters and datasets involved; for instance, training the largest Llama 3 models required Meta to utilize 16,000 GPUs simultaneously [82]. However, once trained, inference with these models becomes more accessible, with options for cloud-based deployment and optimized hardware for specific applications [83].
Comparative Analysis: Performance and Resource Efficiency
Quantitative Performance Benchmarks
Table 1: Performance Comparison of Active Learning Strategies in Materials Science (Regression Tasks) [81]
| Active Learning Strategy Type | Early-Stage Performance (MAE) | Data Efficiency Gain | Time to Convergence |
|---|---|---|---|
| Uncertainty-Driven (LCMD) | Best | 70-95% data savings | Fastest |
| Diversity-Hybrid (RD-GS) | Best | High data savings | Fast |
| Tree-based Uncertainty (Tree-based-R) | Good | Moderate-High data savings | Moderate |
| Geometry-Only (GSx) | Fair | Low data savings | Slow |
| Expected Gradient (EGAL) | Fair | Low data savings | Slow |
| Random Sampling (Baseline) | Poor | Baseline (0% savings) | Slowest |
Table 2: Resource Requirements Comparison
| Resource Factor | Active Learning | Generative AI |
|---|---|---|
| Initial Training Data | Small labeled dataset sufficient | Massive unlabeled dataset required |
| Expert Annotation | Ongoing but optimized (44% time savings reported) [2] | Primarily upfront or minimal for fine-tuning |
| Computational Hardware | Moderate (can use cloud solutions or local clusters) [82] | Extensive (high-end GPUs/TPUs, 128GB+ RAM) [83] |
| Implementation Timeline | Weeks for proof-of-concept [82] | Rapid deployment for existing models |
| Domain Adaptation | Built into the process | Requires fine-tuning or prompt engineering |
Recent benchmark studies demonstrate that active learning strategies can achieve remarkable data efficiency while maintaining model accuracy. In materials science regression tasks, uncertainty-driven and diversity-hybrid approaches have shown particular effectiveness, selecting more informative samples and improving model accuracy, especially during early acquisition phases when data is scarce [81]. The performance gap between strategic active learning methods and random sampling is most pronounced when labeled data is limited, with some studies reporting 70-95% data savings while achieving performance comparable to models trained on full datasets [81].
The effectiveness of different active learning strategies varies throughout the learning cycle. Uncertainty-based methods like LCMD and Tree-based-R excel in early stages when data is scarce, rapidly improving model performance with minimal labeled examples. Diversity-based approaches such as RD-GS maintain effectiveness across multiple stages by ensuring broad coverage of the feature space. As the labeled set grows, the performance advantage of sophisticated active learning strategies gradually diminishes, with most methods converging when sufficient data becomes available [81].
Resource Utilization Patterns
The resource consumption profiles of active learning and generative AI differ significantly both in magnitude and distribution. Active learning implementations typically require more modest computational infrastructure, with options for both local clusters and cloud solutions depending on project needs and duration [82]. The primary resource constraint in active learning is human expertise for annotation, though studies show that AI-assisted administrative tasks can save teachers (as domain experts) 44% of their time on research, lesson planning, and material creation [2].
Generative AI models demand substantial upfront computational investment, with fine-tuning requiring high-performance GPUs and extensive memory [83]. However, inference with pre-trained models has become increasingly efficient, with costs dropping over 280-fold for systems performing at GPT-3.5 levels between 2022 and 2024 [84]. For organizations with limited in-house computational resources, cloud-based solutions provide flexibility, though long-term costs may exceed custom local clusters for continuous use [82].
Experimental Protocols and Methodologies
Benchmarking Active Learning Strategies
Table 3: Essential Research Reagents for Active Learning Experiments
| Component | Function | Implementation Examples |
|---|---|---|
| Base Model Architecture | Core predictive algorithm | Gradient boosting machines, neural networks, support vector regressors |
| Query Strategy | Selects most informative samples | Uncertainty sampling, diversity maximization, expected model change |
| AutoML Framework | Automates model selection and hyperparameter tuning | AutoML systems for pipeline optimization [81] |
| Validation Protocol | Measures performance generalization | 5-fold cross-validation, hold-out test sets |
| Stopping Criterion | Determines when to halt the active learning cycle | Performance plateau, resource exhaustion |
Systematic evaluation of active learning strategies requires carefully designed experimental protocols. The benchmark process typically employs a pool-based active learning framework where an initial dataset is partitioned into labeled and unlabeled pools [81]. The process begins with random sampling of n_init samples from the unlabeled dataset to create an initial labeled dataset. Different active learning strategies then perform multi-step sampling, with the sampled instances added to the labeled pool after simulated "annotation."
At each sampling iteration, an AutoML model is fitted and evaluated on a held-out test set, typically using an 80:20 train-test split with 5-fold cross-validation for robust performance estimation [81]. Key performance metrics include Mean Absolute Error (MAE) and the Coefficient of Determination (R²) for regression tasks, with each strategy compared against random sampling as a baseline. The evaluation focuses particularly on early-stage performance when data is scarcest, as this is where active learning provides the greatest value.
Generative AI Evaluation Metrics
Evaluating generative AI models presents distinct challenges, particularly for scientific applications. Beyond traditional accuracy metrics, assessments typically include:
Benchmark performance on standardized tasks (MMMU, GPQA, SWE-bench) where leading models have shown rapid improvement, with scores increasing by 18.8, 48.9, and 67.3 percentage points respectively in recent years [84]
Factuality and safety measurements using emerging benchmarks like HELM Safety, AIR-Bench, and FACTS [84]
Domain-specific accuracy for specialized applications, which often requires expert validation and careful prompt engineering [27]
For drug development applications, additional evaluation criteria might include synthetic molecule validity, novelty, and docking scores for virtual screening.
Decision Framework and Strategic Implementation
When to Choose Active Learning
Active learning presents distinct advantages for research domains with specific resource constraints. This approach is particularly suitable when:
Expert annotation is available but limited, as active learning optimizes the use of this scarce resource [81]
Data acquisition costs are high, such as in experimental materials science or wet-lab biological validation [81]
Problems are highly domain-specific with technical knowledge requirements that may challenge general-purpose generative models [27]
Data privacy concerns restrict cloud-based solutions, making local model development necessary [27]
Evidence from educational environments demonstrates that AI-powered active learning generates 10 times more engagement than traditional passive methods and improves student outcomes by up to 30% compared to traditional approaches [2], suggesting similar potential for research team training and knowledge acquisition.
When Generative AI Is Preferable
Generative AI approaches offer compelling advantages under different constraint profiles:
Rapid prototyping is needed for problems involving everyday language or common images [27]
Technical expertise in traditional ML is limited, as generative AI "is a democratizing force" that makes advanced capabilities more accessible [27]
Existing foundation models align well with the problem domain, enabling fine-tuning rather than development from scratch
Data generation rather than prediction is the primary objective, such as creating novel molecular structures for screening
Budget allows for substantial computational investment either in fine-tuning or inference at scale
Hybrid Approaches
Increasingly, researchers are finding value in combining both approaches to leverage their complementary strengths:
Using generative AI to create synthetic data for augmenting small experimental datasets [27]
Applying generative models for data preprocessing and cleaning to improve active learning efficiency [27]
Implementing active learning for fine-tuning generative models on domain-specific problems
These hybrid approaches recognize that the choice between methodologies is not necessarily binary, but rather a spectrum of options that can be strategically combined to address specific research constraints.
The comparison between active learning and generative AI reveals a nuanced landscape where neither approach dominates universally across all resource constraint scenarios. Active learning demonstrates clear advantages in data-efficient modeling, particularly when expert annotation is available but computationally expensive, and when problems require specialized domain knowledge. The strategic sample selection in active learning can reduce data requirements by 70-95% while maintaining model performance [81], making it invaluable for resource-constrained research environments.
Conversely, generative AI offers compelling capabilities when rapid deployment is prioritized, when problems align well with pre-trained model capabilities, and when computational resources are more readily available than domain expertise. The dramatic improvements in benchmark performance and rapidly decreasing inference costs make generative AI increasingly accessible [84], though careful attention to domain adaptation remains crucial.
For research domains like drug development, where both computational and expert resources are typically constrained, the strategic integration of both approaches presents promising opportunities. By leveraging active learning for targeted data acquisition and generative AI for data augmentation and rapid prototyping, research teams can navigate resource constraints more effectively, accelerating the pace of scientific discovery while optimizing their limited resources.
The integration of artificial intelligence (AI) into drug development represents a paradigm shift, offering the potential to reduce the decade-long timelines and exorbitant costs traditionally associated with bringing a new drug to market. Two distinct AI-driven approaches—generative AI and active learning—are now at the forefront of this transformation. Generative AI focuses on the de novo creation of novel drug-like molecules and structures, leveraging deep learning models to explore vast chemical spaces that would otherwise remain inaccessible. In parallel, active learning systems employ an iterative, data-driven selection process to guide experimental testing towards the most informative data points, thereby maximizing learning efficiency and minimizing resource expenditure.
Navigating the regulatory and intellectual property landscape is crucial for the successful adoption of these technologies. The U.S. Food and Drug Administration (FDA) provides a framework for the development and approval of new therapeutics, a process that begins with an Investigational New Drug (IND) application [85]. For AI-driven drug discovery, key regulatory considerations include the validation of AI-generated compounds, the adequacy of AI-predicted endpoints, and the use of non-traditional data sources to support applications. The intellectual property framework, meanwhile, must adapt to protect AI-generated inventions, algorithm originality, and the unique data assets used for model training. This guide provides a structured comparison of these approaches within the current regulatory and ethical context to inform researchers, scientists, and drug development professionals.
FDA Regulatory Framework for AI-Driven Drug Development
The Investigational New Drug (IND) Application
The IND application is the critical gateway for initiating clinical trials on a new drug in the United States. Technically, it is an exemption from the federal law that prohibits the shipment of unapproved drugs across state lines, allowing the investigational drug to be distributed to clinical investigators in different states [85]. From a legal standpoint, the submission of an IND marks the point at which a molecule transitions into a "new drug" subject to specific FDA requirements.
An IND application must contain information in three broad areas [85]:
- Animal Pharmacology and Toxicology Studies: Preclinical data to demonstrate that the product is reasonably safe for initial testing in humans.
- Manufacturing Information: Details on the composition, manufacturer, stability, and controls used for producing the drug substance and product, ensuring the sponsor can consistently produce the drug.
- Clinical Protocols and Investigator Information: Detailed protocols for proposed clinical studies to assess whether initial-phase trials will expose subjects to unnecessary risks, along with commitments to obtain informed consent and institutional review board (IRB) approval.
For sponsors using AI in their discovery process, demonstrating the validity and reliability of the AI tools and the data they generate is paramount. This may involve providing additional justification for target selection, compound design, and predictive safety or efficacy models derived from AI.
The FDA provides extensive resources to assist sponsors in the IND process. While not legally enforceable, FDA Guidance Documents represent the agency's current thinking on a particular subject and are invaluable for understanding regulatory expectations [85]. Sponsors can search for relevant guidances on the FDA's website, filtering by topic such as "Drugs" or "Clinical Trials" [86].
Key resources include:
- Pre-IND Consultation Program: This program fosters early communication between sponsors and FDA review divisions to obtain guidance on the data necessary for a successful IND submission [85].
- Laws, Regulations, and Policies: The Code of Federal Regulations (CFR), particularly 21 CFR Part 312, details the legal requirements for INDs. CDER's Manual of Policies & Procedures (MaPPs) provides internal instructions for the FDA's drug review staff [85].
Recent political and organizational changes at the FDA have introduced a degree of uncertainty. A significant reduction in force in April 2025, while excluding drug reviewers, has affected policy offices and support staff, leading to reports of slower communication and delayed meeting schedules [87]. Furthermore, new leadership under Commissioner Marty Makary has expressed interest in reducing animal testing in favor of newer technologies, which could directly impact the preclinical data requirements for INDs, though the specifics of acceptable alternatives are not yet clear [87]. Sponsors are advised to engage with the FDA as early as possible and monitor for new and updated guidance documents.
Intellectual Property Considerations for AI in Pharma
The integration of AI into the drug development pipeline creates novel and complex challenges for intellectual property (IP) protection. The traditional IP framework is being tested by inventions that involve significant AI contribution.
A primary challenge lies in determining patentability for AI-generated inventions. Key questions include the ownership of patents for compounds conceived by an AI system and the threshold for inventiveness when an AI is involved in the discovery process. Furthermore, the data used to train AI models is a critical asset. While raw data is generally not patentable, curated, unique, and high-quality datasets can provide a competitive advantage and may be protected as trade secrets. The algorithms and models themselves are another IP cornerstone. Protecting the underlying AI architecture through patents or copyrights is essential, but the fast-paced evolution of models can render patents obsolete quickly. Finally, the regulatory strategy must be integrated with IP considerations. The scope of patent protection for an AI-discovered drug must be carefully crafted to withstand legal scrutiny, especially if the path to discovery differs significantly from traditional methods. As noted in a review on AI in drug development, establishing more comprehensive intellectual property protections for algorithms remains a challenge that the industry must address [88].
Comparative Analysis: Generative AI vs. Active Learning
While both generative AI and active learning are transformative technologies, they address different stages and challenges in the drug discovery workflow. The table below summarizes their core functions, applications, and key differentiators.
Table 1: High-Level Comparison of Generative AI and Active Learning
| Feature | Generative AI | Active Learning |
|---|---|---|
| Core Function | Creates novel drug candidates, molecules, or data | Selects the most informative experiments to perform |
| Primary Application | De novo drug design, molecule generation, property prediction | Guided screening (e.g., synergistic drug pairs), optimization loops |
| Key Strength | Explores vast, novel chemical space; designs from scratch | Maximizes learning and resource efficiency; handles "rare event" discovery |
| Data Dependency | Requires large initial training datasets | Starts with a small dataset and iteratively expands it |
| Regulatory Focus | Validation of novel compounds & AI-predicted properties | Justification of adaptive trial designs & data selection criteria |
| IP Considerations | Patentability of AI-generated inventions, model architecture | Proprietary selection algorithms, curated experimental datasets |
Experimental Protocols and Performance Data
To objectively compare their performance, it is essential to examine the experimental protocols and quantitative outcomes reported in recent studies.
Active Learning in Synergistic Drug Combination Screening
Protocol Overview: A key study provides a detailed protocol for using active learning to identify synergistic drug pairs [33]. Synergy is a rare event (e.g., 1.47%-3.55% in common datasets), making exhaustive screening prohibitively expensive.
- Objective: To rapidly identify synergistic drug combinations by sequentially selecting small batches of experiments that maximize learning.
- Methodology: The framework involves an AI algorithm (e.g., a neural network) pre-trained on public synergy data. The model uses molecular fingerprints (e.g., Morgan fingerprints) and cellular features (e.g., gene expression profiles) as input. Iteratively, the model selects a batch of unmeasured drug-cell pairs predicted to be most synergistic or informative, these are "tested" (in silico or in vitro), and the new data is used to retrain the model for the next cycle [33].
- Key Parameters: The study highlighted the importance of batch size and the exploration-exploitation trade-off. Smaller batch sizes and dynamic tuning of the selection strategy were found to significantly enhance performance.
Table 2: Experimental Performance of Active Learning in Drug Synergy Screening [33]
| Metric | Performance | Context & Comparison |
|---|---|---|
| Synergy Discovery Rate | Discovered 60% of synergistic pairs by exploring only 10% of the combinatorial space | Without a strategic approach, finding 300 synergistic combinations required 8,253 measurements. |
| Resource Efficiency | Saved 82% of experimental time and materials | Achieved the same goal (300 synergies) with only 1,488 measurements vs. 8,253. |
| Impact of Molecular Encoding | Limited impact on prediction quality | Morgan fingerprint with addition operation performed best. |
| Impact of Cellular Features | Significant performance improvement (0.02-0.06 gain in PR-AUC) | Using gene expression profiles was crucial; as few as 10 relevant genes were sufficient. |
Generative AI in Drug Discovery and Development
Protocol Overview: Generative AI employs deep learning models, such as generative adversarial networks (GANs) or transformer-based models, to design new molecular structures with desired properties.
- Objective: To generate novel, synthetically accessible drug-like molecules with optimized properties for a specific target.
- Methodology: Models are trained on large chemical databases (e.g., ZINC, ChEMBL). They learn the underlying probability distribution of chemical structures and their associated properties. Once trained, they can sample from this distribution to generate new molecules. Techniques like reinforcement learning can further fine-tune the generation towards specific objectives, such as high binding affinity or favorable ADMET (Absorption, Distribution, Metabolism, Excretion, Toxicity) profiles [89] [90].
- Key Applications: This approach is used for de novo molecular design, lead compound optimization, and predicting complex biological properties of molecules, thus reducing reliance on early-stage experimental screening [89].
While direct, head-to-head quantitative comparisons between generative AI and active learning are less common, as they are often used for different tasks, their impacts are profound. Generative AI has revolutionized early-stage hit identification and lead optimization by exploring chemical spaces much more efficiently than human intuition or traditional virtual screening [90]. Active learning, by contrast, excels at optimizing expensive and complex experimental campaigns, such as high-throughput screening, by dramatically reducing the number of experiments needed to achieve a goal [33].
Essential Research Reagent Solutions
The experimental workflows for generative AI and active learning rely on a suite of computational and data resources. The following table details key "reagent solutions" essential for research in this field.
Table 3: Key Research Reagents and Resources for AI-Driven Drug Discovery
| Item Name | Type | Primary Function in Research |
|---|---|---|
| Morgan Fingerprints | Molecular Descriptor | Encodes the structure of a molecule into a fixed-length bit string based on the presence of specific circular substructures; used as input for AI models [33]. |
| Gene Expression Profiles | Cellular Feature | Provides genomic context of the target cell line (e.g., from GDSC database); significantly improves synergy prediction and other cell-specific outcomes [33]. |
| DrugComb / O'Neil Dataset | Curated Database | A meta-database of drug combination screening data; used for pre-training and benchmarking AI models for synergy prediction [33]. |
| Deep Neural Network (DNN) | AI Algorithm | A multi-layered AI model capable of learning complex, non-linear relationships from data; used for both predictive tasks and generative design [89] [33]. |
| ZINC / ChEMBL | Chemical Database | Large, publicly accessible databases of commercially available and bioactive molecules; serve as the training data for generative AI models [89]. |
| Quantitative Structure-Activity Relationship (QSAR) Model | Predictive Model | A computational model that correlates chemical structure with biological activity; AI-based QSAR (e.g., using SVM, Random Forest) accelerates this analysis [89]. |
Workflow and Relationship Visualizations
High-Level AI Drug Discovery Workflow
The following diagram illustrates a generalized workflow integrating both generative AI and active learning approaches within the broader drug development context, including key regulatory and IP milestones.
The Active Learning Iterative Cycle
This diagram details the core, iterative feedback loop that defines the active learning methodology, as applied in areas like synergistic drug pair screening.
The integration of generative AI and active learning into the pharmaceutical research and development pipeline marks a significant evolution in the field. While generative AI acts as a powerful engine for molecular invention, active learning serves as an intelligent guide for experimental efficiency. The choice between or combination of these approaches depends heavily on the specific research problem: creating novel chemical matter versus optimizing a costly experimental campaign.
Navigating the associated regulatory and intellectual property landscapes is a critical component of success. Engaging with the FDA through its Pre-IND program and adhering to its guidance documents is essential for de-risking the path to clinical trials. Simultaneously, a proactive IP strategy that considers the protection of AI-generated discoveries, proprietary algorithms, and unique datasets is necessary to secure the value created by these advanced technologies. As the regulatory environment adapts to scientific advancements and the capabilities of AI continue to mature, these tools are poised to become indispensable in the development of the next generation of therapeutics.
Evidence and Efficacy: Clinical Validation and Strategic Comparative Analysis
The integration of artificial intelligence (AI) into drug discovery represents a paradigm shift in pharmaceutical development, promising to address the sector's persistently high failure rates and unsustainable costs. By mid-2025, the landscape has evolved from theoretical promise to tangible clinical impact, with over 75 AI-derived molecules reaching clinical stages [11]. This review critically examines the clinical track records of leading AI-driven drug discovery platforms, comparing their performance against traditional development approaches and analyzing how their underlying technologies—from generative AI to active learning systems—influence success and failure patterns.
AI has demonstrated remarkable capabilities in compressing early-stage timelines. Several companies have advanced candidates from discovery to Phase I trials in approximately 18 months, a fraction of the traditional ~5-year timeline [11] [91]. However, the ultimate validation requires clinical success, and here the picture is more nuanced. While AI-designed compounds show improved safety profiles—with approximately 90% successfully completing Phase I safety trials compared to <65% for traditional molecules—their ability to demonstrate superior efficacy in later-stage trials remains unproven [91]. This analysis focuses specifically on clinical-stage performance, examining both the accelerated pathways and persistent translational challenges facing AI-designed therapeutics.
Clinical Pipeline Analysis: Quantitative Comparison of AI-Designed Drugs
Table 1: Clinical-Stage AI-Designed Drug Candidates and Their Development Status
| Company/Platform | AI Technology Approach | Key Drug Candidate(s) | Therapeutic Area | Latest Clinical Status | Reported Timeline Efficiency | Key Efficacy Outcomes |
|---|---|---|---|---|---|---|
| Insilico Medicine | Generative AI (GANs + RL), Knowledge Graphs | ISM001-055 (TNK inhibitor) | Idiopathic Pulmonary Fibrosis | Phase IIa (positive results reported) [11] | ~18 months from target to Phase I [11] [92] | Positive Phase IIa results in IPF [11] |
| Exscientia | Generative Design, Automated Precision Chemistry | EXS-21546 (A2A antagonist) | Immuno-oncology | Phase I discontinued (insufficient therapeutic index) [11] | Design cycles ~70% faster, 10x fewer compounds synthesized [11] | Failed due to predicted insufficient therapeutic index [11] |
| Schrödinger | Physics-Enabled ML Design | Zasocitinib (TAK-279, TYK2 inhibitor) | Autoimmune Diseases | Phase III (advanced from Nimbus acquisition) [11] | N/A (acquired after clinical advancement) | Advanced to late-stage testing based on profile [11] |
| Recursion | Phenomics-First AI, Cellular Imaging | REC-1245 | Undisclosed | IND-enabling studies (18-month discovery timeline) [91] | 18 months to IND-enabling studies [91] | Preclinical stage; clinical outcomes pending [91] |
| BenevolentAI | Knowledge Graph Repurposing | Baricitinib repurposing | COVID-19 | Emergency Use Authorization [92] | Accelerated identification for new indication [92] | Successful repurposing for severe COVID-19 [92] |
Table 2: Aggregate Performance Metrics of AI-Designed vs. Traditional Drug Candidates
| Performance Metric | AI-Designed Candidates | Traditional Candidates | Data Source |
|---|---|---|---|
| Phase I Success Rate | ~90% [91] | <65% [91] | Industry analysis of AI-designed molecules [91] |
| Phase II Success Rate | ~40% (early data) [91] | ~40% (historical average) [91] | Limited dataset of AI-designed molecules [91] |
| Preclinical Timeline | 12-18 months (reported peaks) [11] [91] | ~4-5 years (industry average) [11] | Company reports and independent analysis [11] [91] |
| Discovery Cost Reduction | 25-30% (modeled for novel targets) [91] | Baseline | Independent modeling analysis [91] |
| Clinical-stage Attrition | 95% (similar to traditional) [91] | 95% (historical average) [91] | Industry-wide tracking [91] |
The quantitative comparison reveals a mixed clinical track record. AI has unquestionably accelerated preclinical development and improved early-stage safety outcomes. However, the technology has not yet demonstrated a clear ability to overcome the fundamental efficacy challenges that plague traditional drug development, with Phase II success rates remaining comparable to historical averages [91]. This suggests that while AI optimizes molecule design and safety profiling, it may not yet adequately address the translational gap between preclinical models and human therapeutic effects.
Technological Approaches: Generative AI vs. Active Learning in Clinical Translation
Generative AI Platforms
Generative AI platforms typically employ generative adversarial networks (GANs), reinforcement learning (RL), and transformer architectures to design novel molecular structures from scratch. Insilico Medicine's Pharma.AI platform exemplifies this approach, using a combination of policy-gradient-based reinforcement learning and generative models for multi-objective optimization of parameters including potency, toxicity, and novelty [93]. The platform integrates multiple specialized modules: PandaOmics for target identification leveraging 1.9 trillion data points, Chemistry42 for generative molecule design, and inClinico for clinical trial outcome prediction [93].
These systems demonstrate exceptional performance in molecular generation and optimization, with Insilico reporting the design of a novel anti-fibrotic drug and advancement to Phase I trials in approximately 18 months—roughly 50% of traditional timelines [11] [94]. However, this accelerated design capability doesn't necessarily translate to improved clinical efficacy, as these systems often operate with limited human biological context during the design phase [91].
Active Learning and Phenomics Platforms
Active learning platforms employ iterative, data-driven cycles where AI models design experiments, analyze results, and continuously refine their hypotheses. Recursion's platform exemplifies this approach, combining large-scale cellular phenotyping with machine learning in a closed-loop system. Their Phenom-2 model utilizes a 1.9 billion-parameter Vision Transformer trained on approximately 8 billion microscopy images to detect subtle patterns in cellular morphology [93]. This is integrated with MolPhenix for molecule-phenotype effect prediction and MolGPS for molecular property prediction, creating a continuous feedback cycle between computational prediction and experimental validation [93].
The primary strength of active learning approaches lies in their grounding in empirical biological data rather than purely structural or chemical information. However, these systems face challenges of scale and complexity, requiring massive investments in automated laboratory infrastructure and generating enormous datasets (Recursion reports approximately 65 petabytes of proprietary data) [93].
Comparative Clinical Strengths and Limitations
Table 3: Technology Comparison and Clinical Implications
| AI Approach | Clinical-Strengths | Clinical Limitations | Representative Companies |
|---|---|---|---|
| Generative AI | Rapid exploration of chemical space; Multi-parameter optimization; Novel scaffold design | Limited human physiological data integration; Black box design decisions; Questionable translatability | Insilico Medicine, Exscientia, Iambic Therapeutics |
| Active Learning/Phenomics | Grounded in empirical biological data; Continuous experimental validation; Phenotypic relevance | Massive infrastructure requirements; Complex data interpretation; High computational costs | Recursion, Verge Genomics |
| Knowledge Graph Repurposing | Leverages existing clinical data; De-risked compounds; Faster path to clinic | Limited to known biology; Less innovative mechanisms; Dependent on data quality | BenevolentAI |
| Physics-Enabled ML | Incorporates biophysical principles; Better affinity predictions; Interpretable models | Computationally intensive; Limited to well-characterized targets | Schrödinger |
Analysis of Clinical Triumphs: Case Studies
Insilico Medicine's ISM001-055 for Idiopathic Pulmonary Fibrosis
Insilico's TNK inhibitor for idiopathic pulmonary fibrosis represents one of the most advanced validation cases for generative AI. The program advanced from target discovery to Phase I trials in 18 months, leveraging the company's end-to-end generative platform [11]. The target (TNK) was identified using the PandaOmics system analyzing multi-omics and literature data, followed by generative design of the inhibitor through Chemistry42 [93]. The compound demonstrated sufficient promise in Phase I to advance to Phase IIa trials, where it has reportedly shown positive results [11]. This case demonstrates AI's potential to dramatically accelerate the early discovery pipeline while still generating clinically viable candidates.
Baricitinib Repurposing for COVID-19
BenevolentAI's identification of baricitinib as a COVID-19 treatment represents a successful application of AI knowledge graphs for drug repurposing. The company used its knowledge graph technology—integrating scientific literature, clinical trial data, and omics datasets—to identify the JAK1/2 inhibitor as a potential therapeutic for severe COVID-19 [92]. This led to emergency use authorization based on clinical trial data, demonstrating AI's capability to rapidly identify novel therapeutic applications for existing compounds by integrating diverse biological and clinical data sources [92].
Analysis of Clinical Setbacks: Case Studies
Exscientia's EXS-21546 Program Discontinuation
Exscientia's A2A receptor antagonist (EXS-21546) for immuno-oncology represents a notable clinical setback. The compound was designed using the company's generative AI platform and advanced to Phase I trials [11]. However, the program was discontinued in late 2023 after competitor data suggested it would unlikely achieve a sufficient therapeutic index [11]. This case highlights a key limitation in current AI approaches: while they can efficiently design compounds with desired target affinity and pharmacological properties, predicting the complex therapeutic window necessary for clinical success remains challenging. The failure also illustrates how external competitive landscapes can abruptly change a program's viability regardless of its technical success.
The Translational Efficacy Gap
A fundamental challenge facing AI-designed drugs is the persistent gap between preclinical optimization and clinical efficacy. As noted in a 2025 analysis, while AI-designed compounds show excellent safety profiles in early trials, their mechanisms of action are generally comparable to traditional drugs, and they face similar efficacy hurdles in Phase II proof-of-concept studies [91]. This suggests that current AI approaches, while excellent at optimizing molecules against specific targets, may not adequately address the complexity of human disease biology and patient variability.
Experimental Protocols and Methodologies
Protocol: AI-Driven Target Identification and Validation
Objective: Identify novel therapeutic targets using multi-modal data integration and prioritize based on disease relevance and druggability.
Methodology:
- Data Aggregation: Collect and pre-process multi-omics data (genomics, transcriptomics, proteomics) from public repositories (e.g., TCGA, GTEx) and proprietary sources, alongside scientific literature and patent data using NLP extraction [93].
- Knowledge Graph Construction: Build a heterogeneous knowledge graph integrating gene-disease associations, protein-protein interactions, compound-target relationships, and pathway information using systems like Insilico's PandaOmics or Recursion's knowledge graph tools [93].
- Target Prioritization: Apply graph neural networks and attention mechanisms to identify key network nodes with high disease association and druggability potential. Use community detection algorithms to identify disease-relevant biological modules [93].
- Experimental Validation: Design CRISPR-based knockout/knockdown experiments in disease-relevant cell models. Perform high-content phenotypic screening to assess functional impact of target modulation [91] [93].
Protocol: Generative Molecular Design and Optimization
Objective: Design novel chemical entities with optimized binding affinity, selectivity, and ADMET properties.
Methodology:
- Structure-Based Design: Utilize physics-based methods (e.g., molecular dynamics, free energy perturbation) combined with neural network potentials (Iambic's NeuralPLexer) to predict protein-ligand complexes and binding affinities [93].
- Generative Chemical Design: Employ generative models (GANs, VAEs, or autoregressive models) conditioned on target properties to explore chemical space. Implement reaction-aware generation to ensure synthetic feasibility [93].
- Multi-parameter Optimization: Apply reinforcement learning with reward functions balancing potency, selectivity, ADMET properties, and novelty. Use Pareto optimization techniques to identify optimal compound candidates [93].
- Iterative Design-Make-Test-Analyze (DMTA): Establish closed-loop cycles where AI models propose compounds, which are synthesized and tested experimentally, with results fed back to refine the models [11] [93].
Research Reagent Solutions for AI-Driven Discovery
Table 4: Essential Research Reagents and Platforms for AI-Driven Drug Discovery
| Reagent/Platform Category | Specific Examples | Function in AI Drug Discovery |
|---|---|---|
| High-Content Screening Systems | Phenom-2 imaging platform [93] | Generates cellular phenotyping data for training active learning systems |
| Multi-Omics Profiling Tools | RNA sequencing, Proteomics platforms [93] | Provides molecular data for target identification and validation |
| Knowledge Graph Databases | Proprietary and public knowledge bases [93] | Integrates disparate biological data for holistic analysis |
| Generative Chemistry Software | Chemistry42, Magnet (Iambic) [93] | Enables de novo molecular design with multi-parameter optimization |
| Protein Structure Prediction | NeuralPLexer, AlphaFold [92] [93] | Provides structural information for structure-based drug design |
| ADMET Prediction Platforms | MolGPS, Enchant (Iambic) [93] | Predicts clinical properties of candidates before synthesis |
Visualizing AI Drug Discovery Workflows
AI Drug Discovery Workflow Comparison: This diagram illustrates the distinct approaches of generative AI versus active learning/phenomics platforms, converging on clinical translation challenges.
Clinical Performance Factors: This diagram maps the factors influencing the mixed clinical track record of AI-designed drugs, highlighting both successes and persistent challenges.
The clinical track record of AI-designed drugs through 2025 reveals a field in transition, demonstrating remarkable engineering achievements in accelerating preclinical development while facing persistent biological challenges in clinical translation. The technology has proven exceptionally capable at compressing discovery timelines and optimizing molecular properties, with several candidates reaching clinical stages in roughly half the traditional time [11]. However, the fundamental challenge of predicting efficacy in human patients remains largely unsolved, with AI-designed molecules showing similar Phase II failure rates to traditional approaches [91].
The path forward requires greater integration of human-relevant biology into AI platforms. As noted in a 2025 Nature commentary, "If we do not adequately account for human responses and variability before going into the clinic then we should be prepared for our AI drugs to face the same failures in the clinic as traditionally developed molecules" [91]. Promising approaches include the incorporation of high-dimensional functional data from primary human cells, patient-derived tissue models, and real-world evidence [91] [93]. Additionally, causal AI frameworks that move beyond correlation to understand mechanistic biological relationships show potential for improving clinical translatability [95].
The coming 3-5 years will be decisive for AI-designed therapeutics as more candidates reach Phase II and III trials. Their performance in these critical efficacy trials will determine whether AI represents merely an efficiency tool or a transformative technology capable of finally improving the dismal success rates that have plagued drug development for decades.
This guide provides an objective, data-driven comparison of the performance of Generative AI and Active Learning approaches, with a specific focus on applications relevant to scientific research and drug development.
The following table summarizes the key quantitative performance indicators for Generative AI and Active Learning, highlighting their distinct impact areas.
Table 1: High-Level Performance Metric Comparison
| Metric | Generative AI | Active Learning |
|---|---|---|
| Speed / Productivity Gain | - 14% to 56% increase in task completion speed in professional tasks (e.g., coding, writing) [96].- AI tutoring enabled similar learning gains in ~49 minutes vs. 60 minutes for in-person active learning [1]. | - Reduces required labeled data by up to 42% compared to random sampling to achieve similar model accuracy [97]. |
| Cost Reduction | - Procurement functions report 15% to 45% cost reductions in goods and services [98].- Streamlines manual work in key processes by up to 30% [98]. | - Primary value is reducing data labeling cost, a significant bottleneck in machine learning projects [97]. |
| Success Rate / Quality | - In medical education, superior for practical skill development (SMD 0.63) compared to conventional methods [99].- Students reported higher engagement and motivation [1]. | - A rigorous statistical approach is required for comparison; no single strategy is best for all problems ("No Free Lunch" theorem) [100]. |
Detailed Quantitative Comparisons
This section breaks down the performance data across critical dimensions of speed, cost, and output quality.
Speed and Productivity Metrics
Table 2: Quantified Speed and Productivity Gains
| Technology / Approach | Application Context | Measured Outcome | Source |
|---|---|---|---|
| Generative AI | Customer Service (Chatbot) | 14% increase in tasks completed per hour [96]. | |
| Generative AI | Professional Writing | 40% increase in speed and 18% increase in output quality [96]. | |
| Generative AI | Software Development (GitHub Copilot) | 26% - 56% increase in task completion rate/speed [96]. | |
| Generative AI | Management Consulting | 25% increase in speed and 12% increase in task completion [96]. | |
| Generative AI | Procurement Analysis | Executes data analysis ~90% faster than manual processes [98]. | |
| Active Learning | Text Categorization & Document Classification | Can reduce the number of labeled documents required by up to 42% [97]. |
Cost Reduction and Economic Impact
Table 3: Quantified Cost and Economic Impact
| Technology / Approach | Impact Scope | Measured Outcome | Source |
|---|---|---|---|
| Generative AI | Macroeconomic Projection | Projected to increase productivity and GDP by 1.5% by 2035 and 3.7% by 2075 [96]. | |
| Generative AI | Corporate Procurement | 15% to 45% cost reduction in category spending; up to 30% reduction in manual work [98]. | |
| Active Learning | Machine Learning Project Budget | Directly targets and reduces the largest cost component: expert data labeling and annotation [97]. |
Success Rates and Output Quality
Evaluating the "success" of these technologies requires different metrics due to their fundamentally different outputs.
Generative AI Quality Metrics:
- Perplexity (PPL): Measures how well a language model predicts a sample. Lower values indicate better performance and higher confidence (e.g., a PPL of 4.73 for a short sentence is a relatively good score) [101].
- Fréchet Inception Distance (FID): Evaluates the quality of generated images by comparing statistical similarity to real images. Lower scores indicate higher fidelity and diversity [101].
- Bilingual Evaluation Understudy (BLEU): Measures the similarity of machine-generated text to high-quality human reference text [101].
Active Learning Performance Evaluation:
- Area Under the Learning Curve (ALC): A statistical method to compare the overall performance of different active learning strategies across multiple datasets, providing a more robust comparison than visual inspection of learning curves [100].
- Data Efficiency: The primary success metric is the rate of model accuracy improvement per unit of labeled data added. Effective strategies achieve higher accuracy with fewer labeled instances [97] [100].
Experimental Protocols and Methodologies
Protocol: Randomized Controlled Trial (RCT) for Educational Efficacy
This protocol is based on studies comparing AI tutors to active learning classrooms [1].
- Objective: To compare the efficacy of a Generative AI tutor against an in-class active learning session.
- Design: Randomized controlled trial with a crossover design.
- Population: Participants (e.g., 194 students) are divided into two groups.
- Intervention:
- Week 1: Group 1 uses the Generative AI tutor; Group 2 participates in an in-class active learning lesson.
- Week 2: The conditions are reversed.
- Controls: All participants complete a pre-test to establish baseline knowledge. The AI tutor and in-class lesson cover identical core content, informed by the same pedagogical principles.
- Metrics:
- Learning Gain: Post-test score vs. pre-test score.
- Speed: Time-on-task for the AI group (logged by platform) vs. fixed time for the in-class group.
- Perception: Post-lesson surveys on engagement, motivation, and enjoyment using a Likert scale.
- Analysis: Statistical tests (e.g., Mann-Whitney test, linear regression) are used to determine the significance of differences in learning gains and perception, controlling for pre-test scores and other factors [1].
Protocol: Evaluating Active Learning Strategies over Multiple Datasets
This protocol outlines a rigorous method for comparing Active Learning strategies [100].
- Objective: To determine if multiple active learning strategies differ significantly in performance across a range of datasets.
- Design: Benchmarking study.
- Materials:
nselection strategies (e.g., uncertainty sampling, query-by-committee) andmbenchmark datasets. - Workflow:
- For each dataset and strategy, run the active learning iterative process: a model is trained on a small initial set, the strategy selects the most informative instances for labeling, the model is updated, and its performance is recorded.
- This is repeated for a fixed number of iterations or until a performance target is met.
- Metrics:
- Performance Curve: Model accuracy (e.g., F1-score) as a function of the number of labeled instances used.
- Area Under the Learning Curve (ALC): A single aggregate metric representing overall performance.
- Analysis: Use non-parametric statistical tests (e.g., Friedman test with post-hoc Nemenyi test) to compare the ALC or the performance at different iterations across all datasets. This determines if observed differences are statistically significant [100].
The following diagram illustrates the core iterative workflow of an Active Learning system, as described in the experimental protocol.
The Scientist's Toolkit: Research Reagents & Solutions
Table 4: Essential Resources for AI and Machine Learning Research
| Item | Function / Description |
|---|---|
| Pre-trained Foundation Models (e.g., GPT-4, other LLMs) | Large-scale generative models that serve as a base for fine-tuning on specific scientific tasks, such as generating hypotheses or summarizing research papers [64] [99]. |
| Crowdsourcing Platforms (e.g., Amazon Mechanical Turk) | Online platforms used to efficiently collect and label large volumes of data at scale, which is crucial for training and evaluating both generative and active learning models [97]. |
| Active Learning Software Frameworks (e.g., JCLAL) | Java-based frameworks that provide standardized implementations of various active learning strategies, enabling reproducible benchmarking and experimentation [100]. |
| Benchmark Datasets | Curated, publicly available datasets with micro-level crowd votes or expert labels that are essential for the fair and comparative evaluation of different AI strategies [97] [100]. |
| Statistical Comparison Tools | Non-parametric statistical tests (e.g., Friedman, Nemenyi) used to rigorously validate that performance differences between algorithms are statistically significant and not due to chance [100]. |
Integrated Workflow: Combining Generative AI and Active Learning
The most powerful future applications may involve the synergy of both technologies. The following diagram proposes a hybrid workflow for a data-driven research project, such as in drug discovery.
The integration of artificial intelligence into educational methodologies has sparked significant debate regarding its efficacy compared to established teaching approaches. This comparison guide provides an objective analysis of generative AI versus active learning methodologies, examining their respective strengths, weaknesses, and optimal applications within research and scientific education contexts. Current research reveals that generative AI demonstrates superior efficiency and scalability for specific learning outcomes, while active learning maintains distinct advantages in fostering critical collaborative skills. The following comprehensive analysis synthesizes data from randomized controlled trials, meta-analyses, and qualitative studies to guide researchers, scientists, and drug development professionals in strategically deploying these educational approaches.
Active learning represents an established educational approach where students actively engage with the learning process through problem-solving, discussions, and other interactive activities rather than passively receiving information. This methodology has demonstrated significant improvements over traditional passive lectures across science, technology, engineering, and mathematics (STEM) disciplines [1]. In contrast, generative AI refers to artificial intelligence systems capable of creating new content and providing personalized instruction through sophisticated algorithms and large language models. These AI systems can simulate one-on-one tutoring experiences by adapting to individual learning paces and providing immediate feedback [102] [1].
The fundamental distinction between these approaches lies in their core operational mechanisms: active learning relies on human-facilitated collaborative engagement, while generative AI leverages computational power to deliver personalized, self-paced instruction. Understanding their comparative strengths and limitations enables research professionals and educators to make evidence-based decisions about their implementation in scientific training and development environments.
Quantitative Performance Comparison
Recent empirical studies directly comparing generative AI and active learning methodologies reveal distinct performance patterns across multiple metrics. The table below summarizes key quantitative findings from controlled experiments:
Table 1: Comparative Performance Metrics of Generative AI vs. Active Learning
| Performance Metric | Generative AI | Active Learning | Study Details |
|---|---|---|---|
| Knowledge Acquisition | No significant difference (SMD 0.27, p=0.36) [99] | No significant difference [99] | Meta-analysis of 11 RCTs with 786 medical students |
| Practical Skill Development | Significantly superior (SMD 0.63, p=0.02) [99] | Less effective [99] | Meta-analysis of medical education RCTs |
| Learning Gains | Over double the median gains [1] | Baseline median gains [1] | RCT in undergraduate physics (N=194) |
| Time Efficiency | Median 49 minutes (70% spent <60min) [1] | Fixed 60 minutes [1] | Same RCT measuring time on task |
| Student Engagement | Higher (Mean=4.1/5) [1] | Lower (Mean=3.6/5) [1] | Student perceptions on 5-point Likert scale |
| Student Motivation | Higher (Mean=3.4/5) [1] | Lower (Mean=3.1/5) [1] | Student perceptions on 5-point Likert scale |
| Test Score Improvement | 54% higher scores [2] | Baseline scores [2] | Analysis across multiple subjects |
The data indicates a nuanced performance relationship where generative AI demonstrates particular advantages in skill development, efficiency, and engagement, while both approaches show comparable effectiveness for fundamental knowledge acquisition. These findings suggest a complementary rather than exclusionary relationship between the methodologies.
Experimental Protocols and Methodologies
Randomized Controlled Trial in STEM Education
A rigorous RCT conducted at Harvard University provides insightful methodological framework for comparing these educational approaches [1]. The study employed a crossover design where students (N=194) experienced both teaching methodologies consecutively:
- Population: Undergraduate physics students broadly representative of diverse institutional populations
- Intervention: AI tutor delivering content through targeted prompt engineering based on pedagogical best practices
- Control: In-class active learning session employing peer instruction and small-group activities
- Content: Two distinct topics (surface tension and fluid flow) taught in consecutive weeks
- Assessment: Pre-test and post-test measurements for both groups to quantify learning gains
- Secondary Outcomes: Student perceptions of engagement, enjoyment, motivation, and growth mindset using 5-point Likert scales
The AI tutor incorporated seven research-based pedagogical principles: facilitating active learning, managing cognitive load, promoting growth mindset, scaffolding content, ensuring feedback accuracy, providing timely feedback, and allowing self-pacing [1].
Medical Education Meta-Analysis Protocol
A comprehensive meta-analysis compared GAI-based teaching with conventional methods in medical education [99]:
- Search Strategy: Systematic search across PubMed, Cochrane Library, EMBASE, and Web of Science (January 2014-January 2025)
- Selection Criteria: Focus on RCTs involving medical students with GAI interventions versus traditional teaching
- Primary Outcomes: Knowledge acquisition scores and practical skill development
- Secondary Outcomes: Student satisfaction with learning experience
- Risk Assessment: Cochrane Collaboration Risk of Bias tool with GRADE framework for evidence quality
- Statistical Analysis: Random-effects models with subgroup analyses based on geographical location, course classification, learning duration, and assessment timing
The analysis incorporated 11 eligible RCTs comprising 786 medical students, with ChatGPT being the primary GAI tool in 81.8% of studies [99].
Conceptual Relationship and Workflow
The following diagram illustrates the conceptual relationship between generative AI and active learning approaches, highlighting their distinctive characteristics and potential integration points:
Diagram 1: Conceptual Relationship Between Educational Approaches
Experimental Workflow for Comparative Studies
The methodology for conducting rigorous comparisons between generative AI and active learning involves specific experimental sequences:
Diagram 2: Experimental Workflow for Method Comparison
Research Reagent Solutions
For researchers designing studies in this domain, the following table outlines essential methodological components and their functions:
Table 2: Essential Research Components for Educational Methodology Studies
| Research Component | Function & Purpose | Implementation Examples |
|---|---|---|
| Randomized Controlled Trial (RCT) Design | Controls for confounding variables and establishes causality | Crossover designs where participants experience both conditions [99] [1] |
| Pre-Test/Post-Test Assessments | Measures baseline knowledge and learning gains | Standardized content knowledge tests with validated psychometrics [1] |
| Perception Metrics | Quantifies subjective learning experiences | 5-point Likert scales measuring engagement, motivation, and satisfaction [1] |
| Time-on-Task Measurement | Evaluates instructional efficiency | Platform analytics for AI groups; fixed time for in-person sessions [1] |
| Skill Assessment Rubrics | Objectively measures practical competencies | Standardized clinical skill evaluations in medical training [99] |
| Statistical Analysis Models | Determines significance of observed differences | Linear regression with controls for prior knowledge; quantile regression for ceiling effects [1] |
SWOT Analysis of Educational Methodologies
A comprehensive SWOT analysis synthesizes findings from multiple studies, particularly insights from nursing education research [103]:
Table 3: Comprehensive SWOT Analysis of Generative AI in Education
| Internal Factors | External Factors | |
|---|---|---|
| Positive | Strengths: • Personalized learning assistance [103]• Enhanced efficiency & cognitive expansion [103]• Superior practical skill development [99]• Self-paced instruction [1] | Opportunities: • Policy and resource support [103]• Technological advancement [103]• Addresses educational access gaps [2]• Emerging career applications [103] |
| Negative | Weaknesses: • Ethical and legal risks [103]• Potential for inaccurate outputs [103]• Reduced critical thinking development [104]• Technical barriers [103] | Threats: • Digital divide and equity gaps [103]• Job displacement concerns [103]• Educational integrity risks [103]• Plagiarism and academic misconduct [104] |
Discussion and Research Implications
Contextual Application in Scientific Training
For researchers, scientists, and drug development professionals, the comparative analysis between generative AI and active learning offers significant implications for specialized training environments. The superior performance of generative AI in practical skill development [99] suggests particular utility for laboratory technique training, diagnostic procedures, and protocol implementation where repetitive practice with immediate feedback enhances competency.
Conversely, active learning maintains advantages for developing collaborative research skills, interdisciplinary problem-solving, and experimental design capabilities where human interaction and creative negotiation are essential. The highest adoption of generative AI tools among PhD students and early-career academics for research purposes [104] further indicates its growing role in supporting advanced scientific training.
Limitations and Research Gaps
Current research exhibits several limitations requiring further investigation. The low quality of evidence in existing meta-analyses due to substantial heterogeneity across studies [99], inadequate reporting of randomization procedures, and limited implementation of blinding methods necessitates more rigorous experimental designs. Additionally, most studies focus on short-term knowledge retention, with inadequate assessment of long-term educational outcomes and translational impacts on professional practice.
The concentration of studies in specific disciplines (particularly medical education and physics) [99] [1] limits generalizability to specialized scientific domains such as pharmaceutical development and clinical research. Future studies should address these gaps through longitudinal designs, standardized outcome measures, and discipline-specific applications.
The evidence-based comparison between generative AI and active learning reveals a complementary relationship rather than a superiority contest. Generative AI demonstrates clear advantages in efficiency, scalability, and practical skill development, while active learning maintains importance for collaborative competencies and critical thinking skills.
For research organizations and educational institutions, strategic implementation should consider:
- Hybrid models that leverage AI-powered preparation and personalized follow-up while preserving high-value personal interaction
- Domain-specific applications where each methodology demonstrates particular strengths
- Resource optimization by deploying AI for scalable foundational training while reserving human facilitation for advanced conceptual development
- Continuous assessment to evaluate both quantitative learning outcomes and qualitative student experiences
The rapid evolution of generative AI technologies suggests its educational applications will continue to expand, potentially further enhancing its comparative advantages. However, the human-facilitated dimensions of active learning remain essential for developing the collaborative and critical thinking capabilities vital for scientific innovation and drug discovery breakthroughs.
The application of artificial intelligence in drug discovery represents one of the most promising yet challenging frontiers in pharmaceutical research. While traditional AI approaches have demonstrated value in specific tasks, they often operate in isolation, limiting their overall effectiveness. Generative AI, particularly through large language models and diffusion models, has shown remarkable capability in creating novel molecular structures and predicting properties [105] [106]. Simultaneously, active learning frameworks have proven exceptionally efficient at guiding experimental design by iteratively selecting the most informative data points for labeling, thereby maximizing knowledge gain while minimizing resource expenditure [107] [108]. However, when deployed as standalone solutions, both paradigms face significant limitations—generative models may produce chemically viable but practically irrelevant compounds, while active learning strategies can become trapped in local optima without sufficient exploratory mechanisms.
The fusion of these approaches represents a paradigm shift in AI-driven drug discovery. By integrating the creative capacity of generative models with the strategic, data-efficient sampling of active learning, researchers can create synergistic systems that outperform either method alone. This comparative guide examines the experimental evidence supporting this fusion approach, providing drug development professionals with a comprehensive analysis of performance metrics, methodological frameworks, and practical implementation strategies.
Comparative Analysis: Single-Paradigm Systems Versus Fused Approaches
Table 1: Performance comparison of single-paradigm AI systems versus fused approaches in drug discovery applications
| Performance Metric | Generative AI Alone | Active Learning Alone | Fused Approach |
|---|---|---|---|
| Novel Compound Identification Rate | High volume but variable relevance [106] | Limited by initial search space [108] | 60% synergistic pair discovery with only 10% combinatorial space exploration [33] |
| Data Efficiency | Requires extensive training data [105] | 70-95% reduction in data requirements for some endpoints [107] | 82% reduction in experimental materials and time [33] |
| Experimental Validation Success | Variable; limited by synthetic accessibility | Targeted but constrained exploration | 5-10x improvement in detecting highly synergistic combinations [33] |
| Optimization Cycles | Single-directional | Iterative but limited by initial model | Continuous refinement with dynamic exploration-exploitation balancing [108] |
| Resource Requirements | High computational costs for training [106] | Moderate computational, optimized experimental costs | Higher initial computational investment with significantly reduced experimental costs |
Table 2: Molecular property prediction accuracy across AI paradigms
| Prediction Task | Generative AI Performance | Active Learning Performance | Fused Approach Performance |
|---|---|---|---|
| Binding Affinity | Moderate accuracy (varies by model) [109] | Improved with iteration [108] | AGL-EAT-Score achieves high accuracy through graph learning [109] |
| Toxicity (hERG) | Attentive FP achieves high accuracy [109] | Gradual improvement through targeted sampling | CardioGenAI enables early identification and redesign [109] |
| Drug-Induced Liver Injury | Limited by data scarcity | Active data selection improves model robustness | StreamChol provides user-friendly toxicity estimation [109] |
| Solubility | Varies with training data quality | Hyperparameter optimization critical [109] | Preselected hyperparameters prevent overfitting [109] |
Experimental Protocols: Methodologies for Fusion Implementation
Protocol 1: Structure-Based Drug Design with Conditional Generation
The PoLiGenX methodology exemplifies the fusion approach in structure-based design by conditioning ligand generation on reference molecules within specific protein pockets [109]. This protocol integrates generative diffusion models with active learning criteria to ensure synthetic accessibility and binding compatibility.
Experimental Workflow:
- Initialization: Pre-train a generative diffusion model on known protein-ligand complexes from databases like Chembl
- Pocket Identification: Use sequence-based methods like CLAPE-SMB to identify binding sites without 3D structural data
- Conditional Generation: Generate ligand candidates conditioned on both the pocket characteristics and desired molecular properties
- Active Screening: Apply uncertainty-based query strategies (LCMD, Tree-based-R) to prioritize candidates for synthesis
- Iterative Refinement: Incorporate experimental results to retrain both the generative and screening models
Key Advantage: The PoLiGenX approach generates ligands with favorable poses showing reduced steric clashes and lower strain energies compared to those generated with diffusion models alone [109].
Protocol 2: Synergistic Drug Combination Discovery
The RECOVER framework demonstrates how active learning can guide experimental campaigns for identifying synergistic drug pairs, achieving 60% detection of synergistic pairs with only 10% combinatorial space exploration [33].
Experimental Workflow:
- Pre-training: Initialize model (MLP architecture) with public synergy data (Oneil, ALMANAC datasets)
- Feature Selection: Use Morgan fingerprints for molecular representation and gene expression profiles for cellular context
- Batch Selection: Employ exploration-exploitation strategies with dynamic tuning based on current performance
- Experimental Testing: Conduct high-throughput screening of selected batches (typically 50-100 combinations per batch)
- Model Updating: Refine model parameters iteratively using newly acquired experimental data
Critical Finding: Small batch sizes with dynamic exploration-exploitation tuning significantly enhance synergy yield ratios, with optimal performance achieved through uncertainty-driven sampling strategies [33].
Protocol 3: Molecular Optimization with Multi-Objective Active Learning
This protocol, as detailed by Reker et al., combines generative models with multi-objective active learning to simultaneously optimize multiple drug properties while maintaining synthetic feasibility [108].
Experimental Workflow:
- Initial Compound Selection: Start with a diverse set of lead compounds with partial property characterization
- Multi-Objective Model Training: Develop predictive models for each key property (efficacy, toxicity, PK/PD)
- Generative Expansion: Use transformer-based models (CardioGenAI) to generate structural analogs
- Pareto-Optimal Selection: Apply multi-criteria decision making to select compounds balancing all objectives
- Iterative Testing: Prioritize synthesis and testing based on predicted Pareto-frontier improvements
Performance Outcome: This approach has demonstrated successful re-engineering of drugs with known hERG liability while preserving pharmacological activity [109].
The Scientist's Toolkit: Essential Research Reagents and Solutions
Table 3: Key research reagents and computational tools for implementing fused AI approaches
| Tool/Reagent | Function | Application Context |
|---|---|---|
| Morgan Fingerprints | Molecular representation using circular atomic environments [33] | Feature encoding for ML models in synergy prediction and property estimation |
| Gene Expression Profiles (GDSC) | Cellular context features from Genomics of Drug Sensitivity in Cancer database [33] | Incorporating cellular environment into synergy and efficacy predictions |
| CLAPE-SMB | Protein-DNA binding site prediction using only sequence data [109] | Binding site identification without 3D structural information |
| Gnina (v1.3) | CNN-based scoring function for molecular docking poses [109] | Structure-based virtual screening with improved pose selection |
| AGL-EAT-Score | Algebraic graph learning scoring function for binding affinity prediction [109] | Predicting protein-ligand binding affinities using 3D sub-graphs |
| ChemBERTa | Pre-trained molecular representation model [33] | Transfer learning for molecular property prediction with limited data |
| UMAP Splitting | Data splitting strategy for more challenging benchmark evaluations [109] | Creating realistic train-test splits that better evaluate model generalizability |
| Monte Carlo Dropout | Uncertainty estimation technique for regression tasks [107] | Quantifying model uncertainty for active learning query strategies |
Visualization of Integrated Workflows
Discussion: Performance Advantages and Implementation Challenges
The experimental data consistently demonstrates that fused generative AI and active learning systems outperform single-paradigm approaches across multiple drug discovery metrics. The most significant advantage manifests in data efficiency – where traditional methods might require screening thousands of compounds, fused approaches can identify promising candidates with 70-95% fewer experimental measurements [107] [33]. This efficiency gain directly translates to reduced development costs and accelerated timelines.
A critical success factor involves balancing exploration versus exploitation throughout the drug discovery campaign. Early stages benefit from uncertainty-driven and diversity-based query strategies (LCMD, RD-GS), which broadly explore chemical space [107]. As models mature, shifting toward more exploitative strategies that refine promising regions of chemical space yields superior results. The most successful implementations dynamically adjust this balance based on real-time performance metrics.
Despite these advantages, implementation challenges remain. Data quality and representation significantly impact model performance, with molecular encoding strategies and cellular context features playing crucial roles [33]. Additionally, computational infrastructure requirements present barriers, particularly for small organizations, though cloud-based solutions and open-source models are gradually democratizing access [110]. Future developments in automated machine learning (AutoML) and transfer learning promise to further reduce these barriers while improving model performance across diverse drug discovery contexts.
The fusion of generative AI with active learning represents more than a technical improvement – it constitutes a fundamental shift in how drug discovery is approached. By creating a continuous, adaptive cycle of computational prediction and experimental validation, this integrated approach maximizes the value of both computational and experimental resources, ultimately accelerating the delivery of novel therapeutics to patients.
The drug discovery landscape is undergoing a fundamental transformation, moving from a domain reliant on labor-intensive, human-driven workflows to one increasingly powered by artificial intelligence (AI) discovery engines capable of compressing timelines and expanding chemical and biological search spaces [11]. By 2025, AI has progressed from an experimental curiosity to a tangible force, with AI-designed therapeutics now advancing through human trials across diverse therapeutic areas [11]. This shift is characterized by a transition from traditional reductionist approaches, which focus on narrow tasks like fitting a ligand into a protein pocket, toward a more holistic, systems biology level that uses deep learning to integrate multimodal data—including phenotypic, omics, patient data, chemical structures, and text [93]. The core value proposition of modern AI-driven drug discovery (AIDD) platforms lies in their ability to be scalable, to represent biology in silico with sufficient depth and breadth, and to deliver sustainable value in a repeatable, standardized way across R&D workflows [93].
Within this broader transformation, two distinct yet increasingly synergistic technological approaches have emerged: generative AI and active learning (AL). This guide provides an objective comparison of these methodologies, evaluating their performance, applications, and experimental protocols to inform strategic decision-making for researchers, scientists, and drug development professionals.
Comparative Analysis: Generative AI vs. Active Learning
The table below summarizes the core characteristics, strengths, and challenges of generative AI and active learning approaches, providing a high-level comparison for researchers.
Table 1: Core Characteristics of Generative AI and Active Learning in Drug Discovery
| Feature | Generative AI | Active Learning |
|---|---|---|
| Primary Objective | De novo design of novel molecular structures with tailored properties [12]. | Optimized data selection to improve model efficiency and guide experimentation [108]. |
| Core Function | Creates previously unseen molecules from a learned chemical space [12]. | Selects the most informative data points for labeling from a larger dataset [108]. |
| Typical Architecture | Variational Autoencoders (VAEs), Generative Adversarial Networks (GANs), Reinforcement Learning (RL), Transformers [93] [12]. | Iterative feedback loop combining a predictive model with a query strategy [108]. |
| Key Advantage | Explores vast and novel chemical spaces, enabling discovery of unprecedented scaffolds [12]. | Maximizes information gain while minimizing resource use; highly data-efficient [12] [33]. |
| Primary Challenge | Ensuring synthetic accessibility, target engagement, and generalizability beyond training data [12]. | Defining optimal query strategies and managing the exploration-exploitation trade-off [33]. |
| Ideal Application | Hit identification and lead optimization where novel chemistry is required. | Virtual screening and synergistic drug combination discovery in low-data regimes [33]. |
Performance and Clinical Validation
The ultimate validation of any drug discovery technology is its ability to deliver clinically viable candidates. The following table quantifies the performance and track records of leading platforms, many of which integrate both generative and active learning components.
Table 2: Performance Metrics and Clinical Progress of Leading AI-Driven Platforms (2024-2025)
| Company / Platform | Core AI Approach | Key Clinical Progress & Performance Data |
|---|---|---|
| Insilico Medicine (Pharma.AI) | Generative AI (GANs, RL), Knowledge Graphs, NLP [11] [93]. | ISM001-055 (TNIK inhibitor for IPF): From target discovery to Phase I in 18 months; Positive Phase IIa results in 2025 [11]. |
| Exscientia | Generative AI, Centaur Chemist, Patient-derived biology [11]. | AI-designed clinical compounds developed "at a pace substantially faster than industry standards"; Design cycles ~70% faster, requiring 10x fewer synthesized compounds [11]. |
| Schrödinger | Physics-based ML, Molecular Simulation [11]. | Nimbus-originated TYK2 inhibitor (Zasocitinib) advanced to Phase III trials [11]. |
| Recursion (OS Platform) | Phenomic Screening, Deep Learning on Cellular Imaging [11] [93]. | Phenom-2 model trained on 8 billion microscopy images, claims 60% improvement in genetic perturbation separability [93]. |
| BenevolentAI | Knowledge-Graph-Driven Target Discovery [11]. | Identified Baricitinib (RA drug) for repurposing in COVID-19, leading to emergency use authorization [92]. |
| VAE-AL GM Workflow [12] | Generative AI (VAE) nested with Active Learning cycles. | For CDK2: 9 molecules synthesized, 8 showed in vitro activity (1 with nanomolar potency). For KRAS: 4 molecules with potential activity identified in silico [12]. |
| RECOVER (AL Framework) [33] | Active Learning for Synergistic Drug Combinations. | Discovered 60% of synergistic drug pairs by exploring only 10% of the combinatorial space, saving 82% in experimental time and materials [33]. |
Experimental Protocols and Workflows
Integrated Generative AI with Active Learning Framework
A landmark 2025 study demonstrated a robust workflow merging a generative Variational Autoencoder (VAE) with two nested active learning cycles to overcome common limitations of generative models, such as poor target engagement and synthetic inaccessibility [12]. The detailed methodology is as follows and visualized in the diagram below:
- Data Representation & Initial Training: Training molecules are represented as SMILES strings, tokenized, and converted into one-hot encoding vectors. The VAE is first trained on a general dataset and then fine-tuned on a target-specific training set [12].
- Molecule Generation & Inner AL Cycle: The trained VAE is sampled to generate new molecules. An inner AL cycle evaluates these molecules using chemoinformatic oracles (e.g., for drug-likeness, synthetic accessibility). Molecules meeting thresholds are added to a "temporal-specific set" to fine-tune the VAE, prioritizing desired properties [12].
- Outer AL Cycle: After several inner cycles, an outer AL cycle begins. Molecules in the temporal-specific set are evaluated by a physics-based affinity oracle (e.g., molecular docking simulations). High-scoring molecules are transferred to a "permanent-specific set," which is used for further VAE fine-tuning, creating a feedback loop that refines the search for high-affinity compounds [12].
- Candidate Selection: Post-cycling, the most promising candidates from the permanent-specific set undergo stringent filtration and intensive molecular modeling simulations (e.g., PELE, Absolute Binding Free Energy calculations) for in-depth evaluation before synthesis and in vitro testing [12].
Diagram 1: Integrated VAE-AL Workflow
Active Learning for Synergistic Drug Combination Screening
For complex tasks like discovering synergistic drug pairs—a rare phenomenon in a vast combinatorial space—a pure AL framework has proven highly effective. The RECOVER framework provides a detailed protocol [33]:
- Problem Setup & Model Pre-training: The goal is to sequentially select batches of drug combinations to test experimentally. The AL model (e.g., an MLP) is first pre-trained on an existing dataset like Oneil, using molecular fingerprints (e.g., Morgan fingerprints) and cellular features (e.g., gene expression profiles from GDSC) as input to predict synergy scores [33].
- Iterative Batch Selection and Testing:
- Acquisition Function: The trained model evaluates all untested drug pairs. An acquisition function (e.g., expected improvement, upper confidence bound) balances exploration (testing uncertain pairs) and exploitation (testing pairs predicted to be highly synergistic) to select the most "informative" batch for the next round of experiments [33].
- Experimental Testing & Model Update: The selected batch of drug combinations is tested in vitro, and the experimental synergy scores are obtained. This new, high-quality data is added to the training set, and the model is retrained. This iterative feedback loop progressively improves the model's predictive accuracy for the specific experimental context [33].
- Key Parameters: Performance is highly dependent on batch size; smaller batches allow for more dynamic strategy tuning and higher synergy yield. The choice of cellular features (e.g., using gene expression profiles) was found to be more critical for prediction quality than the specific type of molecular encoding [33].
Diagram 2: Active Learning Cycle
The Scientist's Toolkit: Essential Research Reagents and Materials
The successful implementation of the experimental protocols above relies on a suite of computational and empirical tools. The table below details key resources for setting up similar experiments.
Table 3: Essential Research Reagents and Solutions for AI-Driven Experiments
| Item / Resource | Function / Application | Experimental Context |
|---|---|---|
| Molecular Databases (e.g., ChEMBL, DrugComb) | Provide structured bioactivity and drug combination data for model pre-training and benchmarking [33]. | Foundational for all AI workflows; used for initial model training [12] [33]. |
| Fingerprint & Descriptor Tools (e.g., RDKit for Morgan Fingerprints, MAP4) | Encode molecular structures into numerical representations (features) for machine learning models [33]. | Critical pre-processing step; Morgan fingerprints noted for strong performance in synergy prediction [33]. |
| Cellular Feature Data (e.g., GDSC gene expression profiles) | Provide genomic context of the targeted cell line, significantly improving prediction accuracy in phenotypic screens [33]. | Used as model input to account for the cellular environment in synergy prediction and phenotypic analysis [33]. |
| Cheminformatic Oracles | Computational filters for properties like drug-likeness (e.g., Lipinski's Rule of 5) and synthetic accessibility (SA) score [12]. | Used within AL cycles to filter generated molecules before expensive physics-based evaluations [12]. |
| Physics-Based Affinity Oracles | Molecular docking software (e.g., AutoDock) and molecular dynamics simulations (e.g., PELE) to predict binding affinity and pose [12] [111]. | Act as a proxy for experimental affinity measurements within AL cycles; used for candidate prioritization [12]. |
| CETSA (Cellular Thermal Shift Assay) | Validates direct target engagement of drug candidates in intact cells, providing physiologically relevant binding data [111]. | Used post-synthesis for experimental validation, bridging the gap between in silico prediction and cellular efficacy [111]. |
Investment Trends and Strategic Outlook
The investment landscape and strategic direction of AIDD are crystallizing around several key trends, as captured in the 2025 Stanford AI Index Report and industry analyses [84].
- Record Investment and Industry Adoption: Private investment in AI continues to soar, with the U.S. leading at $109.1 billion in 2024—nearly 12 times China's investment [84]. Generative AI alone attracted $33.9 billion globally [84]. Business usage is accelerating, with 78% of organizations reporting AI use in 2024, up from 55% the year before [84].
- Platform Integration and M&A: A significant trend is the consolidation of complementary technologies to create end-to-end platforms. The acquisition of Exscientia by Recursion in a $688 million merger is a prime example, aiming to combine generative chemistry with extensive phenomic data into a unified "AI drug discovery superpower" [11].
- Focus on Clinical Translation and Validation: The field is maturing from proving discovery speed to demonstrating clinical success. The advancement of multiple AI-derived candidates into Phase II and III trials (e.g., from Insilico Medicine, Schrödinger) is a critical focus for investors and partners seeking to de-risk the technology [11] [60].
- Rising Regulatory Attention: As AI's influence grows, so does regulatory scrutiny. In 2024, U.S. federal agencies introduced 59 AI-related regulations—more than double the number in 2023—signaling the need for clear development and validation frameworks [84].
The trajectory of AI-driven drug discovery is defined by a clear evolution from promising tool to essential platform. The comparative analysis reveals that while generative AI and active learning serve distinct primary functions—the former for creative molecular design and the latter for intelligent experimentation—their integration represents the most powerful frontier. Platforms that successfully merge generative chemistry with iterative, data-driven experimental feedback are demonstrating tangible accelerations in preclinical timelines and improved success rates in early clinical trials [11] [12].
For researchers and drug development professionals, the strategic implication is that investment in both capabilities is crucial. Building or partnering with platforms that offer a closed-loop, holistic system—capable of generating novel hypotheses, prioritizing the most informative experiments, and continuously learning from the resulting data—is no longer a speculative venture but a core competitive advantage. The recent surge in investment, M&A activity, and regulatory focus underscores that AI-driven discovery is the foundational paradigm for the future of pharmaceutical R&D [11] [84].
Conclusion
Generative AI and Active Learning are not mutually exclusive but are increasingly powerful when integrated. Generative AI serves as a prolific idea generator, rapidly exploring vast chemical spaces, while Active Learning acts as a precision guide, ensuring resources are allocated to the most informative data points. This synergy is already compressing drug discovery timelines from years to months and demonstrating tangible clinical success. The future of pharmaceutical R&D lies in hybrid models that leverage the creativity of generative design with the efficient, targeted validation of active learning. For researchers and drug development professionals, mastering the strategic application and integration of these technologies is no longer optional but essential for remaining at the forefront of medical innovation and addressing the world's most pressing healthcare challenges.
References
- 1. AI tutoring outperforms in-class active learning [https://www.nature.com/articles/s41598-025-97652-6]
- 2. 20 Statistics on AI in Education to Guide Your Learning ... [https://www.engageli.com/blog/ai-in-education-statistics]
- 3. Active Learning Statistics: Benefits for Education & Training ... [https://www.engageli.com/blog/active-learning-statistics-2025]
- 4. The State of Artificial Intelligence in 2025 [https://www.baytechconsulting.com/blog/the-state-of-artificial-intelligence-in-2025]
- 5. Active Learning Methods for Efficient Data Utilization and ... [https://arxiv.org/abs/2504.16136]
- 6. Impact of AI Tools on Learning Outcomes [https://arxiv.org/html/2510.16019v1]
- 7. How Much does it Cost to Build a Generative AI in 2025 : Aalpha [https://www.aalpha.net/blog/cost-to-build-a-generative-ai/]
- 8. Generative AI Statistics: Insights and Emerging Trends for ... [https://hatchworks.com/blog/gen-ai/generative-ai-statistics/]
- 9. Eroom's law [https://en.wikipedia.org/wiki/Eroom%27s_law]
- 10. Overcoming the Declining Trends in Innovation and ... [https://www.sciencedirect.com/science/article/pii/S2452302X17302231]
- 11. Leading AI-Driven Drug Discovery Platforms [https://www.sciencedirect.com/science/article/abs/pii/S0031699725075118]
- 12. Optimizing drug design by merging generative AI with a ... [https://www.nature.com/articles/s42004-025-01635-7]
- 13. MIT scientists debut a generative AI model that could ... [https://news.mit.edu/2025/mit-scientists-debut-generative-ai-model-that-could-create-molecules-addressing-hard-to-treat-diseases-1125]
- 14. A Survey of Generative AI for de novo Drug Design: New Frontiers in Molecule and Protein Generation [https://arxiv.org/html/2402.08703v1]
- 15. Generative Deep Learning for de Novo Drug Design A ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12308806/]
- 16. [2402.08703] A Survey of Generative AI for de novo Drug Design: New Frontiers in Molecule and Protein Generation [https://arxiv.org/abs/2402.08703]
- 17. Generative artificial intelligence based models optimization towards molecule design enhancement | Journal of Cheminformatics | Full Text [https://jcheminf.biomedcentral.com/articles/10.1186/s13321-025-01059-4]
- 18. Optimal Molecular Design: Generative Active Learning ... [https://pubmed.ncbi.nlm.nih.gov/39225482/]
- 19. Generative AI for drug discovery and protein design [https://www.sciencedirect.com/science/article/pii/S2590098625000107]
- 20. A review of active learning approaches to experimental ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5453429/]
- 21. Training-Free Active Learning Framework in Materials ... [https://arxiv.org/html/2511.19730v1]
- 22. An Interactive Visualization Tool for Understanding Active ... [https://arxiv.org/abs/2111.04936]
- 23. An Introduction to High Contrast Accessibility [https://dev.to/mpriour/an-introduction-to-high-contrast-accessibility-4im4]
- 24. Button states in High contrast mode - General Help [https://community.evolveauthoring.com/t/button-states-in-high-contrast-mode/2166]
- 25. Generative Artificial Intelligence | Center for Teaching Innovation [https://teaching.cornell.edu/generative-artificial-intelligence]
- 26. Effect of Generative Artificial Intelligence on University ... [https://www.sciencedirect.com/science/article/abs/pii/S1747938X25000740]
- 27. Machine learning and generative AI: What are they good for in 2025? | MIT Sloan [https://mitsloan.mit.edu/ideas-made-to-matter/machine-learning-and-generative-ai-what-are-they-good-for]
- 28. The State of AI: Global Survey 2025 [https://www.mckinsey.com/capabilities/quantumblack/our-insights/the-state-of-ai]
- 29. Top Generative AI Skills and Education Trends for 2025 [https://aws.amazon.com/executive-insights/content/top-generative-ai-skills-and-education-trends-for-2025/]
- 30. Generative AI in Education: Use Cases, Benefits, and ... [https://www.fullestop.com/blog/generative-ai-in-education-use-cases-benefits-and-challenges]
- 31. Artificial Intelligence in Small-Molecule Drug Discovery [https://www.mdpi.com/1424-8247/18/9/1271]
- 32. Generative AI in Drug Discovery: Transforming Pharma R&D [https://www.delveinsight.com/blog/generative-ai-drug-discovery-market-impact]
- 33. A guide for active learning in synergistic drug discovery [https://www.nature.com/articles/s41598-025-85600-3]
- 34. Active Learning FEP: Impact on Performance of AL ... [https://pubmed.ncbi.nlm.nih.gov/40243401/]
- 35. Progress of AI-Driven Drug–Target Interaction Prediction and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12563295/]
- 36. STAR- VAE : Latent Variable Transformers for Scalable and... [https://arxiv.org/html/2511.02769v1]
- 37. Deep generative molecular design reshapes drug discovery [https://pmc.ncbi.nlm.nih.gov/articles/PMC9797947/]
- 38. Active Learning Enables Extrapolation in Molecular ... [https://arxiv.org/abs/2501.02059]
- 39. Bayesian, Autoregressive, Comparing , VAEs ... | atalupadhyay GANs [https://atalupadhyay.wordpress.com/2025/01/15/comparing-bayesian-autoregressivevaesgans-and-transformers-in-generative-ai/]
- 40. Generative Models Explained: VAEs, GANs, Diffusion ... [https://bestarion.com/us/generative-models-explained-vaes-gans-diffusion-transformers-autoregressive-models-nerfs/]
- 41. Transformer graph variational autoencoder for generative ... [https://www.sciencedirect.com/science/article/pii/S0006349525000359]
- 42. A Reinforcement Learning-Driven Transformer GAN for ... [https://arxiv.org/html/2503.12796v1]
- 43. Enhancing molecular design efficiency: Uniting language ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11026973/]
- 44. A Review Of Active Learning Strategies [https://www.sagacify.com/news/what-is-active-learning]
- 45. Active Learning Overview: Strategies and Uncertainty ... [https://medium.com/data-science/active-learning-overview-strategies-and-uncertainty-measures-521565e0b0b]
- 46. An Active Learning Approach with Uncertainty ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4144157/]
- 47. Learning with not Enough Data Part 2: Active Learning | Lil'Log [https://lilianweng.github.io/posts/2022-02-20-active-learning/]
- 48. Active Learning and Uncertainty Sampling [https://keymakr.com/blog/uncertainty-sampling-explained/]
- 49. [2506.03817] Survey of Active Learning Hyperparameters [https://arxiv.org/abs/2506.03817]
- 50. Introducing Nested Learning: A new ML paradigm for ... [https://research.google/blog/introducing-nested-learning-a-new-ml-paradigm-for-continual-learning/]
- 51. The Architecture of the Mind: Google's "Nested Learning ... [https://www.executeai.software/the-architecture-of-the-mind-googles-nested-learning-and-the-global-race-for-continual-intelligence/]
- 52. The Active Learning–GenAI Synergy Framework [https://f1000research.com/articles/14-1210]
- 53. Comparative Analysis of Generative Artificial Intelligence ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12288765/]
- 54. How Generative AI Is Reducing Drug Discovery Timelines by ... [https://prajnaaiwisdom.medium.com/how-generative-ai-is-reducing-drug-discovery-timelines-by-70-e86d58f7c780]
- 55. From Start to Phase 1 in 30 Months [https://insilico.com/phase1]
- 56. Insilico Medicine Uses Generative AI to Accelerate Drug ... [https://blogs.nvidia.com/blog/insilico-medicine-uses-generative-ai-to-accelerate-drug-discovery/]
- 57. Insilico Pharma.AI fall launch recap: Understand latest AI ... [https://insilico.com/tpost/003kiu5pv1-insilico-pharmaai-fall-launch-recap-unde]
- 58. Top 10 AI Drug Discovery Platforms in 2025 [https://www.devopsschool.com/blog/top-10-ai-drug-discovery-platforms-in-2025-features-pros-cons-comparison/]
- 59. Exscientia Uses Generative AI to Reimagine Drug Discovery [https://aws.amazon.com/solutions/case-studies/exscientia-generative-ai/]
- 60. From Lab to Clinic: How Artificial Intelligence (AI) Is ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12298131/]
- 61. 12 AI drug discovery companies you should know about [https://www.labiotech.eu/best-biotech/ai-drug-discovery-companies/]
- 62. How AI is transforming drug discovery [https://www.drugdiscoverynews.com/ai-is-transforming-drug-discovery-16706]
- 63. How Generative AI in Healthcare is Transforming Drug ... [https://www.rfidjournal.com/expert-views/how-generative-ai-in-healthcare-is-transforming-drug-discovery-in-2025/222589/]
- 64. Generative AI vs Machine Learning: A Comparison Guide [https://magai.co/generative-ai-vs-machine-learning/]
- 65. AI Driven Drug Discovery: 5 Powerful Breakthroughs in 2025 [https://lifebit.ai/blog/ai-driven-drug-discovery/]
- 66. Editorial: Approaches to improve the performance of virtual ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12378871/]
- 67. How AI Is Irrevocably Altering the DNA of Drug Development [https://www.drugpatentwatch.com/blog/how-ai-is-already-changing-drug-development/?srsltid=AfmBOopKa0ua1F4hy-WjvPjZ-6rBr8njhHIw6aKK_ottMQjG2MO74knp]
- 68. Deep Batch Active Learning for Drug Discovery [https://elifesciences.org/reviewed-preprints/89679v2]
- 69. How successful are AI-discovered drugs in clinical trials? A ... [https://www.sciencedirect.com/science/article/pii/S135964462400134X]
- 70. How in silico trials are reshaping clinical development [https://www.zs.com/insights/in-silico-trials-clinical-research]
- 71. AI Benchmarks 2025: Performance Metrics Show Record ... [https://www.sentisight.ai/ai-benchmarks-performance-soars-in-2025/]
- 72. Systematic literature review on bias mitigation in ... [https://link.springer.com/article/10.1007/s43681-025-00721-9]
- 73. The State Of AI Costs In 2025 [https://www.cloudzero.com/state-of-ai-costs/]
- 74. Integrating artificial intelligence into small molecule ... [https://www.nature.com/articles/s44386-025-00029-y]
- 75. Artificial Intelligence in Small-Molecule Drug Discovery [https://pmc.ncbi.nlm.nih.gov/articles/PMC12472608/]
- 76. SynFrag: Synthetic Accessibility Predictor based on ... [https://chemrxiv.org/engage/chemrxiv/article-details/68e867fddfd0d042d1c5032e]
- 77. Synthetic Accessibility: Definition, Importance, and How to ... [https://neurosnap.ai/blog/post/synthetic-accessibility-definition-importance-and-how-to-assess-it-with-neurosnap/68cb0899cec8a8b395412c77]
- 78. Chemistry-aware AI offers new routes in small molecule ... [https://www.drugtargetreview.com/article/189379/chemistry-aware-ai-offers-new-routes-in-small-molecule-design/]
- 79. Synthetic accessibility scoring and its application to ... [https://pubs.rsc.org/en/content/articlelanding/2025/ta/d5ta01042j]
- 80. Deep Dive into Active Learning Agents for 2025 [https://sparkco.ai/blog/deep-dive-into-active-learning-agents-for-2025]
- 81. A comprehensive benchmark of active learning strategies ... [https://www.nature.com/articles/s41598-025-24613-4]
- 82. Working with Generative AI Models: Requirements & Tips [https://www.iiot-world.com/artificial-intelligence-ml/artificial-intelligence/ow-to-work-with-generative-ai-models/]
- 83. What Hardware Is Needed for AI? - Multimodal [https://www.multimodal.dev/post/what-hardware-is-needed-for-ai]
- 84. The 2025 AI Index Report | Stanford HAI [https://hai.stanford.edu/ai-index/2025-ai-index-report]
- 85. Investigational New Drug (IND) Application [https://www.fda.gov/drugs/types-applications/investigational-new-drug-ind-application]
- 86. Search for FDA Guidance Documents [https://www.fda.gov/regulatory-information/search-fda-guidance-documents]
- 87. How will FDA changes reshape drug approval in 2025 and ... [https://cen.acs.org/pharmaceuticals/drug-development/FDA-changes-reshape-drug-approval/103/web/2025/05]
- 88. The future of pharmaceuticals: Artificial intelligence in drug ... [https://www.sciencedirect.com/science/article/pii/S2095177925000656]
- 89. Artificial intelligence in drug discovery and development - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC7577280/]
- 90. Artificial intelligence in drug development | Nature Medicine [https://www.nature.com/articles/s41591-024-03434-4]
- 91. The AI drug revolution needs a revolution [https://www.nature.com/articles/s44386-025-00013-6]
- 92. Transformative Role of Artificial Intelligence in Drug ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12406033/]
- 93. Beyond Legacy Tools: Defining Modern AI Drug Discovery ... [https://www.biopharmatrend.com/business-intelligence/what-is-ai-drug-discovery/]
- 94. AI in Pharmaceutical Industry: 2025 Guide & Use Cases [https://sranalytics.io/blog/ai-in-pharmaceutical-industry/]
- 95. The next generation of clinical trials with AI [https://www.ddw-online.com/the-next-generation-of-clinical-trials-with-ai-36171-202508/]
- 96. The Projected Impact of Generative AI on Future ... [https://budgetmodel.wharton.upenn.edu/issues/2025/9/8/projected-impact-of-generative-ai-on-future-productivity-growth]
- 97. A review and experimental analysis of active learning over ... [https://link.springer.com/article/10.1007/s10462-021-10021-3]
- 98. From Buzz to Bottom Line - Cost Savings Using GenAI [https://www.bcg.com/publications/2025/from-buzz-to-bottom-line-cost-reductions-using-genai]
- 99. Effectiveness of generative artificial intelligence-based ... [https://bmcmededuc.biomedcentral.com/articles/10.1186/s12909-025-07750-2]
- 100. Statistical comparisons of active learning strategies over ... [https://www.sciencedirect.com/science/article/abs/pii/S0950705118300492]
- 101. The Top 11 AI Metrics for Generative AI [https://encord.com/blog/generative-ai-metrics/]
- 102. Traditional AI vs. Generative AI: What's the Difference? | Illinois [https://education.illinois.edu/about/news-events/news/article/2024/11/11/what-is-generative-ai-vs-ai]
- 103. a qualitative study of undergraduate nursing students - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC12443828/]
- 104. Is generative AI reshaping academic practices worldwide? ... [https://www.sciencedirect.com/science/article/pii/S0306457325002912]
- 105. 2025 – Comparison Guide Between LLM vs Generative AI [https://www.iqudtek.com/blog/2025-comparison-guide-between-llm-vs-generative-ai/]
- 106. The Two Main Types of Generative AI: Comparison and ... [https://academyocean.com/blog/post/the-two-main-types-of-generative-ai-comparison-and-examples]
- 107. A comprehensive benchmark of active learning strategies ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12550062/]
- 108. The present state and challenges of active learning in drug ... [https://www.sciencedirect.com/science/article/abs/pii/S1359644624001107]
- 109. Advanced machine learning for innovative drug discovery [https://pmc.ncbi.nlm.nih.gov/articles/PMC12333061/]
- 110. Top Generative AI Trends to Watch in 2025 and Beyond [https://www.silverwebbuzz.com/blog/top-generative-ai-trends/]
- 111. Top 5 Drug Discovery Trends 2025 Driving Breakthroughs [https://www.pelagobio.com/cetsa-drug-discovery-resources/blog/drug-discovery-trends-2025/]